Abstract
Translation is mediated by precisely orchestrated sequential interactions among translation initiation components, mRNA, and ribosomes. Biochemical, structural, and genetic techniques have revealed the fundamental mechanism that determines what occurs and when, where and in what order. Most mRNAs are circularized via the eIF4E–eIF4G–PABP interaction, which stabilizes mRNAs and enhances translation by recycling ribosomes. However, studies using single-molecule fluorescence imaging have allowed for the visualization of complex data that opposes the traditional “functional circularization” theory. Here, we briefly introduce single-molecule techniques applied to studies on mRNA circularization and describe the results of in vitro and live-cell imaging. Finally, we discuss relevant insights and questions gained from single-molecule research related to translation.
Similar content being viewed by others
Introduction
In protein synthesis, translation initiation is a rate-limiting step because of cascading mRNA and protein (mRNP) formation. Extensive research has revealed that this process is regulated by more than 12 translation initiation factors1,2. During the process, mRNA undergoes functional circularization (or becomes a “closed loop”) through protein–mRNA and protein–protein interactions to positively regulate protein synthesis3,4; that is, functional circularization5 facilitates the recycling of posttermination ribosomes for reinitiation6,7 and shields the mRNA from decay factors8,9. Since all eukaryotic mRNAs, except histone mRNA, have a unique m7G cap structure at the 5ʹ-end and a poly(A) tail at the 3ʹ-end, circularization mostly begins with the recognition of binding proteins to both ends, namely, 5ʹ-cap-eIF4E2,10 and 3ʹ-poly(A) tail-PABP11, respectively. Some mRNAs utilize an internal ribosome entry site (IRES)12,13, N6-methyladenosine (m6A)14,15, or elements related to viruses16,17,18 instead of a cap and/or poly(A) tail to interact with their binding partners. Each binding protein can communicate with other binding proteins at the other end with the help of mediators such as eIF4G19,20 and/or eIF315. Moreover, since circularization is based on stochastic interactions with biomolecules, it may not result in a fixed loop; instead, the loop can change dynamically, suggesting that kinetic/dynamic studies on mRNP components during translation contribute to an integrated understanding of translation21. However, because of technical difficulties, which involve the fluorescent labeling of translation factors, single-particle tracking (SPT) in high-background signals, and spatiotemporal resolution limits, only direct structural evidence has been obtained through atomic force microscopy22. Thus, mRNA circularization remains poorly understood.
Excitingly, single-molecule fluorescence imaging (smFI) has been employed to examine mRNA circularization in live cells, which are composed of numerous transitory interactions23,24. Conformational changes in mRNA are visualized by SPT with high spatiotemporal resolution. It has enabled us to determine the relationship between translation and circularization in cells. Meanwhile, in vitro sm studies on initiation factors have revealed their binding kinetics or conformational change in protein synthesis, thus contributing significantly to our understanding of the molecular processes involved in circularization25,26,27,28,29. In this review, we briefly introduce how smFI techniques have been applied to investigate mRNA circularization and discuss the current knowledge of the relationship between translation and mRNA circularization. Finally, to provide perspective, we address some questions arising from these sm studies.
smFI techniques applied to mRNA circularization studies
The movement of a regulator or mRNA should be explored with high spatiotemporal resolution, or the end-to-end distance should be measured directly to examine functional circularization from interactions between translation factors and/or mRNA. The following smFI techniques are frequently used to obtain these parameters.
smFRET
Single-molecule Förster resonance energy transfer (smFRET) is one of the most widely used imaging techniques in the field. It examines the interaction between molecules or the interdistance of residues in a molecule by measuring the efficiency of transferred nonradiative energy from one fluorophore (donor) to another30,31 (acceptor; Fig. 1a). Because energy transfer occurs when two fluorophores are within the range of several nanometers (1–10 nm), i.e., it rapidly changes within the Förster distance (R0), the strength and length of interactions can be determined in real time by taking advantage of the distance-sensitive feature32,33. This technique has been applied to investigate the binding kinetics or dynamic conformation changes in solutions.
a The relationship between FRET efficiency and interdye distance. When the interdistance R between the donor (green) and acceptor (red) is within 10 nm, the excited energy of the donor is transferred to the acceptor, which subsequently emits fluorescence. The FRET efficiency is inversely proportional to the interdye distance, and 50% of the energy is transferred when R is equal to R0. b Principle of smFISH. Cells are fixed and permeabilized for introducing the probes (fluorescently labeled DNA oligonucleotides with ~20 nt, which is a sufficient length for stable hybridization with RNA at room temperature). Typically, 20–50 probes are used for signal amplification. c Schematic showing the mRNP complex or EGFR immobilized on a PEG-biotin surface via either a biotin-conjugated antibody, biotinylated protein, or SBP fused to the protein of interest. A fluorescently labeled antibody is introduced that binds to the target. The heterogeneous state of a single mRNP or EGFR is investigated quantitatively, including the number or combination of constituents. d Nontranslating (green) or translating (yellow) mRNAs in a cell. As the ribosome synthesizes the epitopes in a coding sequence, the fluorescently labeled antibody (red) binds to the nascent epitope, and the fluorescence intensity depends on the number of nascent epitopes. Fluorescently labeled MCP binds to the MS2 stem‒loop in the 3ʹ-UTR, irrespective of the translational state, and indicates the location of the mRNA.
