Abstract
Background
COVID-19 severity varies widely. Although some demographic and cardio-metabolic factors, including age and obesity, are associated with increasing risk of severe illness, the underlying mechanism(s) are uncertain.
Subjects/methods
In a meta-analysis of three independent studies of 1471 participants in total, we investigated phenotypic and genetic factors associated with subcutaneous adipose tissue expression of Angiotensin I Converting Enzyme 2 (ACE2), measured by RNA-Seq, which acts as a receptor for SARS-CoV-2 cellular entry.
Results
Lower adipose tissue ACE2 expression was associated with multiple adverse cardio-metabolic health indices, including type 2 diabetes (T2D) (P = 9.14 × 10−6), obesity status (P = 4.81 × 10−5), higher serum fasting insulin (P = 5.32 × 10−4), BMI (P = 3.94 × 10−4), and lower serum HDL levels (P = 1.92 × 10−7). ACE2 expression was also associated with estimated proportions of cell types in adipose tissue: lower expression was associated with a lower proportion of microvascular endothelial cells (P = 4.25 × 10−4) and higher proportion of macrophages (P = 2.74 × 10−5). Despite an estimated heritability of 32%, we did not identify any proximal or distal expression quantitative trait loci (eQTLs) associated with adipose tissue ACE2 expression.
Conclusions
Our results demonstrate that individuals with cardio-metabolic features known to increase risk of severe COVID-19 have lower background ACE2 levels in this highly relevant tissue. Reduced adipose tissue ACE2 expression may contribute to the pathophysiology of cardio-metabolic diseases, as well as the associated increased risk of severe COVID-19.
Similar content being viewed by others
Introduction
Since December 2019 the COVID-19 pandemic has swept the world, with over 500 million confirmed cases worldwide and more than six million fatalities reported at the time of writing [1]. A distinctive and challenging aspect of COVID-19 is its wide disease spectrum [2]. While initial reports focused on respiratory symptoms [3, 4], it is now clear that severe COVID-19 also affects the cardiovascular and renal systems substantially [5]. Marked differences in clinical outcomes have been associated with demographic features including age and sex, with older age and male sex having been associated with increased risk of severe COVID-19, as have comorbidities including cardiovascular disease, diabetes, and obesity [2, 4, 5].
ACE2 has become central in understanding COVID-19 pathogenesis, as SARS-CoV-2 employs ACE2 as a receptor for cellular entry [6, 7]. ACE2 is expressed in a number of tissues and cell types [6, 8,9,10,11]. It is currently unclear whether higher ACE2 levels are harmful or beneficial in the context of COVID-19. On the one hand, higher levels of ACE2 may provide additional targets enabling viral invasion of ACE2-expressing cells; on the other, higher ACE2 has beneficial effects in regulation of the renin-angiotensin system (RAS), through controlling hypertension and associated cardio-metabolic disorders [12,13,14].
Obesity, defined as a body mass index (BMI) above 30 kg/m2 [15], is one of the strongest reported risk factors for severe COVID-19 [2, 5], but the mechanism underlying this remains unclear. In the Genotype-Tissue Expression (GTEx) resource, capturing gene expression in 54 tissues, adipose tissue ranks among the highest sites of ACE2 expression in the body [11, 16] (Supplementary Fig. 1). Several potential mechanisms have been proposed for the contribution of adiposity to COVID-19 severity [17,18,19,20,21], which may be multi-factorial. Critically, ACE2 in adipose tissue is key in balancing local adipose RAS, disruption of which may then lead to wider systemic RAS effects [18]. Adipose tissue is also a prime candidate tissue contributor to the widely-reported cytokine storm characteristic of severe COVID-19 [17, 18]. Most importantly, obesity is a potentially modifiable risk factor for COVID-19 severity. Thus, deeper understanding of the contribution of adipose tissue to COVID-19 severity, and to cardio-metabolic risk more generally, may inform better therapeutic strategies, and motivate policies supporting healthy weight achievement and maintenance programmes.
Given the dual relevance of ACE2, in maintenance of cardio-metabolic health generally and as SARS-CoV-2 receptor specifically, it is of interest to explore whether adipose tissue ACE2 expression is associated with specific demographic and phenotypic traits also associated with COVID-19 severity. In this study, we therefore investigated phenotypic and genetic factors associated with ACE2 gene expression in subcutaneous adipose tissue from a total of 1,471 participants drawn from three cohorts [22,23,24], representing the largest meta-analysis of adipose tissue gene expression studies to date.
Materials and methods
Sample collection
TwinsUK
Gene expression data, measured by RNA-Seq, was available for up to 804 female twins from the TwinsUK cohort [25]. Sample collection and processing in TwinsUK has been reported fully elsewhere [25]. Briefly, punch biopsies were taken from a sun-protected area of the abdomen from each participant. Subcutaneous adipose tissue and skin were separated and RNA extracted for each tissue type. Lymphoblastoid cell lines (LCL) were generated from blood samples collected at the time of biopsy. Gene expression in adipose tissue (n = 765), skin (n = 706), LCLs (n = 804), and whole blood (n = 389) was measured by RNA sequencing, as previously described [25].
METSIM
Sample collection and processing has previously been reported for the population-based METSIM cohort, composed of 10,197 males of Finnish ancestry [26]. RNA-sequencing of subcutaneous adipose tissue for 426 male participants collected near the umbilicus by needle biopsy was generated as previously described [23].
FUSION
331 living FUSION participants were recruited from Helsinki, Savitaipale, and Kuopio, as previously described [24]. Adipose tissue biopsies were obtained and processed from 296 of the 331 participants following the same general protocol as previously described for the concurrently-obtained muscle biopsies [24]. Biopsies were taken under local anaesthetic, by surgical scalpel from abdominal subcutaneous fat 5–10 cm lateral of the umbilicus, cleaned of blood and other visible non-adipose tissue, and rinsed with 0.9% sodium chloride. Adipose RNA-sequencing was performed as described for muscle RNA-sequencing [24].
