Abstract
We aimed to explore whether changes in circulating levels of miRNAs according to type 2 diabetes mellitus (T2DM) or prediabetes status could be used as biomarkers to evaluate the risk of developing the disease. The study included 462 patients without T2DM at baseline from the CORDIOPREV trial. After a median follow-up of 60 months, 107 of the subjects developed T2DM, 30 developed prediabetes, 223 maintained prediabetes and 78 remained disease-free. Plasma levels of four miRNAs related to insulin signaling and beta-cell function were measured by RT-PCR. We analyzed the relationship between miRNAs levels and insulin signaling and release indexes at baseline and after the follow-up period. The risk of developing disease based on tertiles (T1-T2-T3) of baseline miRNAs levels was evaluated by COX analysis. Thus, we observed higher miR-150 and miR-30a-5p and lower miR-15a and miR-375 baseline levels in subjects with T2DM than in disease-free subjects. Patients with high miR-150 and miR-30a-5p baseline levels had lower disposition index (p = 0.047 and p = 0.007, respectively). The higher risk of disease was associated with high levels (T3) of miR-150 and miR-30a-5p (HRT3-T1 = 4.218 and HRT3-T1 = 2.527, respectively) and low levels (T1) of miR-15a and miR-375 (HRT1-T3 = 3.269 and HRT1-T3 = 1.604, respectively). In conclusion, our study showed that deregulated plasma levels of miR-150, miR-30a-5p, miR-15a, and miR-375 were observed years before the onset of T2DM and pre-DM and could be used to evaluate the risk of developing the disease, which may improve prediction and prevention among individuals at high risk for T2DM.
Similar content being viewed by others
Introduction
Type 2 diabetes mellitus (T2DM) is a major public health problem, and its prevalence has been increasing over the past few decades with no signs of receding in the near future. In addition to personal affliction, this disease imposes a heavy economic burden on the global health-care system and overall economy1.
In prediabetic conditions, beta cells compensate the insulin-resistant state from target tissues by increasing both the mass of pancreatic islets and the release of insulin in response to dietary glucose and fatty acids2. Chronic stress conditions may induce the activation of adaptation processes that exceed the normal phenotypic flexibility, leading to progressive inflexibility3 and beta-cell dysfunction, followed by elevated fasting glucose (IFG) and/or impaired glucose tolerance (IGT)4,5. Thus, the degree of hyperglycemia reflects the severity of the metabolic process, suggesting that it is an important marker to determine the developmental stage of the disease6,7.
However, T2DM frequently goes undiagnosed for many years because hyperglycemia develops gradually and, in its earlier stages, is not severe enough for the patient to note any of the classic symptoms of diabetes8. For this reason, the tools used currently to screen and diagnose T2DM and to detect individuals with prediabetes (Pre-DM) do not adequately predict the onset of the disease and monitor its progression9. Therefore, more informative biomarkers are needed to identify beta-cell injury and to assess the risk of disease, monitor responses to treatment, personalize therapy and improve patient quality of life10.
Currently, miRNAs are recognized as important regulators of gene expression and central players in the control of several biological and pathological processes, including T2DM11. To this end, in vitro models have shown that miR-30a-5p12 and miR-144 play a role in beta-cell dysfunction13. Moreover, the plasma concentrations of individual14 and multiple circulating miRNAs have been identified as being significantly different between T2DM patients and prediabetic subjects15,16. In addition, alterations in plasma miRNAs levels have been observed in other metabolic diseases, such as metabolic syndrome15,17. However, those experimental designs were not able to identify whether the differences in miRNA levels found were the cause or a consequence of the development of metabolic disease. To the best of our knowledge, only two studies have observed alterations in plasma miRNAs levels before T2DM development18,19.
This imparts a need to conduct long-term follow-up studies in larger populations with incident cases of T2DM to identify the role of miRNA during the development of T2DM and to use that information to assist in the development of predictive biomarkers of T2DM. Therefore, in this study, we aimed to study whether plasma circulating levels of miRNAs according to T2DM or pre-DM status could be used as biomarkers to evaluate the risk of developing the disease.
Materials and methods
Study subjects
This work was conducted within the framework of the CORDIOPREV study, whose rationale, methods and baseline characteristics have been reported by Delgado-Lista et al.20. Briefly, the CORDIOPREV study is an ongoing prospective, randomized, single-blind, controlled dietary intervention trial in 1002 coronary health disease (CHD) patients with a high risk of cardiovascular disease who had their last coronary event more than 6 months before enrollment. In addition to conventional treatment for CHD, the subjects were randomized into two different dietary models (Mediterranean and low-fat diets). The intervention phase is still in progress and will have a median follow-up of seven years. Patients were recruited from November 2009 to February 2012, mostly at the Reina Sofia University Hospital (Córdoba, Spain), but patients from other hospitals from the Córdoba and Jaen provinces were also admitted. In summary, the patients were eligible if they were between 20 and 75 years of age, had established CHD without any clinical events in the last 6 months, were thought to follow a long-term dietary intervention and had no severe diseases or a life expectancy of five years. Details of the trial design were provided in Clinicaltrials.gov (NTC00924937). Written consent was obtained from all subjects prior to recruitment and initiation of the study protocol, and all the amendments were approved by the Ethics Committee of Hospital Reina Sofia, all of which follow the Helsinki Declaration and good clinical practices.