smFISH
Single-molecule fluorescence in situ hybridization (smFISH) is a technique used to detect and localize single mRNAs in a cell by using dye-labeled DNA probes34,35 (Fig. 1b). DNA probes are designed to have a sequence complementary to their mRNA partner, with a 17–22-mer probe used for hybridization. Since smFISH relies on hydrogen bonding between DNA and mRNA, 20–50 probes per transcript are typically used to achieve a sufficient number of base pairs and obtain bright fluorescence signals for high-precision localization. One advantage of smFISH is that even without modifying the target mRNA, the native status of mRNAs in a cell can be determined during translation. Thus, smFISH has been applied to measure the end-to-end distance of mRNAs due to the free labeling positions.
SiMPull
In contrast to conventional blotting assays that show an ensemble-averaged state, the single-molecule pull-down (SiMPull) assay is useful for investigating the heterogeneous state of a target protein complex36,37 (Fig. 1c). Unlike conventional blotting, the target is immobilized on the surface via a specific molecule (antibody36,37, biotin38, or streptavidin-binding peptide39 [SBP]) and detected using fluorescently labeled antibodies. A fluorescence signal indicates whether a specific component is contained in a target. When SiMPull is conducted with multicolor imaging, the combination of fluorescence signals provides the quantitative ratio of components.
NCT or SINAPS
Nascent chain tracking (NCT) and single-molecule imaging of nascent peptides (SINAPS) are novel techniques that visualize nascent proteins synthesized from mRNA in cells via a multimeric array system24,40,41,42,43 (Fig. 1d). A reporter mRNA contains two multimeric arrays in different regions: a repetitive MS2 or PP7 stem‒loop array on the mRNA 3ʹ-UTR and repetitive epitope-coding sequences in an open reading frame. A single mRNA can be visualized by fluorescently labeling the MS2 coat protein (MCP) or PP7 coat protein (PCP), which bind to the MS2 and PP7, respectively. A fluorescence signal is amplified through a repetitive array, which enables the mRNA to be separated from a background signal. Similar to the MS2 or PP7 system, translation with a single mRNA is visualized by using a fluorescently labeled antibody (Fab or scFv) that binds to its epitope. As a ribosome translates the epitope region, the fluorescence signal is increased because more nascent epitopes can be bonded to the antibody. Thus, the location of mRNA and the status of translation in living cells can be visualized using multimeric array systems.
smFI studies in mRNA circularization
Studies have examined mRNA circularization from purified systems to living cells. However, direct observations (end-to-end distance of mRNP) in cells have only been made recently, with several sm studies instead focusing on the association/dissociation or conformational change in these factors and/or mRNA. Here, we describe some aspects related to circularization.
mRNA circularization studies involving purified systems
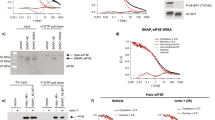
It is believed that circularization begins with the recognition of the 5ʹ-cap of mRNA by eIF4F (eIF4E, eIF4A, and eIF4G), in conjunction with other initiation factors, as the main part of initiation control2. Recognition of eIF4G increases eIF4E’s affinity to a cap or cap analog44,45. To study the 5’ cap-binding kinetics of eIF4E, smFRET was monitored with fluorescently labeled eIF4E in the absence and presence of eIF4G25,46,47 (Fig. 2a). smFRET experiments have demonstrated that full-length yeast eIF4G promotes an alternate conformational state of the RNP complex, causing it to reach the ends of an mRNA. Interestingly, Pab1p, a yeast PABP, has a more significant effect in causing both RNA ends to close even in the absence of poly(A) and/or eIF4G. This observation suggests that PABP may participate in the conformational changes of RNA by directly interacting with mRNA, although its interaction with mRNA itself is relatively weaker than its interaction with the poly(A) tail48,49. A follow-up study on the kinetics of the eIF4E–mRNA interaction has been performed more comprehensively using fluorescently labeled eIF4E, eIF4A, eIF4G, and mRNA immobilized on a surface25. Direct eIF4G–mRNA interactions rather than eIF4E–eIF4G interactions are the main cause for the acceleration of eIF4E-cap binding and the increased association time of eIF4E during translation initiation, which is consistent with a recent in vivo report suggesting that eIF4G may persist in mRNA and facilitate cap-binding activity50. Conversely, another smFRET study found that partial human eIF4G (557-1137) alone slightly affects eIF4E binding kinetics, but the human eIF4F complex substantially changes the eIF4E-cap association46. Interestingly, free eIF4A enhances eIF4E cap accessibility via direct eIF4A-RNA contact, which is similar to the role of eIF4G in eIF4E25. Meanwhile, the binding of eIF4F to RNA results in the dissociation of eIF4E from eIF4F shortly after the formation of the eIF4F-RNA complex, but this process is dependent on ATP hydrolysis (0.07–0.08 s-1; Fig. 2b). Notably, eIF4E and eIF4G are separated simultaneously rather than sequentially, which is consistent with an ensemble study on a yeast system; specifically, eIF4E and 4G preferentially leave mRNA during translation initiation–elongation transition51.