RNA-Seq data processing
TwinsUK and METSIM
RNA-Seq reads were aligned to the hg19 reference genome using STAR [27] version 2.4.0.1 (TwinsUK) and 2.4.2a (METSIM), and quality control conducted as previously described [23, 28]. Gene-level counts were calculated using the quan function from QTLtools [29] and Gencode version 19 [30].
FUSION
RNA-Seq reads were aligned to the hg19 reference genome using procedures as described previously [24]. Six outlier samples were excluded based on low read coverage, and five outliers based on gene diversity. The allelic RNA-Seq read count distribution was compared to single nucleotide polymorphism (SNP) data from genome-wide association study (GWAS) panels available for the same subjects using verifyBamID [31]; two contaminated samples were identified and removed, as well as one pair of sample swaps corrected. Reported sex of the remaining samples was verified by matching the sex inferred using expression of XIST - encoded on the X chromosome and expressed only in cells with a minimum of two X chromosomes—and the mean Y chromosome gene expression. Linear regression of gene expression as a function of age, sex, batch, and RNA integrity number (RIN)—a measure of RNA quality—was performed, in addition to performing principal component analysis (PCA) on the gene expression residuals. The minimum number of PCs were selected to explain 20% of the variance in gene expression. Finally, one non-Finnish subject and one from each of two first-degree relative pairs were removed. A total of 280 samples were retained in the analysis.
Gene inclusion and normalization
For all three studies, RNA-Seq gene expression data were filtered to retain only genes with 5 or more counts per million (CPMs) in greater than or equal to 25% of individuals in each study. Trimmed mean of M-values between-sample normalisation was applied to gene counts [32], and TMM-normalised gene CPMs were then inverse-normalised prior to all downstream analyses (inverse-normalized ACE2).
Estimation of adipose tissue cell type proportions
As adipose tissue comprises adipocytes, immune cells, and vascular endothelial cells, all of which may be present in differing proportions between samples, it is critical to account for between-sample differences in the composition of cell types within biopsies. Adipose tissue cell type proportions were estimated from RNA-Seq data in TwinsUK using CIBERSORT [33], as previously described [34]. Estimated cell types included in our analysis were adipocytes, microvascular endothelial cells (MVEC) and macrophages.
To estimate the proportions of adipose and blood cell types in the adipose biopsies from the FUSION and METSIM studies, a reference transcriptome was created using RNA-Seq of whole blood (GEO accession GSE67488 [35]), and of four types of purified adipose cells (adipocytes, macrophages, CD4 + T cells, and microvascular endothelial cells) described in Glastonbury et al. [34]. For the FUSION study, reference transcriptome reads were aligned to the hg19 reference genome using the same read mapping and quality control procedure as used for the FUSION adipose RNA-Seq data [24]. Blood/cell-type proportions for each FUSION adipose sample were estimated using the unmix function from DESeq2 v1.18.1 [36]. A gene expression signature matrix was constructed and blood/cell-type proportions were estimated for each METSIM adipose sample using CIBERSORT [33].
Phenotypic association analyses
For quantitative phenotypic trait association analyses, in TwinsUK we excluded a subset of participants who had not fasted on the day of biopsy and for whom biochemical measurements were not matched to the day of biopsy, or for whom fasting and time of visit covariates used in our analyses were not available (n = 161). Participants with T2D were excluded from all analyses in all studies except those modelling association of diabetes status with ACE2 expression.
We assessed association between ACE2 gene expression levels (Gencode ID ENSG00000130234.6) and phenotypic traits including age, BMI, serum fasting insulin, fasting plasma glucose, serum HDL and LDL cholesterol, total triglycerides, systolic and diastolic blood pressure, obesity, T2D status (TwinsUK only) and sex (FUSION only). For obesity status, cases (TwinsUK: n = 119; METSIM: n = 67; FUSION: n = 68) were defined as participants with BMI ≥30 kg/m2, while controls (TwinsUK: n = 259; METSIM: n = 140; FUSION: n = 79) were defined as participants with BMI < 25 kg/m2. In TwinsUK, established T2D case status (n = 31) was defined through longitudinal patient self-reports, and longitudinal records of elevated fasting plasma glucose (≥7 mmol/l) and serum fasting insulin levels. In this study, individuals were defined as T2D controls (n = 567) if they had no longitudinal records of elevated fasting plasma glucose nor serum fasting insulin, as well as no self-reports of diabetes. Individuals with insulin resistance, defined as participants with longitudinally-elevated serum fasting insulin levels (>60 pmol/l) with fasting plasma glucose levels consistently below 6.0 mmol/l (n = 130) were excluded from T2D association analyses. In FUSION, study participants underwent a 2-h, four-point oral glucose tolerance test (OGTT) after a 12-h overnight fast, and T2D, impaired fasting glucose (IFG), impaired glucose tolerance (IGT) and normal glucose tolerance (NGT) status was assigned per WHO criteria [37]. We also assessed association of adipose tissue ACE2 expression with use of ACE inhibitors or angiotensin II receptor blockers, as well as adipose tissue estimated cell type proportions, specifically each of microvascular endothelial cell, macrophage and adipocyte proportions, estimated as described above.
We performed rank-based inverse normal transformation of all quantitative phenotypic traits. Association between ACE2 gene expression levels and each phenotypic trait was assessed by fitting linear mixed effects models using the lmer function from the lme4 package [38] (TwinsUK) or linear regression models using the lm function (METSIM and FUSION) in R [39] version 3.5.1 (TwinsUK and METSIM) or version 3.6.3 (FUSION). Inverse normalised ACE2 expression was treated as a continuous dependent variable, and each phenotypic trait in turn was treated as an independent fixed effect.