All patients from the CORDIOPREV-DIAB without T2DM at baseline according to the American Diabetes Association (ADA) diagnostic criteria21 (N = 462) were included in this study22. T2DM and pre-DM were diagnosed according to the ADA diagnostic criteria21 and were evaluated by glucose tolerance tests performed each year during the five years of follow-up. Prediabetic status was defined as having one or more of the following criteria present in a participant: fasting plasma glucose (FPG) concentration ≥100 and <126 mg/dL, impaired fasting glucose (IFG); FPG ≥ 140 and <200 mg/dL 2 h after an oral glucose test (OGTT), impaired glucose tolerance (IGT)23 and glycosylated hemoglobin (HbA1c) ≥5.7 and <6.4%24.
In a median follow-up of 60 months, 78 subjects did not develop T2DM or pre-DM (non-T2DM); 239 subjects were pre-DM at baseline, of which 223 subjects maintained pre-DM status during the follow-up period (pre-DM); 30 subjects were not pre-DM at baseline but developed pre-DM in the follow-up period (incident pre-DM). Finally, 107 of the participating subjects developed T2DM (incident-T2DM) in a median follow-up of 60 months. Moreover, 24 patients were not included in this study due to declining participation, death and withdrawing for other reasons. The baseline characteristics of the subjects in the study are shown in Table 1.
Biochemical measurements of metabolic parameters
Venous blood from the participants was collected in tubes containing EDTA after a 12-h overnight fast. Lipid variables were assessed with the modular auto analyzer DDPPII Hitachi (Roche, Basel, Switzerland) using specific reagents (Boehringer-Mannheim, Mannheim, Germany). Measurements of total cholesterol (TC) and triglycerides (TG) levels were made by colorimetric enzymatic methods25,26; high-density lipoprotein-cholesterol (HDL-c) levels were measured by colorimetric assay27; and low-density lipoprotein (LDL-C) concentrations were calculated by the Friedewald equation, using the following formula: LDL-C = CT - (HDL-C + TG/5). Glucose measurements were performed using the hexokinase method. hs-C-reactive protein (hs-CRP) concentration was determined by high-sensitivity ELISA (BioCheck, Inc., Foster City, CA, USA). Plasma insulin concentrations were measured by microparticle enzyme immunoassay (Abbott Diagnostics, Matsudo-shi, Japan). Non-esterified fatty acid concentrations were measured by enzymatic colorimetric assay (Roche Diagnostics, Penzberg, Germany). ApoA-1 and ApoB concentrations were determined by immunoturbidimetry.
Insulin signaling and release indexes
Before starting the test, the patients abstained from food and medications for 12 h and were asked to refrain from smoking during the fasting period and from alcohol intake during the preceding seven days. They were also requested to avoid strenuous physical activity the day before the test was administered. Oral glucose tolerance test (OGTT): At 8:00 a.m., the patients were admitted to the laboratory, and an OGTT (75 g dextrose monohydrate in 250 mL water, NUTER. TEC GLUCOSA 50) was performed with 0, 30, 60, and 120 min sampling to establish plasma glucose and insulin levels28.
The insulin sensitivity index (ISI) was calculated in the OGTT using the following formula: ISI = 10.000 ÷ √([fasting plasma glucose × fasting plasma insulin]×[mean glucose in OGTT × mean insulin in OGTT])28. HOMA-IR was calculated as previously described by Song, Y., et al.29. Insulin secretion was measured using the insulinogenic index (IGI): IGI = [30 min insulin−fasting insulin (pmol/L)]/[30 min glucose−fasting glucose (mmol/L)30. The disposition index (DI) was estimated as follows: DI = ISI × [AUC30 min insulin/AUC30 min glucose], where AUC30 min is the area under the curve between the baseline and 30 min of the OGTT for insulin (pmol/L) and glucose (mmol/L) measurements, respectively, calculated by the trapezoidal method31. The hepatic insulin resistance index (HIRI) and the muscle insulin sensitivity index (MISI) were used to evaluate tissue-specific IR as described in previous work by our group22, following the methods described by Matsuda and DeFronzo for HIRI and Abdul-Ghani et al. for MISI28,32.
Isolation of circulating miRNAs
Circulating miRNAs were isolated from plasma samples, which were obtained using an EDTA-containing blood tube. In this way, venous blood from the participants was obtained by venipuncture and kept on ice until centrifugation. Immediately, to separate the plasma from the erythrocytes and the buffy coat fractions, EDTA-containing blood tubes were centrifuged at 2.000×g for 10 min at 4 °C. The superior phase corresponding to the plasma was carefully separated using a pasteur pipette, and the sample was stored at −80 °C until use. The plasma samples were then thawed on ice for RNA isolation.
Total RNA from plasma was isolated using an miRNeasy Mini Kit (Qiagen, Hilden, Germany). In short, 200 µL of EDTA-plasma was mixed with 1 mL of Qiazol, incubated for 5 min at room temperature and then mixed with 200 µL of chloroform. We added 2 μg of MS2 RNA carrier (Roche. Mannheim, Germany) before the chloroform protocol step. The organic and aqueous phases were separated by centrifugation at 12,000×g for 15 min at 4 °C. The aqueous phase was collected, and RNA was precipitated by adding 100% ethanol. The mixture was applied to an miRNeasy Mini spin column and centrifuged at 8000×g for 2 min. Next, 700 µL of RWT buffer was added to an RNeasy MinElute spin column at 8000×g for 2 min. The contents in the column were washed with 500 µL of RPE buffer and 500 µL of 80% ethanol. RNA was eluted in 14 μL of RNase-free water. RNA purity and concentration were evaluated by spectrophotometry using a NanoDrop ND-2000 (ThermoFisher, Waltham, MA).