a Schematic of the eIF4E-cap (RNA) binding kinetics in the absence or presence of other factors (eIF4G, eIF4A, and Pab1p) alongside the FRET value. Capped RNA is immobilized on the PEG-biotin surface via streptavidin-biotin interactions. FRET indicates that Cy5-eIF4E binds to the cap of Cy3-RNA. The FRET value increases in the presence of other factors. b Schematic and intensity-time trace representing the comprehensive eIF4F-cap (mRNA) binding kinetics during translation initiation. eIF4F (eIF4E, eIF4A and eIF4G) binds to the mRNA cap (coarrival). Shortly after (~0.08 s-1), eIF4E dissociates from mRNA with eIF4G (codeparture). Meanwhile, eIF4E-cap binding is enhanced by the interaction between “free” eIF4A and mRNA, independent of ATP hydrolysis. c The effect of structure and length of mRNA on eIF4E association, which depends on the mRNA length and complexity but not on the free energy. d Left: Schematic representation of an smFRET system that directly monitors the bent conformation of fluorescently labeled RRM1-4 in the absence (left) or presence (right) of poly(A)_25 RNA alongside a histogram of the FRET efficiency in the absence or presence of poly(A)_25 RNA. Right: Schematic figure representing the role of bent conformation in mRNA circularization for translation. It enhances the eIF4G–PABP interaction, enhancing the formation of 80S in an in vitro translation system. e Direct measurement of the end-to-end distance of mRNA. The interdistance of two dyes (d1) does not reflect the actual end-to-end distance (d2). The fluorophore linker length is considered to obtain an exact value, producing 5–9 nm among the investigated mRNAs. f Schematic of a single polysome tethering assay that visualizes the heterogeneous state of a single mRNP complex for a monosome (left) and a polysome (right). The components of the mRNP complex were captured by fluorescently labeled antibodies. A pie chart indicating that most mRNAs contain only one factor.
The effect of the structural diversity of mRNA52 or regulators28,29 on translation heterogeneities has recently been demonstrated. Indeed, translation is regulated by the secondary structure53, length54,55, mRNA sequence56, posttranscriptional modifications (e.g., m6A modification57), and poly(A) length29. Wang et al. demonstrated that the addition of a small stem loop (ΔG = − 4.8 kcal/mol) in the 5ʹ-UTR is sufficient to perturb the state in translation initiation and delay translation58, which is not detectable in an ensemble assay. The innate 5ʹ-UTR structure affects protein–mRNA dynamics59. The eIF4E–mRNA association is inversely proportional to the degree of secondary structure formation on the 5ʹ-UTR of the mRNA (Fig. 2c). Interestingly, the rate of eIF4E–mRNA association depends not on the free energy of the secondary structure but on the degree of complexity of the secondary structure. The rate of association between eIF4E and mRNA tends to increase as the total length of mRNA shortens, and the eIF4G–mRNA interaction accelerates the association rate of eIF4E in proportion to the length of mRNA; consequently, less deviation occurs among mRNAs25. Conformational changes in a regulator can play a crucial role in circularization. According to our smFRET study, conformational changes in PABP, specifically in the RRM2–RRM3 region, are induced when PABP binds to a poly(A) tail of RNA, and this conformational change affects PABP–poly(A)29 and PABP–eIF4G28 interactions (Fig. 2d). Mutation of the region inhibits the PABP–eIF4G interaction and inhibits the efficient formation of 80S ribosomes. Consistent with the canonical view, this result suggests the importance of PABP binding to the poly(A) tail in an end-proximity form of mRNA. Recently, this bent conformation of PABP was re-examined via cryo-EM60.