Covariates included in the TwinsUK study were: age, BMI (for all phenotypes except BMI and obesity), number of hours fasted, and the RNA-Seq technical covariates sample median transcript integrity number (TIN—a measure of sample quality) [40], median insert size, and mean GC content included as fixed effects. Random effects included were date of RNA sequencing, morning versus afternoon visit time, RNA extraction batch, RNA-Seq primer index, family and zygosity. Family and zygosity are both random effects that describe the family a twin belongs to, and their clonality (MZ/DZ status). Each family identifier is specific to a twin pair, and each zygosity label is identical for each pair of MZ twins, while individual DZ co-twins are each given a unique zygosity label. In TwinsUK, the full model was then compared to a null model where the phenotypic trait of interest was omitted, using a 1 degree of freedom ANOVA.
Covariates included in the METSIM study were: age, BMI (for all phenotypes except BMI and obesity), blood cell type proportion, and RNA-Seq technical covariates read deletion size, mean read insertion size, TIN, and sequencing batch.
Covariates included in the FUSION study were: age, sex, BMI (for all phenotypes except BMI and obesity), collection site, blood cell type proportion, four genetic PCs, and RNA-Seq technical covariates mean GC content, median read insertion size, RNA integrity number (RIN), TIN, and sequencing batch included as fixed effects. In the FUSION study, which included both males and females, analyses were also performed in males and females separately, excluding the sex covariate in the models.
Sensitivity analyses assessing the impact of adjusting for body fat distribution as opposed to BMI were conducted in all three studies. In TwinsUK, dual-energy X-ray absorptiometry (DEXA) scans were used to quantify fat volume in android and gynoid regions following standard manufacturer’s recommendations (DXA; Hologic QDR 4500 plus), from which android/gynoid ratio was calculated. In each of the METSIM and FUSION studies, waist/hip ratio (WHR) was used as a measure of body fat distribution. Sensitivity analyses were run by including either android/gynoid ratio (TwinsUK) or WHR (METSIM and FUSION) as a covariate instead of BMI in the models above. Analyses in the FUSION study were conducted in males and females separately.
Sensitivity analyses were conducted for all three studies, including estimated MVEC cell type proportion, then both estimated MVEC and macrophage cell type proportions, as an additional covariate(s) in the models above.
Meta-analysis of phenotypic associations with adipose tissue ACE2 expression
Given the demographic differences between our study samples, we selected a meta-analysis approach that permited the true effect to vary between studies. We thus conducted a random-effects inverse-variance weighted meta-analysis of up to 1,237 participants, combining ACE2 gene expression-phenotype association results from the TwinsUK, METSIM and FUSION studies (See Supplementary Tables 1–6 for trait-specific numbers of participants). Meta-analysis was conducted for 13 out of 15 traits using the rma function (restricted maximum likelihood method) from the metafor package [41] in R version 3.5.1 [39]. Association of T2D with ACE2 was assessed in TwinsUK only, as it was the only study with participants with established T2D at time of biopsy. Association of ACE2 expression with sex was assessed in FUSION only, given that the TwinsUK and METSIM studies included only females or males, respectively, to avoid biasing of estimates that may result from confounding by study-specific technical differences.
In order to take into account the number of tests conducted, a multiple testing correction threshold of P < 3.33 × 10−3, calculated using a Bonferroni correction for 15 traits (0.05/15), was used to assess significance of phenotypic associations with ACE2 expression.
Detection of COVID-19 antibodies in TwinsUK participants
Enzyme linked immunosorbent assay (ELISA)
Presence of immunoglobulin G (IgG) to SARS-CoV-2 nucleoprotein (N) and spike (S) in serum from TwinsUK participants was measured using ELISA as previously described [42, 43]. Serum samples were diluted to 1:50. Optical density (OD) values >4-fold above assay background was used as a cut-off for seropositivity [42].
Lateral flow assays
Fortress COVID-19 Total Ab Device (Fortress Diagnostics Ltd) was used for qualitative detection of total antibodies against SARS-CoV-2, employing a chromatographic lateral flow device within a cassette format. In total, 10 μl of serum was added to the specimen window immediately followed by two drops of diluent buffer to the buffer window. Each cassette was left for exactly 10 min before being read by a trained person. Seropositivity was indicated by the presence of a visible red line in the Test Zone and results were classified as either IgG positive only, IgM positive only, IgG & IgM positive, negative or invalid. Prior to and during the cassette readings the personnel responsible for readings undertook a verification process looking at inter-rater and intra-rater reading variability.
Identification of SARS-CoV-2 seropositive individuals in TwinsUK
The presence of SARS-CoV-2 antibodies was assessed in 459 TwinsUK subjects for whom adipose tissue gene expression data were available, using either ELISA or lateral flow SARS-CoV-2 antibody tests as described. Individuals were classified as SARS-CoV-2 seropositive if they received a positive antibody test result from either method.
Longitudinal COVID-19 symptom reporting in TwinsUK
Experience of potential COVID-19 symptoms during the period between February-July 2020 was collected from TwinsUK participants using detailed postal health questionnaires. The symptoms which the participants were asked to report on are shown in Supplementary Table 7.
Association of adipose tissue ACE2 expression with COVID-19 symptom presentation in SARS-CoV-2 seropositive participants from TwinsUK
We assessed association between adipose tissue ACE2 gene expression levels and presentation of one or more core COVID-19 symptoms (fever, persistent cough, anosmia) and/or one or more of the more severe COVID-19 symptoms (shortness of breath, chest pain or chest tightness) in a sample of 32 unrelated subjects, without diabetes, from TwinsUK who were found to be seropositive for SARS-CoV-2 antibodies, and for whom COVID-19 symptom reports were available.
Association was assessed by fitting a logistic regression model using the glm function in R version 3.5.1. The presence or absence of core and/or severe COVID-19 symptoms was treated as a binary dependent variable; ACE2 expression, adjusted for RNA-Seq technical covariates (sample median transcript integrity number (TIN), median insert size, mean GC content, date of sequencing and primer index) as well as age and BMI at time of biopsy, was treated as an independent fixed effect, with BMI and age at antibody assessment included as covariates in the model.
Correlation of phenotypic traits
The correlations between phenotypic traits were assessed in TwinsUK, which was the largest study included in our analyses. Phenotypic trait correlations were assessed using a Spearman correlation in unrelated participants from TwinsUK (n = 441), in R version 3.5.1 [39].