Retrotranscription and preamplification of miRNAs
The study of miRNAs expression was carried out on four miRNAs (miR-150; miR-30a-5p; miR-15a and miR-375) selected on the basis of previous evidence of their participation in the biomechanisms involved in T2DM pathogenesis–in particular, beta cell function, insulin secretion and insulin sensitivity12,33,34,35,36,37. The retrotranscription of RNA was carried out using a TaqMan MicroRNA Reverse Transcription Kit (Life Technologies, Carlsbad, CA, USA). The RT mix contained 2 μL of RNA and 3 µL of RT custom primer pool in a final volume of 7.5 μL. The RT primer pool was customized by selecting specific primers for our set of target miRNAs in the database (https://www.thermofisher.com/es/en/home/life-science/pcr/real-time-pcr/real-time-pcr-assays/mirna-ncrna-taqman-assays.html). The plates were incubated in an iQ5 thermocycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA) at 16 °C for 30 min, followed by 42 °C for 30 min and, finally, 85 °C for 5 min. In this step, the cDNA could be stored at −20 °C for one week. We then prepared a mixture containing 10 μL of a PreAmp custom primer pool specific to our set of target miRNAs, 7.5 µL of RT mix and 20 μL of TaqMan PreAmp Master Mix (Life Technologies, Carlsbad, CA, USA) to make a final volume of 40 µL. The mixture was then incubated in a iQ5 Thermocycler under the following conditions: denaturation at 95 °C for 10 min; 55 °C for 2 min, 72 °C for 2 min; 20 cycles of amplification (15 s at 95 °C, 4 min at 60 °C); and finally 99.9 °C for 10 min. The preamplified products were then diluted with RNase-free water at a ratio of 1:40 and used for real-time RT-PCR reactions.
Measuring levels of circulating miRNAs using real-time PCR
The circulating levels of miRNAs were measured using the OpenArray® platform (Life Technologies, Carlsbad, CA, USA) following the manufacturer’s instructions. As a normalization method, we first selected the miRNAs that showed the least variability in their CT values in all samples. For this, we used the NormFinder Bioinformatic tool (MOMA-Department of Molecular Medicine, Aarhus University Hospital, Denmark)38 (software extensively used in expression studies39,40. The application showed that the most stable miRNAs were miR-143 and miR-144. Second, we used the BestKeeper method to calculate the geometric mean of the pair-wise Ct values (Ct values of miR-143 and miR-144)41. The relative expression data were analyzed by OpenArray® Real-Time qPCR Analysis Software and with the relative quantification powered by Thermo Fisher Cloud (Life Technologies, Carlsbad, CA, USA). miRNAs levels are expressed in arbitrary units (AU), calculated with the Ct of each target miRNA with reference to the BestKeeper value.
Relationship of insulin signaling and release indexes with circulating miRNA levels
To analyze the relationship between baseline plasma miRNA levels and both insulin signaling and release indexes, the subjects were categorized by the median of their plasma miRNA levels. Thus, for each miRNA studied, two groups were created: one low-level group and one high-level group. The evolution of signaling and release indexes was evaluated between baseline and a median follow-up of 60 months, according to the levels of miRNAs expression (low or high).
Statistical analysis
The quantitative data were evaluated using the Kolmogorov–Smirnov test to determine whether they followed a normal distribution. The parameters were taken as normally distributed if p > 0.05. A comparative analysis of baseline plasma miRNAs levels between the pre-DM or T2DM groups was carried out using ONE-WAY ANOVA. Statistical significance for non-normal distribution variables was assessed by non-parametric tests using the Mann–Whitney U test. The effect of baseline levels of miRNAs on insulin resistance, insulin secretion and DI after a median follow-up of 60 months was assessed using the Wilcoxon test. Non-T2DM, pre-DM and incident pre-DM subjects were grouped, and the probability of developing T2DM (incident-T2DM) was evaluated by Cox regression analysis to test the potential predictive value of the miRNAs studied. The subjects were categorized into tertiles of plasma levels for each miRNA analyzed. Thus, low (T1), intermediate (T2) and high (T3) plasma levels were the three groups defined. The hazard ratio (HR) in the Cox regression analysis of each miRNA studied was analyzed by comparing T1 vs. T2 and T1 vs. T3. P < 0.05 was considered to be significant. All analyses were adjusted for age, diet, gender, body mass index, glycosylated hemoglobin, HDL-C, triglycerides and waist circumference. Receiver operating characteristic curve (ROC) analysis was performed after including the study variables in a logistic binary regression analysis and with the statistical residuals (probabilities). The statistical analysis was carried out using SPSS (now PASW Statistics for Windows (version 21.0) (IBM. Chicago, Illinois, USA).
Results
Baseline characteristics of patients
In a median follow-up of 60 months, 78 subjects did not develop T2DM or pre-DM (non-T2DM); 239 subjects were pre-DM at baseline, of which 223 subjects maintained pre-DM during the follow-up period (pre-DM); 30 subjects were not pre-DM at baseline but developed pre-DM in the follow-up period (Incident pre-DM). Finally, 107 of the participating subjects developed T2DM (Incident-T2DM) in a median follow-up of 60 months. BMI, waist circumference, total cholesterol, TG, HbA1c, fasting glucose, insulin, glucose after 2 h of OGTT, HOMA-IR and HIRI were all higher in incident-T2DM than in pre-DM, incident pre-DM and non-T2DM subjects (all p < 0.05). In contrast, ISI, IGI, and DI were lower in incident-T2DM subjects than in pre-DM subjects and non-T2DM subjects after a median follow-up of 60 months (Table 1).