Despite structure-driven circularization, the direct measurement of the RNA end-to-end distance shows the inherent proximity of the mRNA ends. An smFRET study was conducted to experimentally determine the end-to-end distances of mRNAs of various lengths (500–5,500 nt) from viruses and fungi61 (Fig. 2e). mRNAs were labeled with a dye at each end, and the interdye distance was measured from the FRET efficiency. Interestingly, all mRNAs used in this study have an end-to-end distance within 5–9 nm without any protein, irrespective of length, origin, and secondary structure. Before this initial experimental demonstration, several theoretical calculations had estimated that the end-to-end distance of mRNA should be small and not dependent on mRNA length62,63,64. More recently, the inherent end-to-end distance was more systematically investigated with human mRNA and lncRNA65. Computational analysis and smFRET have shown that proximity is attributed not only to base pairing in the 5ʹ- or 3ʹ-UTR but also to stem loops formed in the whole sequence. Furthermore, even in the firefly luciferase ORF without a 5ʹ- and 3ʹ-UTR, the end-to-end distance was 4.9 nm. This inherent closeness of both ends of RNA in the absence of proteins is consistent with reports on cells described in the next section.
mRNA circularization in single polysomes
Many in vitro studies demonstrate functional circularization via translation factors. However, no direct investigation has focused on whether the circularized mRNP is formed via a 5ʹ-cap-eIF4E–eIF4G–PABP–poly(A) interaction in cells. Therefore, an sm blotting experiment was conducted using polysomal or subpolysomal fractions to find direct evidence of functional circularization39 (Fig. 2f). Immobilization of the cross-linked polysomal fraction to a PEG-biotin surface was performed using biotin-conjugated or fluorescently labeled antibodies against eIF4E, eIF4G or PABP. Strikingly, the results exhibited rare colocalization between factors in all fractions. Specifically, most mRNAs have only one member. Because endogenous mRNPs were cross-linked under normal conditions, this result suggested that mRNA circularization cannot occur in an active translation state. Furthermore, even in the case of an m6A-containing reporter mRNA, which was reported previously, the mRNA is more likely circularized through the interaction of the m6A reader protein YTHDF family14 or METTL315 with eIFs; we could not find an increase in colocalization between factors39. Although a different m6A reporter was used, m6A-containing mRNA likely increases the translation efficiency, but it does not increase the level of mRNA circularization during translation.
mRNA circularization in cells
Recent advances in single-RNA cellular imaging techniques have enabled the visualization of mRNA circularization during translation. Although it is primitive, the most direct method to investigate circularization is by measuring the distance of the mRNA compartment. Morisaki et al. quantified the size of polysomes with three different lengths (125, 374, and 1544 a.a.) by measuring the distance between nascent peptide chains and the 3ʹ-UTR of mRNA24 (Fig. 3a). Interestingly, NCT experiments have shown that the distance is shorter (65–105 nm) than expected, suggesting that polysomes are compact rather than extended. This is consistent with previous reports involving ET66 or EM67 (or cryo-EM) describing that polysomes have either a rosette68,69, helical70 or spiral shape71. However, the size was not correlated with the length of mRNA in this experiment. More recently, two groups reported a striking result: a closed loop is not a stable state of translating mRNA72,73. smFISH and SunTag signals were monitored under normal or stress (nontranslating) conditions to determine the architecture of mRNP (Fig. 3b). An smFISH experiment with a hybridizing probe at both ends of mRNA demonstrated that mRNP compaction depends on the translation state. More specifically, ribosome release is the main cause of compaction because compaction occurs in the 5ʹ- to 3ʹ-direction when translation initiation is inhibited72. Only under stress conditions, such as treatment with arsenite, heat, or translation inhibitors, does mRNP take a globular form, which is likely to cause functional circularization. The translating mRNP also possesses a compact (or globular) conformation, although to a lesser extent than under stress conditions. These findings are consistent with the results of the above NCT experiment; however, it is insufficient to cause both ends of the mRNA to connect. Interestingly, both studies have demonstrated the dependency of end-to-end distance on the ribosome occupancy of mRNA. Indeed, the longer the ORF, the greater the degree of ribosome elongation because the end-to-end distance is correlated with the length of mRNA. However, this result is different from that of the NCT experiment. Furthermore, the breakage of the eIF4G–PABP interaction does not lead to changes in compaction73, suggesting that the interaction between translation factors may not be a critical factor for the circularization of mRNA and that the proximity of the ends is an intrinsic feature of mRNA. This idea is also supported by the fact that long noncoding (lnc) RNA has compactness similar to that of nontranslating mRNA65. In addition, nuclear mRNA is slightly less compact than translationally inhibited mRNA because of the binding of RBPs73.