Results
We quantified adipose ACE2 expression levels in bulk RNA-Seq data from TwinsUK [22, 44, 45] (765 female mono- and dizygotic twins), 426 males from the Metabolic Syndrome in Men (METSIM) study [23], and 149 Finnish males and 131 Finnish females from the Finland-United States Investigation of NIDDM Genetics (FUSION) Tissue Biopsy Study [24]. The cohorts differ in country of origin, male:female composition, age distribution, and underlying ascertainment criteria (Table 1). Study participants ranged in age from 35 to 85 years, with variation between studies (Table 1 and Supplementary Fig. 2). While TwinsUK and METSIM are representative of the UK [22] and Kuopio Finland [23] populations respectively, the FUSION study design was enriched for individuals with abnormal glycaemic indices at time of biopsy.
We hypothesised that the true effect sizes of ACE2 gene expression-phenotypic associations may vary between cohorts that differ in their demographic characteristics, and due to technical and methodological differences between the studies. We therefore assessed association of ACE2 gene expression with demographic and phenotypic traits in each of the TwinsUK, METSIM and FUSION studies, adjusting for age and BMI, and then combined the association results from all three studies by conducting a random-effect inverse-variance-weighted meta-analysis. Results of meta-analysis of the three studies showed consistency in association signals across cohorts for some traits, but heterogeneity for others.
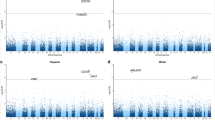
Adipose tissue ACE2 expression was relatively low across all three cohorts (Table 1, Supplementary Figs. 3 and 4). In our meta-analysis, lower adipose tissue ACE2 expression levels were associated with higher serum fasting insulin (β [95% CI] = −0.12 [−0.18, −0.05]; P = 5.32 × 10−4), and higher body mass index (BMI) (β [95% CI] = −0.10 [−0.16, −0.04]; P = 3.94 × 10−4) (Fig. 1 and Supplementary Table 1). Consistent with the association observed for BMI, participants with obesity had lower adipose tissue ACE2 expression compared to normal-weight controls (β [95% CI] = −0.34[−0.50, −0.17]; P = 4.81 × 10−5) (Fig. 1 and Supplementary Table 1).
Squares with error bars represent the standardised β coefficients and their 95% confidence intervals for association of ACE2 expression levels with each trait (derived from linear/linear mixed effects regression models), with meta-analysis effect sizes and 95% confidence intervals (random-effects meta-analysis) shown as black diamonds. N represents the sample size for each analysis. a serum fasting insulin, (b) fasting plasma glucose, (c) BMI (d) serum HDL cholesterol, (e) serum LDL cholesterol, (f) serum total triglycerides, (g) systolic blood pressure, (h) diastolic blood pressure, (i) age, (j) obesity status. Association of ACE2 expression with (k) type 2 diabetes status in TwinsUK and (l) sex in the FUSION study. Boxplots (k, l) display the median and inter-quartile range (IQR), with whiskers corresponding to ±1.5*IQR.
TwinsUK includes a subset of individuals diagnosed with T2D prior to biopsy. Within TwinsUK, T2D status was associated with lower adipose tissue ACE2 expression compared to normoglycaemic controls (β[95% CI] = −0.84[−1.21, −0.47]; P = 9.14 × 10−6). No association was observed with fasting glucose in our study (P > 0.05) (Fig. 1 and Supplementary Table 1).
Lower ACE2 expression was strongly associated with lower serum HDL cholesterol (β [95% CI] = 0.16 [0.10, 0.21]; P = 1.92 × 10−7) (Fig. 1 and Supplementary Table 1). Association of total serum triglycerides with adipose tissue ACE2 expression showed heterogeneity between studies (heterogeneity indices: I2 = 73.24; Q-test P = 0.01). Results of meta-analysis of all three studies showed only nominally significant association between lower ACE2 expression and higher triglyceride levels (β [95% CI] = −0.13[−0.25, 0.00]; P = 4.15 × 10−2), but this association was not significant after multiple testing correction (MTC threshold P < 3.33 × 10−3). In TwinsUK, lower ACE2 expression in adipose tissue was strongly associated with higher total triglyceride levels (β [95% CI] = −0.23 [−0.31, −0.14]; P = 1.63 × 10−7). Adipose tissue ACE2 expression was not associated with serum LDL cholesterol, nor with systolic or diastolic blood pressure (P > 0.05) (Fig. 1 and Supplementary Table 1).
Severity of COVID-19 is associated with increasing age [2,3,4, 46]. Upon meta-analysis, higher adipose ACE2 expression levels were nominally associated with older age (β [95% CI] = 0.15 [0.05, 0.26]; P = 5.23 × 10−3), but not significant after MTC (Fig. 1 and Supplementary Table 1). However, we observed heterogeneity in age effects between studies (Heterogeneity I2 = 71.38; Cochrane’s Q-test P = 0.02) potentially driven by differences in the age distribution of the studies. In the study, with the largest age range, TwinsUK (38–85 years), higher adipose tissue ACE2 expression was strongly associated with older age (β [95% CI] = 0.25[0.17, 0.32]; P = 6.31 × 10−9). Higher ACE2 expression in adipose tissue from older individuals was consistent with previous reports of age-dependent ACE2 expression in nasal epithelium [47]. However, in skin samples from the same TwinsUK participant, lower ACE2 expression was associated with increased age (β [95% CI] = −0.23[−0.31, −0.14]; P = 4.77 × 10−7), supporting heterogeneity of effects between tissues [11] (Supplementary Fig. 5). We also noted that adipose tissue gene expression levels of ACE2 were not associated with use of ACE inhibitors or angiotensin II receptor blockers (P > 0.05; Supplementary Table 2).
Located on the X chromosome, ACE2 has been shown to escape from X inactivation [48]. It is therefore of interest to consider whether ACE2 expression differences may contribute to the observed differences in COVID-19 severity and clinical outcomes. Association of ACE2 expression with sex was assessed only in the FUSION study, which included both males and females. Consistent with reports from GTEx [11], ACE2 expression was nominally higher in females in FUSION (β [95% CI] = 0.32 [0.02, 0.61]; P = 0.037), but this difference was not significant after multiple testing correction (Fig. 1 and Supplementary Table 1).
