Abstract
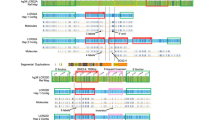
The mechanism of chromosomal rearrangement associated with inverted–duplication–deletion (INV–DUP–DEL) pattern formation has been investigated by many researchers, and several possible mechanisms have been proposed. Currently, fold-back and subsequent dicentric chromosome formation has been established as non-recurrent INV–DUP–DEL pattern formation mechanisms. In the present study, we analyzed the breakpoint junctions of INV–DUP–DEL patterns in five patients using long-read whole-genome sequencing and detected 2.2–6.1 kb copy-neutral regions in all five patients. At the end of the INV–DUP–DEL, two patients exhibited chromosomal translocations, which are recognized as telomere capture, and one patient showed direct telomere healing. The remaining two patients had additional small-sized intrachromosomal segments at the end of the derivative chromosomes. These findings have not been previously reported but they may only be explained by the presence of telomere capture breakage. Further investigations are required to better understand the mechanisms underlying this finding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Shimada S, Shimojima K, Okamoto N, Sangu N, Hirasawa K, Matsuo M, et al. Microarray analysis of 50 patients reveals the critical chromosomal regions responsible for 1p36 deletion syndrome-related complications. Brain Dev. 2015;37:515–26.
Yamamoto-Shimojima K, Kouwaki M, Kawashima Y, Itomi K, Momosaki K, Ozasa S, et al. Natural histories of patients with Wolf-Hirschhorn syndrome derived from variable chromosomal abnormalities. Congenit Anom (Kyoto). 2019;59:169–73.
Ballif BC, Wakui K, Gajecka M, Shaffer LG. Translocation breakpoint mapping and sequence analysis in three monosomy 1p36 subjects with der(1)t(1;1)(p36;q44) suggest mechanisms for telomere capture in stabilizing de novo terminal rearrangements. Hum Genet 2004;114:198–206.
Zuffardi O, Bonaglia M, Ciccone R, Giorda R. Inverted duplications deletions: underdiagnosed rearrangements?? Clin Genet. 2009;75:505–13.
Shimokawa O, Kurosawa K, Ida T, Harada N, Kondoh T, Miyake N, et al. Molecular characterization of inv dup del(8p): analysis of five cases. Am J Med Genet A 2004;128a:133–7.
Vibert R, Mignot C, Keren B, Chantot-Bastaraud S, Portnoï MF, Nouguès MC, et al. Neurodevelopmental phenotype in 36 new patients with 8p inverted duplication-deletion: genotype-phenotype correlation for anomalies of the corpus callosum. Clin Genet. 2022;101:307–16.
Yurchenko DA, Minzhenkova ME, Dadali EL, Markova ZG, Rudenskaya GE, Matyushchenko GN, et al. Clinical manifestations of various molecular cytogenetic variants of eight cases of "8p inverted duplication/deletion syndrome". Biomedicines .2022;10:567.
García-Santiago FA, Martínez-Glez V, Santos F, García-Miñaur S, Mansilla E, Meneses AG, et al. Analysis of invdupdel(8p) rearrangement: Clinical, cytogenetic and molecular characterization. Am J Med Genet A. 2015;167a:1018–25.
Kato T, Inagaki H, Miyai S, Suzuki F, Naru Y, Shinkai Y, et al. The involvement of U-type dicentric chromosomes in the formation of terminal deletions with or without adjacent inverted duplications. Hum Genet. 2020;139:1417–27.
Rowe LR, Lee JY, Rector L, Kaminsky EB, Brothman AR, Martin CL, et al. U-type exchange is the most frequent mechanism for inverted duplication with terminal deletion rearrangements. J Med Genet. 2009;46:694–702.
Yu S, Graf WD. Telomere capture as a frequent mechanism for stabilization of the terminal chromosomal deletion associated with inverted duplication. Cytogenet Genome Res. 2010;129:265–74.
Kibe T, Mori Y, Okanishi T, Shimojima K, Yokochi K, Yamamoto T. Two concurrent chromosomal aberrations involving interstitial deletion in 1q24.2q25.2 and inverted duplication and deletion in 10q26 in a patient with stroke associated with antithrombin deficiency and a patent foramen ovale. Am J Med Genet A. 2011;155a:215–20.