Schematic representing nontranslating or translating mRNA and the measurement of its end-to-end distance in a cell (fixed or live). a The distance between the nascent epitope and the PP7 stem‒loop indicates the compactness of the translating mRNP. b Direct end-to-end distance measurement of mRNA during translation depending on the translation state through the utilization of a smFISH probe in a fixed cell. As ribosome occupancy increases, the distance between the two ends also increases. c End-to-end distance measurement of mRNA during cap or IRES translation (emerald) or both (orange). Cap translation leads to a longer distance than IRES translation resulting from ribosome occupancy.
mRNA does not only undergo circularization when in the presence of eIF4E–eIF4G–PABP. Recently, Koch et al. demonstrated IRES-mediated mRNA compaction23. They constructed a reporter mRNA molecule that contains a cap and IRES downstream of the cap-ORF, which produced a 10x FLAG epitope and a 24x SunTag by the cap and IRES, respectively (Fig. 3c). In this study, the distances of the cap-ORF to 3ʹ-UTR and IRES-ORF to 3ʹ-UTR were measured in cap- or IRES-only translation or cap+IRES translation. Interestingly, the distance of the IRES-ORF to the 3ʹ-UTR was similar to that of the cap to the 3ʹ-UTR, suggesting that the IRES-ORF is compact when it is idle, possibly because of the lack of ribosomes on the IRES-ORF. The distance of ORF to 3ʹ-UTR in both cases is proportional to intensity, that is, the number of ribosomes, which is consistent with previous studies on fixed cells72,73. Furthermore, in the case of cap+IRES translation, the distance from the cap-ORF to the 3ʹ-UTR is slightly decreased, but the distance from the IRES-ORF to the 3ʹ-UTR is significantly decreased compared to cap- or IRES-only translation. This decrease occurs because there are fewer available ribosomes in cap+IRES translation than in cap- or IRES-only translation; the decreased number of ribosomes loaded on mRNA is represented by the distance decrease, which also supports the effect of ribosomes on mRNA compaction.
Future perspectives
SmFI has provided an opportunity to examine mRNA circularization in view of the interaction between proteins and mRNAs, especially in relation to time. If translation or ribosome occupancy is the criterion for mRNA circularization, then how can recent sm data be reconciled with a considerable amount of ensemble data? Biochemically, circularization should occur on the basis of the binding affinities of protein–protein or protein–mRNA interactions and the concentration of proteins in cells74,75. During translation elongation, initiation also occurs. Then, how can ribosomes be reinitiated after termination? Does the communication of both ends of mRNA occur only in a nontranslating state? Is the extension of mRNA by loaded ribosomes sufficient to break the eIF4G–PABP or the eIF4E–eIF4G interaction? Or is there an unknown factor? Another interesting question is when functional circularization occurs. Indeed, a recent study has shown that the initiation rate increases only after the first ribosome has completed translation to capped and polyadenylated mRNA3. Here, it is possible that the first ribosome drags eIF4F and meets PABP at the 3ʹ-end, thus making the physical link. In addition to classical circularization via 5ʹ-cap-eIF4E–eIF4G–PABP-3ʹ-poly(A), mRNA circularization mediated by other structural IRESs12,13, m6A14,15, or 3’ cap-independent translational enhancers16,76 should be addressed for a comprehensive understanding. The smFI methods will help answer these questions. Since the development of NCT or SINAPS, the sm study of translational kinetics/dynamics in living cells has focused on various subjects77,78,79,80,81. Nevertheless, it is still difficult to track single or multiple translation regulators at the same time owing to the high intracellular concentration of the targets74,75, although one example has been reported50. Further advances in protein labeling strategies are also required beyond the several methods that have been developed, including small peptide tags and their cognate antibodies82,83,84, the incorporation of unnatural amino acids85 or chemical labeling without genetic manipulation86. Because of its high specificity and efficiency, the Halo-87 or SNAP-tag88 is still the preferred strategy despite the potential problems caused by the large tag size. Additionally, technical advances in protein purification are required for experiments in an in vitro system. Currently, to our knowledge, no purification of full-length human eIF4G has been reported46. In addition to the advancement of techniques, multiple approaches can be integrated. Recently, a study combined two methods, namely, fluorescence correlation spectroscopy (FCS) and SPT, to investigate cap-dependent translation initiation in living cells50. Each method complements the other’s shortcomings and reveals the temporal regulation of cap-binding activity at extraordinary resolutions.
Concluding remarks
Functional mRNA circularization based on protein–protein or RNA–protein interactions has been considered a canonical dogma in translational studies. However, recent sm studies have shown that mRNA appears circularized, yet it cannot be physically connected by proteins in active translation. Although the smFI technique does not provide a complete solution, it is currently the most optimal method used for visualizing the dynamic behavior of single mRNP in intact cells. In the future, smFI may contribute to a comprehensive understanding of how we can change interactions in an mRNP complex and the resulting conformation of mRNA throughout translation.
References
Dever, T. E., Kinzy, T. G. & Pavitt, G. D. Mechanism and regulation of protein synthesis in Saccharomyces cerevisiae. Genetics 203, 65–107 (2016).
Jackson, R. J., Hellen, C. U. & Pestova, T. V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 11, 113–127 (2010).
Alekhina, O. M., Terenin, I. M., Dmitriev, S. E. & Vassilenko, K. S. Functional cyclization of eukaryotic mRNAs. Int. J. Mol. Sci. 21, 1677 (2020).
