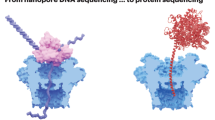
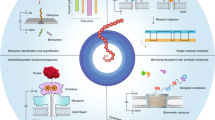
Abstract
Nanopore sequencing is one of the most exciting new technologies that undergo dynamic development. With its development, a growing number of analytical tools are becoming available for researchers. To help them better navigate this ever changing field, we discuss a range of software available to analyze sequences obtained using nanopore technology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Eid J, Fehr A, Gray J, Luong K, Lyle J, Otto G, et al. Real-time DNA sequencing from single polymerase molecules. Science. 2009;323:133–8.
Kasianowicz JJ, Brandin E, Branton D, Deamer DW. Characterization of individual polynucleotide molecules using a membrane channel. Proc Natl Acad Sci. 1996;93:13770–3.
Leggett RM, Clark MD. A world of opportunities with nanopore sequencing. J Exp Bot. 2017;68:5419–29.
Loman NJ, Quinlan AR. Poretools: a toolkit for analyzing nanopore sequence data. Bioinformatics. 2014;30:3399–401.
Rang FJ, Kloosterman WP, de Ridder J. From squiggle to basepair: computational approaches for improving nanopore sequencing read accuracy. Genome Biol. 2018;19:90.
Boza V, Brejova B, Vinar T. DeepNano: deep recurrent neural networks for base calling in MinION nanopore reads. PLoS ONE. 2017;12:e0178751.
Teng HT, Cao MD, Hall MB, Duarte T, Wang S, Coin LJM. Chiron: translating nanopore raw signal directly into nucleotide sequence using deep learning. Gigascience. 2018;7:giy037.
Wick RR, Judd LM, Holt KE. Performance of neural network basecalling tools for Oxford nanopore sequencing. Genome Biology. 2019;20:129.
Simpson JT, Workman RE, Zuzarte PC, David M, Dursi LJ, Timp W. Detecting DNA cytosine methylation using nanopore sequencing. Nat Methods. 2017;14:407.
Needleman SB, Wunsch CD. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol. 1970;48:443–53.
Smith TF, Waterman MS. Identification of common molecular subsequences. J Mol Biol. 1981;147:195–7.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Shang J, Zhu F, Vongsangnak W, Tang Y, Zhang W, Shen B. Evaluation and comparison of multiple aligners for next-generation sequencing data analysis. Biomed Res Int. 2014;2014:309650.
Kielbasa SM, Wan R, Sato K, Horton P, Frith MC. Adaptive seeds tame genomic sequence comparison. Genome Res. 2011;21:487–93.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26:589–95.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–6.
Staden R. A strategy of DNA sequencing employing computer programs. Nucleic Acids Res. 1979;6:2601–10.
Hernandez D, Francois P, Farinelli L, Osteras M, Schrenzel J. De novo bacterial genome sequencing: Millions of very short reads assembled on a desktop computer. Genome Res. 2008;18:802–9.
Simpson JT, Durbin R. Efficient construction of an assembly string graph using the FM-index. Bioinformatics 2010;26:i367–i73.
Gremme G, Steinbiss S, Kurtz S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE ACM Trans Comput Biol Bioinform. 2013;10:645–56.
Pevzner PA, Tang H, Waterman MS. An Eulerian path approach to DNA fragment assembly. Proc Natl Acad Sci USA. 2001;98:9748–53.
Pevzner PA, Tang H, Tesler G. De novo repeat classification and fragment assembly. Genome Res. 2004;14:1786–96.
Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017;27:722–36.
Myers EW, Sutton GG, Delcher AL, Dew IM, Fasulo DP, Flanigan MJ, et al. A whole-genome assembly of Drosophila. Science. 2000;287:2196–204.
Ma ZW, Hu JC. Complete genome sequence of a marine-sediment-derived bacterial strain Bacillus velezensis SH-B74, a cyclic lipopeptides producer and a biopesticide. 3 Biotech. 2019;9:162.
Brejova B, Lichancova H, Brazdovic F, Hegedusova E, Jakubkova MF, Hodorova V, et al. Genome sequence of the opportunistic human pathogen Magnusiomyces capitatus. Curr Genet. 2019;65:539–60.