Baseline levels of circulating miRNAs in the study
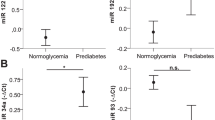
We observed that baseline plasma levels of miR-150 and miR-30a-5p were higher and that miR-15a and miR-375 were lower in incident-T2DM subjects than in non-T2DM subjects, with intermediate values in the pre-DM and incident pre-DM patients (p < 0.001; p < 0.001; p = 0.001; and p = 0.044, respectively). We also observed that the plasma levels of miR-150 were higher in the pre-DM subjects than in non-T2DM subjects (p = 0.020) (Fig. 1).
Values are expressed as the mean ± standard error. Variables were calculated using ONE-WAY ANOVA through SPSS (now PASW Statistics for Windows (version 21.0) (IBM. Chicago, Illinois)). Subjects did not develop T2DM or pre-DM (non-T2DM); subjects maintained pre-DM during the follow-up period (pre-DM); subjects not pre-DM at baseline but developed pre-DM in the follow-up period (Incident pre-DM); and subjects developed T2DM (Incident-T2DM). Significance was assessed by non-parametric tests using the Mann-Whitney U test. * p < 0.05
Relationship between miRNAs levels and insulin signaling and release indexes
We studied the changes according to the baseline miRNAs levels in the insulin signaling and release indexes (DI, ISI, HIRI and MISI) after 4 years of follow-up. The subjects were categorized according to the median basal plasma levels for each of the four miRNAs studied. We observed that DI decreased after 4 years in subjects with high plasma levels of miR-150 and miR-30a-5p (p = 0.047 and p = 0.007, respectively) compared with subjects with low plasma levels of these miRNAs. In contrast, no significant differences were observed in miR-375 and miR-15a (Fig. 2). The DI was lower in patients with high plasma levels of miR-150 compared with that in patients with low plasma levels for this miRNA at 4 years of follow-up (p = 0.029) (Fig. 2).
Values are expressed as the mean ± standard error. Variables were calculated by Wilcoxon test using SPSS (now PASW Statistics for Windows (version 21.0) (IBM. Chicago, Illinois)). The dotted gray line shows subjects with low levels of miRNA at baseline, and the continuous black line shows subjects with high levels of miRNA at baseline. §p < 0.05 baseline vs. 4 y. #p < 0.05 Low expression levels vs. high expression levels groups
Moreover, the ISI and MISI indexes decreased after the follow-up period in patients with high baseline plasma levels of miR-150 (p < 0.001 and p = 0.020, respectively) when compared with the low plasma levels group (Table 2). Likewise, patients with high baseline plasma levels of miR-375, compared to the group with low levels of this miRNA, showed a decrease in the HIRI after the follow-up period (p < 0.001) (Table 2).
Analysis of the probability of developing T2DM based on miRNAs levels using the Cox regression model
The risk of developing T2DM based on plasma miRNAs levels was evaluated by Cox regression analysis. We categorized patients by tertiles of the baseline levels for each miRNA: low levels (T1), intermediate levels (T2), and high levels (T3). Our results revealed that patients with high levels (T3) of miR-150 and miR-30a-5p are at a higher risk of developing T2DM than those with low levels (T1) (miR-150, HR = 4.218; 95% CI: 2.370–7.507 and miR-30a-5p, HR = 2.527; 95% CI: 1.552–4.116). In contrast, patients with low levels (T1) of miR-15a and miR-375 are at a higher risk of developing T2DM than those with high levels (T3) (miR-15a, HR = 3.269; 95% CI: 1.941–5.507 and miR-375, HR = 1.604; 95% CI: 0.957–2.687) (Fig. 3).
Subjects were divided into low (T1), intermediate (T2) and high (T3) baseline levels of miRNAs. The dotted black line represents subjects from T1, the dotted gray line represents subjects from T2, and the continuous black line represents subjects from T3. Analyses were adjusted for age, diet, gender, body mass index, glycosylated hemoglobin, HDL, triglycerides, and waist circumference
Comparison between miRNAs-based model and established biomarkers of diabetic risk
We compared the predictive value of our model to the previously established biomarkers and FINDRISC and ADA scores in order to identify the added predictive power of the miRNAs. We performed a receiver operating characteristic (ROC) curve analysis, combining our model with the previously established biomarkers and scores. We observed an AUC of 0.714 when clinical variables were used (age, gender, BMI, HDL-C, TG, HbA1c, fasting glucose and fasting insulin), an AUC = 0.759 when we added the OGTT-derived index (DI, MISI, ISI, HIRI and IGI), an AUC = 0.793 when the clinical variables were combined with the miRNAs, an AUC of 0.754 when the FINDRISC score was combined with the miRNAs and an AUC of 0.750 when the ADA score was combined with the miRNAs (Fig. 4).