Kahvejian, A., Roy, G. & Sonenberg, N. The mRNA closed-loop model: the function of PABP and PABP-interacting proteins in mRNA translation. Cold Spring Harb. Symp. Quant. Biol. 66, 293–300 (2001).
Pelletier, J. & Sonenberg, N. The organizing principles of eukaryotic ribosome recruitment. Annu. Rev. Biochem. 88, 307–335 (2019).
Marshall, E., Stansfield, I. & Romano, M. C. Ribosome recycling induces optimal translation rate at low ribosomal availability. J. R. Soc. Interface 11, 20140589 (2014).
Chou, T. Ribosome recycling, diffusion, and mRNA loop formation in translational regulation. Biophys. J. 85, 755–773 (2003).
Schoenberg, D. R. & Maquat, L. E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012).
Garneau, N. L., Wilusz, J. & Wilusz, C. J. The highways and byways of mRNA decay. Nat. Rev. Mol. Cell Biol. 8, 113–126 (2007).
Walters, B. & Thompson, S. R. Cap-Independent translational control of carcinogenesis. Front. Oncol. 6, 128 (2016).
Deo, R. C., Bonanno, J. B., Sonenberg, N. & Burley, S. K. Recognition of polyadenylate RNA by the poly(A)-binding protein. Cell 98, 835–845 (1999).
Yang, Y. & Wang, Z. IRES-mediated cap-independent translation, a path leading to hidden proteome. J. Mol. Cell. Biol. 11, 911–919 (2019).
Hellen, C. U. & Sarnow, P. Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev. 15, 1593–1612 (2001).
Wang, X. et al. N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Choe, J. et al. mRNA circularization by METTL3-eIF3h enhances translation and promotes oncogenesis. Nature 561, 556–560 (2018).
Simon, A. E. & Miller, W. A. 3’ cap-independent translation enhancers of plant viruses. Annu. Rev. Microbiol. 67, 21–42 (2013).
Groft, C. M. & Burley, S. K. Recognition of eIF4G by rotavirus NSP3 reveals a basis for mRNA circularization. Mol. Cell 9, 1273–1283 (2002).
Vende, P., Piron, M., Castagne, N. & Poncet, D. Efficient translation of rotavirus mRNA requires simultaneous interaction of NSP3 with the eukaryotic translation initiation factor eIF4G and the mRNA 3’ end. J. Virol. 74, 7064–7071 (2000).
Imataka, H., Gradi, A. & Sonenberg, N. A newly identified N-terminal amino acid sequence of human eIF4G binds poly(A)-binding protein and functions in poly(A)-dependent translation. EMBO J. 17, 7480–7489 (1998).
Tarun, S. Z. Jr & Sachs, A. B. Association of the yeast poly(A) tail binding protein with translation initiation factor eIF-4G. EMBO J. 15, 7168–7177 (1996).
Wagner, S. et al. Selective translation complex profiling reveals staged initiation and co-translational assembly of initiation factor complexes. Mol. Cell 79, 546–560.e547 (2020).
Wells, S. E., Hillner, P. E., Vale, R. D. & Sachs, A. B. Circularization of mRNA by eukaryotic translation initiation factors. Mol. Cell 2, 135–140 (1998).
Koch, A., Aguilera, L., Morisaki, T., Munsky, B. & Stasevich, T. J. Quantifying the dynamics of IRES and cap translation with single-molecule resolution in live cells. Nat. Struct. Mol. Biol. 27, 1095–1104 (2020).
Morisaki, T. et al. Real-time quantification of single RNA translation dynamics in living cells. Science 352, 1425–1429 (2016).
Cetin, B. & O’Leary, S. E. mRNA- and factor-driven dynamic variability controls eIF4F-cap recognition for translation initiation. Nucleic Acids Res. 50, 8240–8261 (2022).
Wang, J. et al. eIF5B gates the transition from translation initiation to elongation. Nature 573, 605–608 (2019).
Andreou, A. Z., Harms, U. & Klostermeier, D. Single-stranded regions modulate conformational dynamics and ATPase activity of eIF4A to optimize 5’-UTR unwinding. Nucleic Acids Res. 47, 5260–5275 (2019).
Hong, K. Y. et al. The bent conformation of poly(A)-binding protein induced by RNA-binding is required for its translational activation function. RNA Biol. 14, 370–377 (2017).
Lee, S. H. et al. Poly(A) RNA and Paip2 act as allosteric regulators of poly(A)-binding protein. Nucleic Acids Res 42, 2697–2707 (2014).
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl Acad. Sci. USA 93, 6264–6268 (1996).
Algar, W. R., Hildebrandt, N., Vogel, S. S. & Medintz, I. L. FRET as a biomolecular research tool - understanding its potential while avoiding pitfalls. Nat. Methods 16, 815–829 (2019).