Karageorgiou C, Gamez-Visairas V, Tarrio R, Rodriguez-Trelles F. Long-read based assembly and synteny analysis of a reference Drosophila subobscura genome reveals signatures of structural evolution driven by inversions recombination-suppression effects. BMC Genomics. 2019;20:223.
Wang MJ, Tu LL, Yuan DJ, Zhu D, Shen C, Li JY, et al. Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nat Genet. 2019;51:224.
Xiao YS, Xiao ZZ, Ma DY, Liu J, Li J. Genome sequence of the barred knifejaw Oplegnathus fasciatus (Temminck & Schlegel, 1844): the first chromosome-level draft genome in the family Oplegnathidae. Gigascience. 2019;8:giz013.
Lin Y, Yuan J, Kolmogorov M, Shen MW, Chaisson M, Pevzner PA. Assembly of long error-prone reads using de Bruijn graphs. P Natl Acad Sci USA. 2016;113:E8396–E405.
Quick J, Loman NJ, Duraffour S, Simpson JT, Severi E, Cowley L, et al. Real-time, portable genome sequencing for Ebola surveillance. Nature. 2016;530:228–32.
Zeng Y, Chen T. DNA methylation reprogramming during mammalian development. Genes. 2019;10:257.
Rand AC, Jain M, Eizenga JM, Musselman-Brown A, Olsen HE, Akeson M, et al. Mapping DNA methylation with high-throughput nanopore sequencing. Nat Methods. 2017;14:411.
Liu Q, Fang L, Yu G, Wang D, Xiao CL, Wang K. Detection of DNA base modifications by deep recurrent neural network on Oxford nanopore sequencing data. Nat Commun. 2019;10:2449.
Tardaguila M, de la Fuente L, Marti C, Pereira C, Pardo-Palacios FJ, del Risco H, et al. SQANTI: extensive characterization of long-read transcript sequences for quality control in full-length transcriptome identification and quantification. Genome Res. 2018;28:396–411.
Tang AD, Soulette CM, Baren MJV, Hart K, Hrabeta-Robinson E, Wu CJ, et al. Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns. bioRxiv. 2018:410183.
Byrne A, Beaudin AE, Olsen HE, Jain M, Cole C, Palmer T, et al. Nanopore long-read RNAseq reveals widespread transcriptional variation among the surface receptors of individual B cells. Nat Commun. 2017;8:16027.
Cook DE, Valle-Inclan JE, Pajoro A, Rovenich H, Thomma BPHJ, Faino L. Long-Read Annotation: Automated Eukaryotic Genome Annotation Based on Long-Read cDNA Sequencing. Plant Physiol. 2019;179:38–54.
Yang C, Chu J, Warren RL, Birol I. NanoSim: nanopore sequence read simulator based on statistical characterization. Gigascience. 2017;6:gix010.
Rodríguez-Pérez H, Hernández-Beeftink T, Lorenzo-Salazar JM, Roda-García JL, Pérez-González CJ, Colebrook M, et al. NanoDJ: a dockerized jupyter notebook for interactive Oxford Nanopore MinION sequence manipulation and genome assembly. BMC Bioinformatics. 2019;20:234.
Mitsuhashi S, Frith MC, Mizuguchi T, Miyatake S, Toyota T, Adachi H, et al. Tandem-genotypes: robust detection of tandem repeat expansions from long DNA reads. Genome Biol. 2019;20:58.
Edwards HS, Krishnakumar R, Sinha A, Bird SW, Patel KD, Bartsch MS. ReAl-time Selective Sequencing with RUBRIC: read until with basecall and reference-informed criteria. BMC Bioinformatics. 2019;20:234.
Shabardina V, Kischka T, Manske F, Grundmann N, Frith MC, Suzuki Y, et al. NanoPipe-a web server for nanopore MinION sequencing data analysis. Gigascience. 2019;8:giy169.
Aristotle. The nicomachean ethics. Oxford; New York: Oxford University Press; 2009. xliii, p. 277.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Makałowski, W., Shabardina, V. Bioinformatics of nanopore sequencing. J Hum Genet 65, 61–67 (2020). https://doi.org/10.1038/s10038-019-0659-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-019-0659-4