Given the well-documented differences in body fat distribution between males and females, we sought to conduct a sensitivity analysis to assess whether phenotypic associations may be confounded by these differences. We therefore adjusted for available measures of body fat distribution in each of the three studies in place of BMI, adjusting for DEXA-derived android/gynoid ratio in TwinsUK, and waist/hip ratio in each of METSIM and FUSION. Sensitivity analyses in the FUSION study were conducted in males and females separately. Phenotypic associations were robust to these adjustments, and were in some cases strengthened (Supplementary Table 3).
Adipose tissue consists of multiple cell types, gene expression of all of which is represented in bulk RNA-Seq data, and to our knowledge, it is still unclear which adipose cell types are responsible for expression of ACE2. We therefore assessed the relationship between ACE2 expression in subcutaneous adipose tissue and estimated adipose tissue cell type proportions. Consistent with differences in biopsy methods employed, significant heterogeneity in cell type composition was observed between the three studies (Fig. 2). Despite these differences, a higher proportion of microvascular endothelial cells (MVEC) was associated with higher ACE2 expression (β [95% CI] = 0.15 [0.07, 0.23]; P = 4.25 × 10−4), suggesting microvascular endothelial cell subpopulations contribute to ACE2 expression levels in adipose tissue (Fig. 2 and Supplementary Table 4). Conversely, lower ACE2 expression was associated with higher macrophage proportion (β [95% CI] = −0.18 [−0.26, −0.09]; P = 2.74 × 10−5) (Fig. 2 and Supplementary Table 4). Increased macrophage infiltration in adipose tissue has been associated with a pro-inflammatory state [49]. ACE2 expression was not associated with adipocyte proportion (P > 0.05; Fig. 2 and Supplementary Table 4).
Density plots of estimated cell type proportions of (a) microvascular endothelial cells (MVEC), (b) macrophages, (c) adipocytes in each of the TwinsUK, METSIM, and FUSION studies. Vertical dashed lines correspond to the study mean. Association of adipose tissue ACE2 expression levels with estimated (d) MVEC, (e) macrophage and (f) adipocyte proportions. Squares with error bars represent the standardised β coefficients and their 95% confidence intervals for association of ACE2 expression levels with each of the traits (derived from linear/linear mixed effects regression models), with the meta-analysis effect sizes and 95% confidence intervals (random-effects meta-analysis) shown as black diamonds. N represents the sample size for each analysis.
To establish whether phenotypic associations with ACE2 expression were simply reflecting differences in adipose tissue cell type composition [34], we conducted sensitivity analyses including the estimated proportion of MVECs as a covariate in our models. While strength of association was attenuated for some phenotypes, the associations were largely robust to adjustment for MVEC proportion (Supplementary Table 5), suggesting phenotypic associations were not driven solely by differences in the proportions of ACE2-expressing cell types. Furthermore, correlation patterns between phenotypic traits, including age and each of adipose tissue ACE2 expression and MVEC proportions, were notably different (Supplementary Fig. 6). In contrast, phenotypic associations were notably attenuated upon inclusion of both MVEC and macrophage proportions as covariates (Supplementary Table 6).
We next sought to investigate whether background adipose tissue ACE2 expression levels were associated with COVID-19 clinical presentation in individuals confirmed to have experienced SARS-CoV-2 infection. As part of the TwinsUK COVID-19 research initiative, we assessed SARS-CoV-2 antibody status and COVID-19 symptom reports in TwinsUK participants (Supplementary Table 7), of whom 459 overlapped our adipose tissue gene expression study sample. In unrelated TwinsUK participants seropositive for SARS-CoV-2 antibodies (N = 32), we assessed whether background ACE2 expression was associated with presentation of one or more core COVID-19 symptoms (fever, persistent cough, anosmia) and/or one or more of the more severe COVID-19 symptoms (shortness of breath, chest pain or chest tightness). Adipose tissue ACE2 expression was nominally lower in SARS-CoV-2 seropositive subjects who exhibited one or more classic or severe symptoms of COVID-19 compared to seropositive subjects who did not exhibit any of these symptoms (β [95% CI] = −1.31 [−2.53, −0.09]; P = 0.035; Fig. 3). Further studies in larger sample sizes will be required to confirm these findings.
Adipose tissue ACE2 expression residuals (adjusted for age, BMI and RNA-Seq technical covariates) are shown in SARS-CoV-2 seropositive TwinsUK participants who reported presenting one or more classic or severe COVID-19 symptoms compared to seropositive individuals who did not display any of the classic or severe COVID-19 symptoms. Boxplots display the median and inter-quartile range (IQR), with whiskers corresponding to ±1.5*IQR. Effect size and P value were calculated using a logistic regression model adjusting for current age and BMI.
Finally, despite a heritability of 0.32 [95% CI = 0.14–0.50] in TwinsUK, cis- and trans-eQTL meta-analyses of all three studies did not reveal any significant eQTLs associated with adipose tissue ACE2 expression (Supplementary Information and Supplementary Fig. 7). Similarly, no proximal methylation probes were associated with genetic variants in cis or trans in the TwinsUK sample (Supplementary Information and Supplementary Fig. 8).