a ROC model including clinical variables (age, gender, BMI, HDL-C, TG, HbA1c, fasting glucose and fasting insulin); (b) ROC model including clinical variables and OGTT-derived indexes (DI, MISI, ISI, HIRI, and IGI); (c) ROC model including clinical variables and studied miRNAs; (d) ROC model including studied miRNAs and FINDRISC score; and (e) ROC model including studied miRNAs and ADA score. The analyses were performed using SPSS (now PASW Statistics for Windows (version 21.0) (IBM. Chicago, Illinois))
Discussion
Our study showed that miR-150 and miR-30a-5p, as well as miR-15a and miR-375, were deregulated in plasma several years before the diagnosis of T2DM. Specifically, we demonstrated for the first time that the baseline plasma levels of miR-150 and miR-30a-5p were higher and miR-15a and miR-375 were lower in incident-T2DM subjects compared with non-T2DM subjects after a median follow-up of 60 months, with intermediate levels in incident pre-DM and pre-DM subjects. Moreover, our study showed that a higher risk of developing T2DM is associated with low baseline plasma levels of miR-15a and miR-375 and high baseline plasma levels of miR-150 and miR-30a-5p. Our study also revealed the potential use of circulating miRNAs-based tools for predicting type 2 diabetes development in clinical practice, showing a small but higher added predictive value to the usual clinical variables used, such as age, gender, BMI, HDL-C, TG, HbA1c, fasting glucose and fasting insulin, than the predictive value added by indices derived from OGTT, such as DI, MISI, ISI, HIRI and IGI, to these variables.
Circulating miRNAs in human biofluids, such as blood42, have led to their use as non-invasive biomarkers for multiple pathologies, including CVD and T2DM43,44,45. This is based on the evidence that miRNAs play a major role in different mechanisms involved in T2DM development, such as insulin production, secretion and action, which supports their use as biomarkers for T2DM diagnosis46. However, the alteration in the miRNA pattern may precede or appear at early stages of diabetes or may be a consequence of the onset of diabetes47.
In this study, we have shown that the plasma levels of four miRNAs are deregulated in prediabetic states, several years preceding the development of T2DM, suggesting their potential use in the development of circulating miRNAs-based tools for T2DM prediction. When comparing the AUC of the ROC curves, we observed an improvement in the predictive capacity of the parameters used in clinical practice when the miRNAs were added to the model. However, this improvement was slightly higher than the improvement observed when OGTT-derived indexes were added to the clinical variables. Nevertheless, these findings suggest a potential use of miRNAs in clinical practice as they can be determined by a single blood collection, whereas OGTT requires continuous blood sampling (every half-hour for 2 h) to calculate the indexes.
In addition, the combined use of the FINDRISC and ADA scores with miRNAs yielded lower predictive power, probably because both scores include parameters not specific for T2DM and/or that they are self-reported, which diminishes their reliability and specificity for the prediction of T2DM status.
In fact, Cox proportional hazards regression analysis showed that patients with high levels of miR-15a and miR-375 are at low risk of developing T2DM (HR 3.269 and 1.604, respectively) whereas patients with high levels of miR-150 and miR-30a-5p are at high risk of developing T2DM (HR 4.218 and 2.527, respectively). The idea of the usefulness of miRNAs plasma levels to assess the risk of T2DM development is supported by other studies, in which alterations in plasma miRNAs levels have been observed before the development of T2DM18,19.
However, whereas the study by Zampetaki et al. is a case-control study in which matched controls were selected, our incidence study included all subjects from the CORDIOPREV cohort without T2DM at baseline, and we used all the non-T2DM patients after the follow-up period as controls.
Whereas our study evaluates the incidence of T2DM, the Zampetaki et al. study compares the differences between groups. The two study designs have their pros and cons: for example, in a case-control study, the recruitment of controls could be prone to selection bias—that is, the controls are systematically different from the population they are meant to represent. However, careful matching, aimed at eliminating any possible confounding factors, leads to efficiency in the study. Otherwise, incidence studies are usually the preferred approach for studying the causes of disease because they use all the available information on the source population over the risk period. However, they are often extremely costly in terms of time and resources, and equivalent results can be achieved more efficiently by using an incidence case–control study design. Although there is a difference in design, both studies demonstrate the potential of miRNAs as biomarkers of T2DM.
The study by Willeit et al. and the present work are incidence studies in which statistical and predictive analysis can be performed to evaluate the risk of T2DM development. Nevertheless, in contrast to the study by Willeit et al.18, which focused on miR-122 to evaluate the risk of T2DM, our study included four miRNAs, which were combined with several clinical variables, such as age, gender, BMI, HDL-C, TG, HbA1c, fasting glucose and fasting insulin; this allowed us to build predictive models by adding miRNAs, which increased the predictive power of the traditional clinical parameters.
In addition, in our study, we followed ADA diagnostic criteria, whereas the studies by Zampetaki et al. and by Willeit et al. followed the World Health Organization guidelines, which do not take into account plasma levels of glycosylated hemoglobin as diagnostic criteria for diabetes, but it was used for the clinical validation of new cases of diabetes. Although these two methods are not very different, when the World Health Organization guidelines are followed, new cases of diabetes diagnosed by glycosylated hemoglobin are not detected.
Overall, miR-150, with a potential role in the main tissues for glucose homeostasis, and miR-15a, involved in insulin production and secretion, showed better risk assessment than miR-30a-5p and miR-375. However, analyses based on indexes of insulin signaling and release have shown that in patients with high circulating levels of miR-150, DI, ISI and MISI decreased during the follow-up period.