Kaneko S, Shimbo A, Irabu H, Yamamoto T, Shimizu M. Inverted-duplication-deletion of chromosome 10q identified in a patient with systemic lupus erythematosus. Pediatr Int. 2022;65:e15396.
Tamura T, Yamamoto Shimojima K, Okamoto N, Yagasaki H, Morioka I, Kanno H, et al. Long-read sequence analysis for clustered genomic copy number aberrations revealed architectures of intricately intertwined rearrangements. Am J Med Genet A. 2022;191a:112–9.
Yanagishita T, Imaizumi T, Yamamoto-Shimojima K, Yano T, Okamoto N, Nagata S, et al. Breakpoint junction analysis for complex genomic rearrangements with the caldera volcano-like pattern. Hum Mutat. 2020;41:2119–27.
Imaizumi T, Yamamoto-Shimojima K, Yanagishita T, Ondo Y, Nishi E, Okamoto N, et al. Complex chromosomal rearrangements of human chromosome 21 in a patient manifesting clinical features partially overlapped with that of Down syndrome. Hum Genet. 2020;139:1555–63.
Imaizumi T, Yamamoto-Shimojima K, Yanagishita T, Ondo Y, Yamamoto T. Analyses of breakpoint junctions of complex genomic rearrangements comprising multiple consecutive microdeletions by nanopore sequencing. J Hum Genet. 2020;65:735–41.
Clark BE, Shooter C, Smith F, Brawand D, Thein SL. Next-generation sequencing as a tool for breakpoint analysis in rearrangements of the globin gene clusters. Int J Lab Hematol. 2017;39:111–20.
Cer RZ, Donohue DE, Mudunuri US, Temiz NA, Loss MA, Starner NJ, et al. Non-B DB v2.0: a database of predicted non-B DNA-forming motifs and its associated tools. Nucleic Acids Res. 2013;41:D94–d100.
Voet T, Vanneste E, Van der Aa N, Melotte C, Jackmaert S, Vandendael T, et al. Breakage-fusion-bridge cycles leading to inv dup del occur in human cleavage stage embryos. Hum Mutat. 2011;32:783–93.
Hermetz KE, Newman S, Conneely KN, Martin CL, Ballif BC, Shaffer LG, et al. Large inverted duplications in the human genome form via a fold-back mechanism. PLoS Genet. 2014;10:e1004139.
Bonaglia MC, Giorda R, Massagli A, Galluzzi R, Ciccone R, Zuffardi O. A familial inverted duplication/deletion of 2p25.1-25.3 provides new clues on the genesis of inverted duplications. Eur J Hum Genet. 2009;17:179–86.
Weckselblatt B, Rudd MK. Human structural variation: mechanisms of chromosome rearrangements. Trends Genet. 2015;31:587–99.
Lopes M, Foiani M, Sogo JM. Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol Cell. 2006;21:15–27.
Burssed B, Zamariolli M, Bellucco FT, Melaragno MI. Mechanisms of structural chromosomal rearrangement formation. Mol Cytogenet. 2022;15:23.
Burssed B, Zamariolli M, Favilla BP, Meloni VA, Goloni-Bertollo EM, Bellucco FT, et al. Fold-back mechanism originating inv-dup-del rearrangements in chromosomes 13 and 15. Chromosome Res. 2023;31:10.
Schlade-Bartusiak K, Tucker T, Safavi H, Livingston J, van Allen MI, Eydoux P, et al. Independent post-zygotic breaks of a dicentric chromosome result in mosaicism for an inverted duplication deletion 9p and terminal deletion 9p. Eur J Med Genet. 2013;56:229–35.
Pedurupillay CR, Misceo D, Gamage TH, Dissanayake VH, Frengen E. Post-zygotic breakage of a dicentric chromosome results in mosaicism for a telocentric 9p marker chromosome in a boy with developmental delay. Gene 2014;533:403–10.
Kostiner DR, Nguyen H, Cox VA, Cotter PD. Stabilization of a terminal inversion duplication of 8p by telomere capture from 18q. Cytogenet Genome Res. 2002;98:9–12.