Discussion
Taken together, our results suggest that lower adipose tissue expression of ACE2 is associated with multiple adverse cardio-metabolic health indices, all of which are risk factors for severe COVID-19 [2, 4, 5]. Lower adipose tissue ACE2 expression was associated with diabetes and obesity status, increased serum fasting insulin and triglyceride levels, BMI, and with increased macrophage infiltration in adipose tissue, a marker of inflammation [49]. ACE2 expression was positively correlated with HDL levels, age and adipose tissue MVEC proportion. Exploratory analyses in a small sample of SARS-CoV-2 seropositive individuals from TwinsUK suggested association of adipose tissue ACE2 expression levels with symptom presentation. While the limited sample size warrants cautious interpretation, these findings highlight the importance of assessing the contribution of adipose tissue ACE2 expression to COVID-19 severity in larger sample sizes. As previously noted for ACE2 associations reported in other tissues [50], and given its association with adipose tissue cell type proportions, it remains unclear to what extent these associations are mediated through differences in underlying cell type composition. While sensitivity analyses adjusting for MVEC cell type proportion estimates suggested that the ACE2 expression-trait associations were robust, estimation of cell type composition from bulk RNA-Seq data in non-blood tissues remains in its infancy. Adipose tissue single-cell RNA sequencing (scRNA-Seq) datasets will be critical to address this question fully. Furthermore, as shown by the opposing directions of effect in skin versus adipose tissue for association with age in the TwinsUK sample, ACE2 expression-trait associations may differ between tissues, supporting the need for caution in their interpretation and extrapolation. The contrast between our observation that adipose tissue ACE2 levels were positively correlated with age - a strong risk factor for severe COVID-19 - yet negatively correlated with COVID-19 cardio-metabolic risk factors may reflect the multi-systemic nature of COVID-19. There are likely a number of mechanisms that underlie increased risk of severe COVID-19, and these are not necessarily shared by all its demographic and cardio-metabolic risk factors. While adipose tissue may be a key driver of increased susceptibility to severe COVID-19 for some risk factors such as cardio-metabolic risk factors, other tissues and mechanisms may be more important for other traits such as age.
Despite evidence of heritability of adipose tissue ACE2 expression levels and our meta-analysis sample size, we did not yet identify cis- or trans-eQTLs associated with adipose ACE2 expression. Increased sample sizes may be needed to detect genetic factors influencing expression of ACE2 in this important tissue.
Increasing evidence suggests that COVID-19 severity may be modulated by widespread microvascular damage and increased predisposition to thrombotic events [51,52,53], in which ACE2, and the RAS system more widely, may play a critical role. Adipose tissue is an active endocrine organ which secretes hormones, chemokines and cytokines that contribute to regulation of inflammatory and immune responses [54, 55]. Given the association of ACE2 expression with cardio-metabolic risk factors for severe COVID-19, it is intriguing to consider whether low levels of adipose ACE2 expression may contribute to COVID-19 severity in at-risk individuals. SARS-CoV infection reduces ACE2 levels post-infection [56,57,58,59]. Our observation that individuals suffering from co-morbidities recognised to be associated with severe COVID-19 may have lower pre-infection ACE2 levels in some tissues may support a theory that pre-existing low ACE2 levels in at risk-individuals are then reduced further upon SARS-CoV-2 infection, contributing to the dysregulation of the RAS characteristic of severe COVID-19 [60]. Further studies will be needed to clarify the contribution of adipose tissue ACE2 expression levels to COVID-19 severity and beyond.
Data availability
TwinsUK RNA-Seq data are deposited in the European Genome-phenome Archive (EGA) under accession EGAS00001000805. METSIM RNASeq data have been deposited in the Gene Expression Omnibus (GEO) under accession GSE135134. The dbGaP accession number for the FUSION Tissue Biopsy Study is phs001048.v2.p1.
References
Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis. 2020;20:533–4.
Docherty AB, Harrison EM, Green CA, Hardwick HE, Pius R, Norman L, et al. Features of 16,749 hospitalised UK patients with COVID-19 using the ISARIC WHO Clinical Characterisation Protocol. medRxiv. 2020;10:2020.04.23.20076042.
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395:507–13.
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506.
Intensive care national audit & research centre. ICNARC report on COVID-19 in critical care - 04 April 2020. 2020:1–16.
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–74.
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271–280.e8.
Ziegler CGK, Allon SJ, Nyquist SK, Mbano IM, Miao VN, Tzouanas CN, et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell. 2020;181:1016–1035.e19.
Seow JJW, Pai R, Mishra A, Shepherdson E, Lim TKH, Goh BKP, et al. scRNA-seq reveals ACE2 and TMPRSS2 expression in TROP2+ liver progenitor cells: implications in COVID-19 associated liver dysfunction. Front Med. 2021;8:603374.
Fan C, Lu W, Li K, Ding Y, Wang J. ACE2 expression in kidney and testis may cause kidney and testis infection in COVID-19 patients. Front Med. 2021;7:563893.
He Y Tissue specific expression and genetic regulation of SARS-CoV-2 receptors ACE2 and TMPRSS2. https://medium.com/@yuanhe716/tissue-specific-expression-and-genetic-regulation-of-sars-cov-2-receptors-ace2-and-tmprss2-d75d76bf45fb (accessed 13 Jun 2020).
Schmieder RE, Hilgers KF, Schlaich MP, Schmidt BM. Renin-angiotensin system and cardiovascular risk. Lancet. 2007;369:1208–19.
Tikellis C, Thomas MC. Angiotensin-converting enzyme 2 (ACE2) is a key modulator of the renin angiotensin system in health and disease. Int. J. Pept. 2012;2012:256294.
Crackower MA, Sarao R, Oliveira-dos-Santos AJ, Da Costa J, Zhang L. Angiotensin-converting enzyme 2 is an essential regulator of heart function. Nature. 2002;417:822–8.
Arner P, Spalding KL. Fat cell turnover in humans. Biochem Biophys Res Commun. 2010;396:101–4.
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Malavazos AE, Corsi Romanelli MM, Bandera F, Iacobellis G. Targeting the adipose tissue in COVID-19. Obesity. 2020;28:1178–9.
Ryan PMD, Caplice NM. Is adipose tissue a reservoir for viral spread, immune activation, and cytokine amplification in Coronavirus Disease 2019? Obesity. 2020;28:1191–4.
Ryan DH, Ravussin E, Heymsfield S. COVID 19 and the patient with obesity – the editors speak out. Obesity. 2020;28:847.