In fact, miR-150 is highly expressed in the main tissues for glucose homeostasis, such as adipose, skeletal muscle and liver13. Moreover, this miRNA might come from hematopoietic cells, and the increase in miR-150 could be a secondary phenomenon due to low-grade inflammation. This idea is supported by the involvement of this miRNA in the activation of B cells and other immune cells in adipose tissue, which, in turn, increases the insulin resistance of this tissue37. In addition, our study showed a relationship between miR-150 plasma levels and OGTT-derived indexes, which are related to insulin signaling and release. However, further studies are needed to clarify which tissues are responsible for releasing miR-150 into the blood stream and the mechanisms that regulate this miRNA.
In contrast, miR-15a positively regulates insulin biosynthesis by inhibiting endogenous UCP-2 (uncoupling protein-2) expression, leading to higher ATP levels in islets and improving glucose-stimulated insulin secretion (GSIS). This increase in intracellular ATP closes ATP-sensitive potassium channels, causing plasma membrane depolarization, the influx of Ca2+ and GSIS35,48. Thus, the lower levels of miR-15a that we observed in the patients who develop prediabetes and T2DM suggest that an impairment of stimuli for insulin secretion occurs before the development of T2DM. Moreover, our results are in line with previous studies, which have shown that circulating miR-15a levels decreased significantly before the onset of type 2 diabetes mellitus14,19.
Regarding miRNAs functions, the potential role of miR-150 in insulin resistance in obesity by controlling adipose tissue inflammation seems to have a higher physiological impact than the change in insulin signaling by miR-15a, as suggested by the close relationship found between miR-150 and the DI analyzed in our study. This idea is supported by the fact that the high plasma levels of miR-30a-5p, which have previously been shown to modulate beta cell function, were both associated with a decrease in DI. In fact, the overexpression of miR-30a-5p has been linked to the suppression of BETA2/NeuroD, which plays a key role in the regulation of insulin secretion and pancreatic beta cell dysfunction during glucotoxicity12. Beta2/NeuroD is a transcription factor that binds to the E element of the insulin gene49 and modulates K+ channels to regulate insulin secretion50.
Studies in animal models have shown the relationship of miR-375 with the development of T2DM. In fact, miR375KO mice have a mass reduction in beta cells and an increase in the number of α cells, leading to islet instability34. Moreover, it has also been shown that the overexpression of miR-375 in INS-E cells inhibits insulin gene expression in response to glucose by down-regulating the PDK1 gene, leading also to PI3K pathway inhibition51. In this sense, more research is needed to clarify the relationship between miR-375 and the development of T2DM.
Certain limitations of the current study must be mentioned. The first limitation lies in the fact that we performed a focused bibliographic search of miRNAs associated with insulin sensitivity, insulin secretion and growth and proliferation of beta cells, and other potential T2DM-related miRNAs that have not been described were not included in our study. Another limitation lies in the fact that we analyzed plasma levels of miRNAs, and this approach, while suitable for assessing disease risk, may not accurately reflect the involvement and directionality in the interactions of these miRNAs in processes occurring at cellular levels. Moreover, the prevention of T2DM was not the primary endpoint of the CORDIOPREV trial but was a secondary analysis conducted in the subgroup of cardiovascular patients without T2DM at baseline (CORDIOPREV-DIAB study). The study included a large number of older patients with acute myocardial infarction, which limits our findings to people with these characteristics and precludes generalization to healthy people.
In conclusion, our study showed that deregulated plasma levels of miR-150, miR-30a-5p, miR-15a, and miR-375 were observed years before the onset of T2DM and pre-DM and could be used to evaluate the risk of developing the disease, which may improve prediction and prevention among individuals at high risk of T2DM.
References
United Nations. Political Declaration of the High-Level Meeting of the General Assembly on the Prevention and Control of Noncommunicable Diseases. (Sixty-sixth session of the United Nations General Assembly, New York, 2011).
Prentki, M. & Nolan, C. J. Islet beta cell failure in type 2 diabetes. J. Clin. Invest. 116, 1802–1812 (2006).
Van Ommen, B., van der Greef, J., Ordovas, J. M. & Daniel, H. Phenotypic flexibility as key factor in the human nutrition and health relationship. Genes Nutr. 9, 423–431 (2014).
American Diabetes Association. Classification and diagnosis of diabetes. Diabetes Care 40, S11–S24 (2017).
Kanat, M. et al. Distinct beta-cell defects in mpaired fasting glucose and impaired glucose tolerance. Diabetes 61, 447–453 (2012).
Burant, C. & Young, L. Medical Managements of Type 2 Diabetes Mellitus, 7th edn. (American Diabetes Association, Arlington, 2012).
Giugliano, D., Ceriello, A. & Esposito, K. Glucose metabolism and hyperglycemia. Am. J. Clin. Nutr. 87, 217S–222S (2008).
American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 32(Suppl 1), S62–S67 (2009).
Sebastiani, G. et al. Circulating microRNAs and diabetes mellitus: a novel tool for disease prediction, diagnosis, and staging? J. Endocrinol. Invest. 40, 591–610 (2017).
Seyhan, A. A. et al. Pancreas-enriched miRNAs are altered in the circulation of subjects with diabetes: a pilot cross-sectional study. Sci. Rep. 6, e31479 (2016).
Flynt, A. S. & Lai, E. C. Biological principles of microRNA-mediated regulation: shared themes amid diversity. Nat. Rev. Genet. 9, 831–842 (2008).
Kim, J. W. et al. miRNA-30a-5p-mediated silencing of Beta2/NeuroD expression is an important initial event of glucotoxicity-induced beta cell dysfunction in rodent models. Diabetologia 56, 847–855 (2013).