Al-Zain A, Nester MR, and Symington LS Double-strand breaks induce inverted duplication chromosome rearrangements by a DNA polymerase δ and Rad51-dependent mechanism. bioRxiv. 2023.
Li BZ, Putnam CD, Kolodner RD. Mechanisms underlying genome instability mediated by formation of foldback inversions in Saccharomyces cerevisiae. Elife .2020;9:e58223.
Buysse K, Antonacci F, Callewaert B, Loeys B, Fränkel U, Siu V, et al. Unusual 8p inverted duplication deletion with telomere capture from 8q. Eur J Med Genet. 2009;52:31–6.
Knijnenburg J, Uytdewilligen MEW, van Hassel D, Oostenbrink R, Eussen BHJ, de Klein A, et al. Postzygotic telomere capture causes segmental UPD, duplication and deletion of chromosome 8p in a patient with intellectual disability and obesity. Eur J Med Genet. 2017;60:445–50.
Milosevic J, El Khattabi L, Roubergue A, Coussement A, Doummar D, Cuisset L, et al. Inverted duplication with deletion: first interstitial case suggesting a novel undescribed mechanism of formation. Am J Med Genet A. 2014;164a:3180–6.
Chabchoub E, Rodríguez L, Galán E, Mansilla E, Martínez-Fernandez ML, Martínez-Frías ML, et al. Molecular characterisation of a mosaicism with a complex chromosome rearrangement: evidence for coincident chromosome healing by telomere capture and neo-telomere formation. J Med Genet. 2007;44:250–6.
Hannes F, Van Houdt J, Quarrell OW, Poot M, Hochstenbach R, Fryns JP, et al. Telomere healing following DNA polymerase arrest-induced breakages is likely the main mechanism generating chromosome 4p terminal deletions. Hum Mutat. 2010;31:1343–51.
Yatsenko SA, Brundage EK, Roney EK, Cheung SW, Chinault AC, Lupski JR. Molecular mechanisms for subtelomeric rearrangements associated with the 9q34.3 microdeletion syndrome. Hum Mol Genet. 2009;18:1924–36.
Kockler ZW, Osia B, Lee R, Musmaker K, Malkova A. Repair of DNA breaks by break-induced replication. Annu Rev Biochem. 2021;90:165–91.
Rossi E, Riegel M, Messa J, Gimelli S, Maraschio P, Ciccone R, et al. Duplications in addition to terminal deletions are present in a proportion of ring chromosomes: clues to the mechanisms of formation. J Med Genet. 2008;45:147–54.
Murmann AE, Conrad DF, Mashek H, Curtis CA, Nicolae RI, Ober C, et al. Inverted duplications on acentric markers: mechanism of formation. Hum Mol Genet. 2009;18:2241–56.
Guilherme RS, Meloni VF, Kim CA, Pellegrino R, Takeno SS, Spinner NB, et al. Mechanisms of ring chromosome formation, ring instability and clinical consequences. BMC Med Genet. 2011;12:171.
Acknowledgements
We would like to express our gratitude to the patients and their families for their cooperation. We also thank M. Kiyooka and W. Chen for the PacBio sequencing and GeneBay, Inc. for the Nanopore PromethION sequencing.
Funding
This study was supported by JSPS KAKENHI, Grant Numbers 21K07873 (JSPS) and 16H06279 (PAGS).
Author information
Authors and Affiliations
Contributions
TY designed the study. KSY organized the study and drafted the manuscript. TT, NO, EN, AN, IT, YS, and MS contributed to acquisition of data. YM and AT analyzed the data. HK reviewed the manuscript critically. All authors contributed to the data interpretation. All authors agree to be accountable for all aspects of the work and ensure that questions related to the accuracy or integrity of any part of the study are appropriately investigated and resolved.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shimojima Yamamoto, K., Tamura, T., Okamoto, N. et al. Identification of small-sized intrachromosomal segments at the ends of INV–DUP–DEL patterns. J Hum Genet 68, 751–757 (2023). https://doi.org/10.1038/s10038-023-01181-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-023-01181-x