Goossens GH, Dicker D, Farpour-Lambert NJ, Fruhbeck G, Mullerova D, Woodward E, et al. Obesity and COVID-19: a perspective from the European association for the study of obesity on immunological perturbations, therapeutic challenges, and opportunities in obesity. Obes Facts. 2020;13:439–52.
Akoumianakis I, Filippatos T. The renin–angiotensin–aldosterone system as a link between obesity and coronavirus disease 2019 severity. Obes Rev. 2020;21:e13077.
Moayyeri A, Hammond CJ, Hart DJ, Spector TD. The UK adult twin registry (twinsUK resource). Twin Res Hum Genet. 2013;16:144–9.
Raulerson CK, Ko A, Kidd JC, Currin KW, Brotman SM, Cannon ME, et al. Adipose tissue gene expression associations reveal hundreds of candidate genes for cardiometabolic traits. Am J Hum Genet. 2019;105:773–87.
Scott LJ, Erdos MR, Huyghe JR, Welch RP, Beck AT, Wolford BN, et al. The genetic regulatory signature of type 2 diabetes in human skeletal muscle. Nat Commun. 2016;7:11764.
Buil A, Brown AA, Lappalainen T, Viñuela A, Davies MN, Zheng H-F, et al. Gene-gene and gene-environment interactions detected by transcriptome sequence analysis in twins. Nat Genet. 2015;47:88–91.
Civelek M, Wu Y, Pan C, Raulerson CK, Ko A, He A, et al. Genetic regulation of adipose gene expression and cardio-metabolic traits. Am J Hum Genet. 2017;100:428–43.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Glastonbury CAA, Viñuela A, Buil A, Halldorsson GHH, Thorleifsson G, Helgason H, et al. Adiposity-dependent regulatory effects on multi-tissue transcriptomes. Am J Hum Genet. 2016;99:567–79.
Delaneau O, Ongen H, Brown AA, Fort A, Panousis NI, Dermitzakis ET. A complete tool set for molecular QTL discovery and analysis. Nat Commun. 2017;8:15452.
Frankish A, Diekhans M, Ferreira A-M, Johnson R, Jungreis I, Loveland J, et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 2019;47:767.
Jun G, Flickinger M, Hetrick KN, Romm JM, Doheny KF, Abecasis GR, et al. Detecting and estimating contamination of human DNA samples in sequencing and array-based genotype data. Am J Hum Genet. 2012;91:839–48.
Chen Y, Mccarthy D, Ritchie M, Robinson M, Smyth GK edgeR: differential expression analysis of digital gene expression data User’s Guide. http://bioconductor.org/packages/release/bioc/vignettes/edgeR/inst/doc/edgeRUsersGuide.pdf (accessed 3 May 2017).
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7.
Glastonbury CA, Couto Alves A, El-Sayed Moustafa JS, Small KS. Cell-type heterogeneity in adipose tissue is associated with complex traits and reveals disease-relevant cell-specific eQTLs. Am J Hum Genet. 2019;104:1013–24.
Tabassum R, Sivadas A, Agrawal V, Tian H, Arafat D, Gibson G. Omic personality: Implications of stable transcript and methylation profiles for personalized medicine. Genome Med. 2015;7:88–88.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
WHO. Definition and Diagnosis of Diabetes Mellitus and Intermediate Hyperglycemia: report of a WHO/IDF consultation. 2006.
Bates D, Mächler M, Bolker B, Walker S. Fitting linear mixed-effects models using lme4. J Stat Softw. 2015;67:51.
R Development Core Team. R: A Language and Environment for Statistical Computing. R Found Stat Comput Vienna Austria 2013; 0.
Wang L, Wang S, Li W, Li Y, Eckel-Passow JE, Dasari S, et al. RSeQC: quality control of RNA-seq experiments. Bioinformatics. 2012;28:2184–5.
Viechtbauer W. Conducting meta-analyses in R with the metafor. J Stat Softw. 2010;36:1–48.
Pickering S, Betancor G, Galão RP, Merrick B, Signell AW, Wilson HD, et al. Comparative assessment of multiple COVID-19 serological technologies supports continued evaluation of point-of-care lateral flow assays in hospital and community healthcare settings. PLoS Pathog. 2020;16:e1008817.
Seow J, Graham C, Merrick B, Acors S, Pickering S, Steel KJA, et al. Longitudinal observation and decline of neutralizing antibody responses in the three months following SARS-CoV-2 infection in humans. Nat Microbiol. 2020;5:1598–607.
Grundberg E, Small KS, Hedman ÅK, Nica AC, Buil A, Keildson S, et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat Genet. 2012;44:1084–9.
Buil A, Brown AA, Lappalainen T, Viñuela A, Davies MN, Zheng H-F, et al. Gene-gene and gene-environment interactions detected by transcriptome sequence analysis in twins. Nat Genet. 2015;47:88–91.
Intensive care national audit & research centre. ICNARC report on COVID-19 in critical care - 04 April 2020. 2020.
Bunyavanich S, Do A, Vicencio A Nasal Gene Expression of Angiotensin-Converting Enzyme 2 in Children and Adults. JAMA - J. Am. Med. Assoc. 2020. https://doi.org/10.1001/jama.2020.8707.
Tukiainen T, Villani AC, Yen A, Rivas MA, Marshall JL, Satija R, et al. Landscape of X chromosome inactivation across human tissues. Nature. 2017;550:244–8.
Weisberg SP, McCann D, Desai M, Rosenbaum M, Leibel RL, Ferrante AW. Obesity is associated with macrophage accumulation in adipose tissue. J Clin Invest. 2003;112:1796–808.
Patel AB, Verma A. Nasal ACE2 Levels and COVID-19 in Children. J Am Med Assoc. 2020;323:2386–7.
Han H, Yang L, Liu R, Liu F, Wu KL, Li J, et al. Prominent changes in blood coagulation of patients with SARS-CoV-2 infection. Clin Chem Lab Med. 2020;58:1116–20.
Lodigiani C, Iapichino G, Carenzo L, Cecconi M, Ferrazzi P, Sebastian T, et al. Venous and arterial thromboembolic complications in COVID-19 patients admitted to an academic hospital in Milan, Italy. Thromb Res. 2020;191:9–14.