Karolina, D. S. et al. MicroRNA 144 impairs insulin signaling by inhibiting the expression of insulin receptor substrate 1 in type 2 diabetes mellitus. PLoS One 6, e22839 (2011).
Al-Kafaji, G. et al. Peripheral blood microRNA-15a is a potential biomarker for type 2 diabetes mellitus and pre-diabetes. Mol. Med. Rep. 12, 7485–7490 (2015).
Karolina, D. S. et al. Circulating miRNA profiles in patients with metabolic syndrome. J. Clin. Endocrinol. Metab. 97, E2271–E2276 (2012).
Kong, L. et al. Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: a clinical study. Acta Diabetol. 48, 61–69 (2011).
Chen, W. M. et al. Modulation of microRNA Expression in Subjects with Metabolic Syndrome and Decrease of Cholesterol Efflux from Macrophages via microRNA-33-Mediated Attenuation of ATP-Binding Cassette Transporter A1 Expression by Statins. PLoS. One. 11, e0154672 (2016).
Willeit, P. et al. Circulating MicroRNA-122 Is Associated With the Risk of New-Onset Metabolic Syndrome and Type 2 Diabetes. Diabetes 66, 347–357 (2017).
Zampetaki, A. et al. Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ. Res. 107, 810–817 (2010).
Delgado-Lista, J. et al. CORonary Diet Intervention with Olive oil and cardiovascular PREVention study (the CORDIOPREV study): Rationale, methods, and baseline characteristics: a clinical trial comparing the efficacy of a Mediterranean diet rich in olive oil versus a low-fat diet on cardiovascular disease in coronary patients. Am. Heart J. 177, 42–50 (2016).
American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 34(Suppl 1), S62–S69 (2011).
Blanco-Rojo R. et al. The insulin resistance phenotype (muscle or liver) interacts with the type of diet to determine changes in disposition index after 2 years of intervention: the CORDIOPREV-DIAB randomised clinical trial. Diabetologia https://doi.org/10.1007/s00125-015-3776-4 (2015).
Nathan, D. M. et al. Impaired fasting glucose and impaired glucose tolerance: implications for care. Diabetes Care 30, 753–759 (2007).
Lauritzen, T., Sandbaek, A., Skriver, M. V. & Borch-Johnsen, K. HbA1c and cardiovascular risk score identify people who may benefit from preventive interventions: a 7 year follow-up of a high-risk screening programme for diabetes in primary care (ADDITION), Denmark. Diabetologia 54, 1318–1326 (2011).
Allain, C. C., Poon, L. S., Chan, C. S., Richmond, W. & Fu, P. C. Enzymatic determination of total serum cholesterol. Clin. Chem. 20, 470–475 (1974).
Bucolo, G. & David, H. Quantitative determination of serum triglycerides by the use of enzymes. Clin. Chem. 19, 476–482 (1973).
Briggs, C. J., Anderson, D., Johnson, P. & Deegan, T. Evaluation of the polyethylene glycol precipitation method for the estimation of high-density lipoprotein cholesterol. Ann. Clin. Biochem. 18, 177–181 (1981).
Matsuda, M. & DeFronzo, R. A. Insulin sensitivity indices obtained from oral glucose tolerance testing: comparison with the euglycemic insulin clamp. Diabetes Care 22, 1462–1470 (1999).
Song, Y. et al. Insulin sensitivity and insulin secretion determined by homeostasis model assessment and risk of diabetes in a multiethnic cohort of women: the Women’s Health Initiative Observational Study. Diabetes Care 30, 1747–1752 (2007).
Hanson, R. L. et al. Evaluation of simple indices of insulin sensitivity and insulin secretion for use in epidemiologic studies. Am. J. Epidemiol. 151, 190–198 (2000).
Tang, W. et al. The association between serum uric acid and residual beta -cell function in type 2 diabetes. J. Diabetes Res. 2014, 709691 (2014).
Abdul-Ghani, M. A., Matsuda, M., Balas, B. & DeFronzo, R. A. Muscle and liver insulin resistance indexes derived from the oral glucose tolerance test. Diabetes Care 30, 89–94 (2007).
Plaisance, V., Waeber, G., Regazzi, R. & Abderrahmani, A. Role of microRNAs in islet beta-cell compensation and failure during diabetes. J. Diabetes Res. 2014, 1–12 (2014).
Poy, M. N. et al. miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proc. Natl Acad. Sci. USA 106, 5813–5818 (2009).
Sun, L. L. et al. MicroRNA-15a positively regulates insulin synthesis by inhibiting uncoupling protein-2 expression. Diabetes Res. Clin. Pract. 91, 94–100 (2011).
Xia, H. Q., Pan, Y., Peng, J. & Lu, G. X. Over-expression of miR375 reduces glucose-induced insulin secretion in Nit-1 cells. Mol. Biol. Rep. 38, 3061–3065 (2011).
Ying, W. et al. miR-150 regulates obesity-associated insulin resistance by controlling B cell functions. Sci. Rep. 6, 20176 (2016).
Andersen, C. L., Jensen, J. L. & Orntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Danese, E. et al. Reference miRNAs for colorectal cancer: analysis and verification of current data. Sci. Rep. 7, 8413 (2017).
De Spiegelaere, W. et al. Reference gene validation for RT-qPCR, a note on different available software packages. PLoS ONE 10, e0122515 (2015).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper--Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515 (2004).
Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin. Chem. 56, 1733–1741 (2010).