Liu, J. et al. Cascading MutS and MutL sliding clamps control DNA diffusion to activate mismatch repair. Nature 539, 583–587 (2016).
Ha, T. et al. Single-molecule fluorescence spectroscopy of enzyme conformational dynamics and cleavage mechanism. Proc. Natl Acad. Sci. USA 96, 893–898 (1999).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Raj, A., van den Bogaard, P., Rifkin, S. A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Yeom, K. H. et al. Single-molecule approach to immunoprecipitated protein complexes: insights into miRNA uridylation. EMBO Rep. 12, 690–696 (2011).
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011).
Kim, K. L. et al. Pairwise detection of site-specific receptor phosphorylations using single-molecule blotting. Nat. Commun. 7, 11107 (2016).
Kim, B. et al. Single polysome analysis of mRNP. Biochem. Biophys. Res. Commun. 618, 73–78 (2022).
Yan, X., Hoek, T. A., Vale, R. D. & Tanenbaum, M. E. Dynamics of translation of single mRNA molecules in vivo. Cell 165, 976–989 (2016).
Wu, B., Eliscovich, C., Yoon, Y. J. & Singer, R. H. Translation dynamics of single mRNAs in live cells and neurons. Science 352, 1430–1435 (2016).
Wang, C., Han, B., Zhou, R. & Zhuang, X. Real-time imaging of translation on single mRNA transcripts in live. Cells Cell 165, 990–1001 (2016).
Pichon, X. et al. Visualization of single endogenous polysomes reveals the dynamics of translation in live human cells. J. Cell Biol. 214, 769–781 (2016).
Moerke, N. J. et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 128, 257–267 (2007).
von Der Haar, T., Ball, P. D. & McCarthy, J. E. Stabilization of eukaryotic initiation factor 4E binding to the mRNA 5’-Cap by domains of eIF4G. J. Biol. Chem. 275, 30551–30555 (2000).
Hong, H.J., Guevara, M.G., Lin, E. & O’Leary, S.E. Single-molecule dynamics of SARS-CoV-2 5’ cap recognition by human eIF4F. bioRxiv https://doi.org/10.1101/2021.05.26.445185 (2021).
O’Leary, S. E., Petrov, A., Chen, J. & Puglisi, J. D. Dynamic recognition of the mRNA cap by Saccharomyces cerevisiae eIF4E. Structure 21, 2197–2207 (2013).
Kini, H. K., Silverman, I. M., Ji, X., Gregory, B. D. & Liebhaber, S. A. Cytoplasmic poly(A) binding protein-1 binds to genomically encoded sequences within mammalian mRNAs. RNA 22, 61–74 (2016).
Sladic, R. T., Lagnado, C. A., Bagley, C. J. & Goodall, G. J. Human PABP binds AU-rich RNA via RNA-binding domains 3 and 4. Eur. J. Biochem. 271, 450–457 (2004).
Gandin, V. et al. Cap-dependent translation initiation monitored in living cells. Nat. Commun. 13, 6558 (2021).
Wang, X. et al. Stoichiometry and change of the mRNA closed-loop factors as translating ribosomes transit from initiation to elongation. Plos One 11, e0150616 (2016).
Sonneveld, S., Verhagen, B. M. P. & Tanenbaum, M. E. Heterogeneity in mRNA translation. Trends Cell Biol. 30, 606–618 (2020).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Paek, K. Y. et al. Translation initiation mediated by RNA looping. Proc. Natl Acad. Sci. USA 112, 1041–1046 (2015).
Santhanam, A. N. et al. Role of 3’UTRs in the translation of mRNAs regulated by oncogenic eIF4E-a computational inference. Plos ONE 4, e4868 (2009).
Hanson, G. & Coller, J. Codon optimality, bias and usage in translation and mRNA decay. Nat. Rev. Mol. Cell Biol. 19, 20–30 (2018).
Meyer, K. D. m(6)A-mediated translation regulation. Biochim. Biophys. Acta Gene Regul. Mech. 1862, 301–309 (2019).
Wang, H., Sun, L., Gaba, A. & Qu, X. An in vitro single-molecule assay for eukaryotic cap-dependent translation initiation kinetics. Nucleic Acids Res 48, e6 (2020).
Cetin, B., Song, G. J. & O’Leary, S. E. Heterogeneous dynamics of protein-RNA interactions across transcriptome-derived messenger RNA populations. J. Am. Chem. Soc. 142, 21249–21253 (2020).
Schafer, I. B. et al. Molecular basis for poly(A) RNP architecture and recognition by the Pan2-Pan3 deadenylase. Cell 177, 1619–1631.e21 (2019).
Leija-Martinez, N. et al. The separation between the 5 ‘-3 ‘ ends in long RNA molecules is short and nearly constant. Nucleic Acids Res. 42, 13963–13968 (2014).