Klok FA, Kruip MJHA, van der Meer NJM, Arbous MS, Gommers DAMPJ, Kant KM, et al. Incidence of thrombotic complications in critically ill ICU patients with COVID-19. Thromb Res. 2020;191:145–7.
Scheja L, Heeren J. The endocrine function of adipose tissues in health and cardiometabolic disease. Nat Rev Endocrinol. 2019;15:507–24.
Makki K, Froguel P, Wolowczuk I. Adipose tissue in obesity-related inflammation and insulin resistance: cells, cytokines, and chemokines. ISRN Inflamm. 2013;2013:1–12.
Kuba K, Imai Y, Rao S, Gao H, Guo F, Guan B, et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat Med. 2005;11:875–9.
Haga S, Yamamoto N, Nakai-Murakami C, Osawa Y, Tokunaga K, Sata T, et al. Modulation of TNF-α-converting enzyme by the spike protein of SARS-CoV and ACE2 induces TNF-α production and facilitates viral entry. Proc Natl Acad Sci USA. 2008;105:7809–14.
Imai Y, Kuba K, Rao S, Huan Y, Guo F, Guan B, et al. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature. 2005;436:112–6.
Zheng YY, Ma YT, Zhang JY, Xie X. COVID-19 and the cardiovascular system. Nat Rev Cardiol. 2020;17:259–60.
Alghatrif M, Cingolani O, Lakatta EG. The dilemma of coronavirus disease 2019, aging, and cardiovascular disease: insights from cardiovascular aging science. JAMA Cardiol. 2020;5:747–8.
Acknowledgements
KSS acknowledges funding from the Medical Research Council (MR/M004422/1 and MR/R023131/1). TwinsUK is funded by the Wellcome Trust, Medical Research Council, Versus Arthritis, European Union Horizon 2020, Chronic Disease Research Foundation (CDRF), Zoe Ltd and the National Institute for Health and Care Research (NIHCR) Clinical Research Network (CRN) and Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust in partnership with King’s College London. The FUSION and METSIM studies were supported by US National Institutes of Health (NIH) grants U01DK062370, 1-ZIA-HG000024, R01DK093757, R01DK072193, P01HL28481, and Academy of Finland Grants 271961, 272741 and 258753. SMB was supported in part by a grant from the National Institute of General Medical Sciences under award 5T32 GM007092 and acknowledges the Global Partnership Award and UNC Global for travel funds. SV is supported by the National Council of Science and Technology of Mexico (CONACYT). SARS-CoV-2 serological analyses were supported by a COVID urgent response grant from the Chronic Disease Research Foundation (CDRF, CJS) and a King’s Together Rapid COVID-19 Call award (MHM, KJD, CJS). Questionnaire material for the assessment of COVID-19 symptoms was developed as part of the Wellcome Longitudinal Population Study COVID-19 Steering Group and Secretariat (supported by Wellcome funding 221574/Z/20/Z). Data on TwinsUK participants are available to bona fide researchers under managed access due to governance and ethical constraints. Raw data should be requested via our website (https://twinsuk.ac.uk/resources-for-researchers/access-our-data/) and requests are reviewed by the TwinsUK Resource Executive Committee (TREC) regularly. The authors acknowledge use of the research computing facility at King’s College London, Rosalind (https://rosalind.kcl.ac.uk), which is delivered in partnership with the National Institute for Health Research (NIHR) Biomedical Research Centres at South London & Maudsley and Guy’s & St. Thomas’ NHS Foundation Trusts, and part-funded by capital equipment grants from the Maudsley Charity (award 980) and Guy’s & St. Thomas’ Charity (TR130505). The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHCR, King’s College London, or the Department of Health and Social Care. The authors thank Dr. Jörg Saßmannshausen and Dr Lukasz Zalweski from the Rosalind support team for their computational support.
Author information
Authors and Affiliations
Contributions
JSE-SM and KSS conceived and designed the study. JSE-SM conducted statistical analyses of the TwinsUK study data and meta-analyses. AUJ and LG conducted statistical analyses of the FUSION study data. SMB conducted statistical analyses of the METSIM study data. JSE-SM, AUJ, LG, SMB, LJS, KLM and KSS interpreted the data. SV performed analysis of the TwinsUK methylation data, supervised by JTB. MF advised on statistical analyses. ALR, AZ, LB, MRE, NN, HMS, RW, TY, TL, SP, JT, FSC, PP, MB, HAK, ML, JS, CG, IH, SA, NK, KJD, MHM, SW, ELD, and CJS participated in sample collection/data generation/data processing. JSE-SM, AUJ, LG, SMB, ALR, LJS, KLM and KSS wrote the manuscript. All co-authors provided critical feedback and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics declaration
The TwinsUK project was approved by the ethics committee at St Thomas’ Hospital London, where all the biopsies were carried out. Volunteers gave informed consent and signed an approved consent form prior to the biopsy procedure. Volunteers were supplied with an appropriate detailed information sheet regarding the research project and biopsy procedure by post prior to attending for the biopsy. The FUSION study was approved by the coordinating ethics committee of the Hospital District of Helsinki and Uusimaa, and written informed consent was obtained from all participants. The METSIM study was approved by the Ethics Committee of the University of Eastern Finland and Kuopio University Hospital in Kuopio, Finland, and written informed consent was obtained from all participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El-Sayed Moustafa, J.S., Jackson, A.U., Brotman, S.M. et al. ACE2 expression in adipose tissue is associated with cardio-metabolic risk factors and cell type composition—implications for COVID-19. Int J Obes 46, 1478–1486 (2022). https://doi.org/10.1038/s41366-022-01136-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-022-01136-w
This article is cited by
-
The interplay of aging, adipose tissue, and COVID-19: a potent alliance with implications for health
GeroScience (2024)
-
Comparison of different prognostic scores in estimating short- and long-term mortality in COVID-19 patients above 60 years old in a university hospital in Belgium
European Geriatric Medicine (2023)