Guay, C. & Regazzi, R. Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat. Rev. Endocrinol. 9, 513–521 (2013).
Jansen, F. et al. MicroRNA expression in circulating microvesicles predicts cardiovascular events in patients with coronary artery disease. J. Am. Heart Assoc. 3, e001249 (2014).
Kantharidis, P., Wang, B., Carew, R. M. & Lan, H. Y. Diabetes complications: the microRNA perspective. Diabetes 60, 1832–1837 (2011).
Fernandez-Valverde, S. L., Taft, R. J. & Mattick, J. S. MicroRNAs in beta-cell biology, insulin resistance, diabetes and its complications. Diabetes 60, 1825–1831 (2011).
Tang, X., Tang, G. & Ozcan, S. Role of microRNAs in diabetes. Biochim. Biophys. Acta 1779, 697–701 (2008).
Chan, C. B. et al. Increased uncoupling protein-2 levels in beta-cells are associated with impaired glucose-stimulated insulin secretion: mechanism of action. Diabetes 50, 1302–1310 (2001).
Naya, F. J., Stellrecht, C. M. & Tsai, M. J. Tissue-specific regulation of the insulin gene by a novel basic helix-loop-helix transcription factor. Genes Dev. 9, 1009–1019 (1995).
Kim, J. W. et al. Transactivation of the mouse sulfonylurea receptor I gene by BETA2/NeuroD. Mol. Endocrinol. 16, 1097–1107 (2002).
El Ouaamari, A. et al. miR-375 targets 3’-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes 57, 2708–2717 (2008).
Acknowledgements
The CIBEROBN is an initiative of the Instituto de Salud Carlos III, Madrid, Spain. The CORDIOPREV study is supported by the Fundación Patrimonio Comunal Olivarero, Junta de Andalucía (Consejería de Salud, Consejería de Agricultura y Pesca, Consejería de Innovación, Ciencia y Empresa), Diputaciones de Jaén y Córdoba, Centro de Excelencia en Investigación sobre Aceite de Oliva y Salud and Ministerio de Medio Ambiente, Medio Rural y Marino, and Gobierno de España. The study was also partly supported by research grants from the Ministerio de Ciencia e Innovacion (AGL2009-122270 to J.L.-M.); Ministerio de Economía y Competitividad (AGL2012/39615, PIE14/00005, PIE 14/00031 to J.L.-M.; and AGL2015-67896-P to J.L.-M. and A.C.); Consejería de Innovación, Ciencia y Empresa, Proyectos de Investigación de Excelencia, Junta de Andalucía (CVI-7450 to J.L.-M.); and Fondo Europeo de Desarrollo Regional (FEDER). This work was conducted within the project Fatty Acid Metabolism Interlinking Diet with Chronic Disease Risk, funded by research grant from the European Union (PCIN-2016-084 to J.L.-M.) and the U.S. Department of Agriculture-Agricultural Research Service (ARS), under Agreement No. 58-1950-4-003 (O.-J.M.). Antonio Camargo is supported by an ISCIII research contract (Programa Miguel-Servet CP14/00114). We would like to thank the Córdoba node of the Biobank of the Sistema Sanitario Publico de Andalucía (Andalucía, Spain) for providing the human biological samples. We would also like to thank the EASP (Escuela Andaluza de Salud Publica), Granada, Spain, which performed the randomization process for this study.
Author information
Authors and Affiliations
Contributions
The authors’ responsibilities were as follows: J.-L.R., R.-Z.O.A., and L.-M.J. conceived and designed the experiments. A.-D.J.F., D.-L.J., P.-M.P., and L.-M.J. participated in the recruitment and carried out the clinical and nutritional control of the volunteers. J.-L.R., C.-A., and R.-Z.O.A. performed the experiments and collected the data. J.-L.R., C.-A., A.-D.J.F., R.-Z.O.A., and L.-M.J. analyzed and interpreted the data. J.-L.R., C.-A., R.-Z.O.A., and L.-M.J. drafted the manuscript. L.-M.J. conceived and designed the study. v.O.-B., L.-R.M., O.-J.M., and L.-M.J. provided critical revision of the paper for important intellectual content. R.-Z.O.A., D.-L.J., P.-M.P. and L.-M.J. had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. All the authors were involved in writing the paper and gave their final approval to the submitted and published versions. None of the authors have any conflict of interest that could affect the performance of the work or the interpretation of the data.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiménez-Lucena, R., Camargo, A., Alcalá-Diaz, J.F. et al. A plasma circulating miRNAs profile predicts type 2 diabetes mellitus and prediabetes: from the CORDIOPREV study. Exp Mol Med 50, 1–12 (2018). https://doi.org/10.1038/s12276-018-0194-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-018-0194-y
This article is cited by
-
CAPN2 correlates with insulin resistance states in PCOS as evidenced by multi-dataset analysis
Journal of Ovarian Research (2024)
-
Metabolic memory: mechanisms and diseases
Signal Transduction and Targeted Therapy (2024)
-
The link between glycemic control measures and eye microvascular complications in a clinical cohort of type 2 diabetes with microRNA-223-3p signature
Journal of Translational Medicine (2023)
-
Human umbilical cord mesenchymal stromal cell small extracellular vesicle transfer of microRNA-223-3p to lung epithelial cells attenuates inflammation in acute lung injury in mice
Journal of Nanobiotechnology (2023)
-
A comprehensive overview on Micro RNA signature in type 2 diabetes Mellitus and its complications
Indian Journal of Clinical Biochemistry (2023)