Yoffe, A. M., Prinsen, P., Gelbart, W. M. & Ben-Shaul, A. The ends of a large RNA molecule are necessarily close. Nucleic Acids Res. 39, 292–299 (2011).
Fang, L. T. The end-to-end distance of RNA as a randomly self-paired polymer. J. Theor. Biol. 280, 101–107 (2011).
Clote, P., Ponty, Y. & Steyaert, J. M. Expected distance between terminal nucleotides of RNA secondary structures. J. Math. Biol. 65, 581–599 (2012).
Lai, W. J. C. et al. mRNAs and lncRNAs intrinsically form secondary structures with short end-to-end distances. Nat. Commun. 9, 4328 (2018).
Skoglund, U., Andersson, K., Strandberg, B. & Daneholt, B. Three-dimensional structure of a specific pre-messenger RNP particle established by electron microscope tomography. Nature 319, 560–564 (1986).
Batisse, J., Batisse, C., Budd, A., Bottcher, B. & Hurt, E. Purification of nuclear poly(A)-binding protein Nab2 reveals association with the yeast transcriptome and a messenger ribonucleoprotein core structure. J. Biol. Chem. 284, 34911–34917 (2009).
Madin, K. et al. Formation of circular polyribosomes in wheat germ cell-free protein synthesis system. FEBS Lett. 562, 155–159 (2004).
Wettstein, F. O., Staehelin, T. & Noll, H. Ribosomal aggregate engaged in protein synthesis: characterization of the ergosome. Nature 197, 430–435 (1963).
Brandt, F., Carlson, L. A., Hartl, F. U., Baumeister, W. & Grunewald, K. The three-dimensional organization of polyribosomes in intact human cells. Mol. Cell 39, 560–569 (2010).
Palade, G. E. A small particulate component of the cytoplasm. J. Biophys. Biochem. Cytol. 1, 59–68 (1955).
Khong, A. & Parker, R. mRNP architecture in translating and stress conditions reveals an ordered pathway of mRNP compaction. J. Cell Biol. 217, 4124–4140 (2018).
Adivarahan, S. et al. Spatial organization of single mRNPs at different stages of the gene expression pathway. Mol. Cell 72, 727–738.e725 (2018).
Gu, S. et al. The flip-flop configuration of the PABP-dimer leads to switching of the translation function. Nucleic Acids Res. 50, 306–321 (2022).
Merrick, W. C. & Pavitt, G. D. Protein synthesis initiation in eukaryotic cells. Cold Spring Harb. Perspect. Biol. 10, a033092 (2018).
Guo, L., Allen, E. M. & Miller, W. A. Base-pairing between untranslated regions facilitates translation of uncapped, nonpolyadenylated viral RNA. Mol. Cell 7, 1103–1109 (2001).
Kobayashi, H. & Singer, R. H. Single-molecule imaging of microRNA-mediated gene silencing in cells. Nat. Commun. 13, 1435 (2022).
Cialek, C. A. et al. Imaging translational control by Argonaute with single-molecule resolution in live cells. Nat. Commun. 13, 3345 (2022).
Cialek, C. A., Koch, A. L., Galindo, G. & Stasevich, T. J. Lighting up single-mRNA translation dynamics in living cells. Curr. Opin. Genet. Dev. 61, 75–82 (2020).
Boersma, S. et al. Translation and replication dynamics of single RNA viruses. Cell 183, 1930–1945.e1923 (2020).
Bruurs, L. J. M. et al. Heterogeneity in viral replication dynamics shapes the antiviral response. bioRxiv https://doi.org/10.1101/2022.06.08.495262 (2022).
Liu, Y. et al. Visualizing looping of two endogenous genomic loci using synthetic zinc-finger proteins with anti-FLAG and anti-HA frankenbodies in living cells. Genes Cells 26, 905–926 (2021).
Traenkle, B., Segan, S., Fagbadebo, F. O., Kaiser, P. D. & Rothbauer, U. A novel epitope tagging system to visualize and monitor antigens in live cells with chromobodies. Sci. Rep. 10, 14267 (2020).
Gotzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Lee, K. J., Kang, D. & Park, H. S. Site-specific labeling of proteins using unnatural amino acids. Mol. Cell. 42, 386–396 (2019).
Shiraiwa, K., Cheng, R., Nonaka, H., Tamura, T. & Hamachi, I. Chemical tools for endogenous protein labeling and profiling. Cell Chem. Biol. 27, 970–985 (2020).
Los, G. V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Acknowledgements
This research was supported by the National Research Foundation of Korea funded by the Ministry of Science and ICT (2020R1A2C1012345 to J.-B.L).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, B., Seol, J., Kim, Y.K. et al. Single-molecule visualization of mRNA circularization during translation. Exp Mol Med 55, 283–289 (2023). https://doi.org/10.1038/s12276-023-00933-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-023-00933-1