Abstract
The focal facial dermal dysplasias (FFDDs) are a group of rare inherited developmental disorders characterized by congenital scar-like atrophic lesions in the bitemporal (FFDD1, 2, and 3) or preauricular (FFDD4) areas. FFDD4 is an autosomal-recessive trait characterized by preauricular skin defects without additional dysmorphic findings. Previously, only two CYP26C1 mutations in four unrelated patients with FFDD4 were reported. Here, we report two additional unrelated FFDD4 patients with four CYP26C1 mutations including three novel lesions: a missense mutation, c.230G>C (p.Arg77Pro), and two splice-site mutations, c.1191+1G>T (IVS5(+1)G>T) and c.1191+2insT (IVS5(+2)insT). In silico analyses predicted all three mutations as pathogenic. Compound heterozygosity was validated through parental studies. These results provide further evidence that CYP26C1 mutations are the molecular genetic basis of FFDD4. Identification of additional cases by dermatologists, pediatricians, and medical geneticists will lead to further understanding of the clinical spectrum of FFDD4 and define its molecular genetic heterogeneity.
Similar content being viewed by others
Introduction
The focal facial dermal dysplasias (FFDDs) are a group of inherited developmental disorders characterized by congenital scar-like atrophic lesions in the bitemporal (FFDD1, FFDD2, and FFDD3) or preauricular (FFDD4) areas [1,2,3,4,5]. FFDD4 (MIM614974) is characterized by preauricular skin defects without additional dysmorphic findings and is inherited as an autosomal-recessive trait [4,5,6]. To date, only ~20 families have been reported worldwide [1, 5,6,7,8,9,10,11].
In FFDD4, the facial skin lesions at birth typically present as multiple, round, or oval-shaped, and vesicular, in the preauricular region along the developmental fusion line of the maxillary and mandibular prominences [5, 10]. These vesicles are filled with fluid for a few years of life and then change into hypopigmented macules, sometimes surrounded by a hyperpigmented rim with fine hairs (hair collar sign). Additional features may include polyps on the buccal mucosa along the same line as the external skin defects. Development and intelligence are usually normal, but neurologic complications have been reported in one patient due to a perinatal intracranial hemorrhage [10].
Recently, our group identified two CYP26C1 mutations in a family with two affected siblings with FFDD4 by exome sequencing [5]. A seven-base-pair (CCATGCA) duplication, c.844_851dup (p.Gln284fs*128, previously erroneously designated as p.Glu284fs*128), was inherited from the mother and a missense mutation, c.1433G>A encoding p.Arg478His, was inherited from the father. Both resulted in loss of CYP26C1 enzyme function when evaluated in an in vitro expression assay [5], establishing the pathogenicity of these mutations. Additional sequencing of four other unrelated patients with FFDD4 revealed that three of the four patients were homozygous for the mutation encoding p.Gln284fs*128.
Here, we describe two unrelated Caucasian families with FFDD4. The probands were screened for CYP26C1 mutations, and compound heterozygosity for three novel CYP26C1 mutations was identified, thereby extending the molecular genetic heterogeneity of CYP26C1 mutations causing FFDD4.
Materials and methods
Patients
The two unrelated probands and their family members were examined and their clinical findings were recorded. This study was approved by Pellegrin-Enfants, Bordeaux, France, and written informed consent was obtained for each subject. This study was also approved by the Institutional Review Board of the Icahn School of Medicine at Mount Sinai, New York, NY.
Molecular and structural analysis
Genomic DNA was isolated from venous blood obtained from each subject. The six exons and their intronic boundaries of the CYP26C1 gene (NM_183374.2) were sequenced as previously described [5].
Results
Clinical description
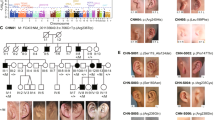
Patient 1 was a male, first evaluated at 3 months of age for bilateral abnormal facial skin lesions (Figs. 1a, b). The unrelated parents had normal facial characteristics and the family history was negative for facial skin findings. The parents also had an unaffected 4-year-old son. The patient was born at term after an uneventful pregnancy, labor, and vaginal delivery. Birth weight was 3.2 kg (25th–50th percentile), height was 50 cm (25th–50th percentile), and head circumference was 35 cm (50th–75th percentile). On dermatologic examination, he had three round skin lesions measuring ~7 mm on both cheeks in a preauricular distribution. Areas of cutis aplasia were present on both cheeks and were bordered by a rim with thin hairs (Figs. 1a, b). No lesions were present in the temporal regions or on his buccal mucosa. His psychomotor development was normal for age. The remainder of his physical examination was unremarkable.
Patient 2 was a female, first seen at 24 months of age for bilateral facial skin abnormalities (Figs. 1c, d). The unrelated parents had normal facial appearances with no lesions, and the family history was negative for abnormal facial dermatologic findings. The parents had an unaffected 4-year-old daughter. Patient 2 was born at term after an uneventful pregnancy, labor, and delivery. She walked at 1 year of age and her psychomotor development was normal. On examination, she had three round skin lesions measuring ~8 mm on the right cheek and two on the left cheek, both in a preauricular distribution. She had a right preauricular tag and fistula associated with an area of long thin hair. The left lesions were atrophic and pale (Fig. 1d). The remainder of her physical examination was unremarkable.
Molecular genetic studies
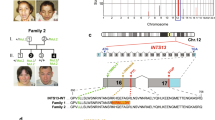
In Patient 1, sequencing CYP26C1 identified two novel mutations: a missense, c.230G>C encoding p.Arg77Pro, and an insertion, c.1191+2insT (IVS5+2insT), located at the consensus splice site of the exon 5/intron 5 junction. The c.230G>C mutation was inherited from his father, whereas the c.1191+2insT lesion was from his mother (Fig. 2a).
Partial sequences of CYP26C1 in Patients 1 and 2. a Patient 1 had a novel missense mutation, c.230G>C, encoding p.Arg77Pro and a novel insertion mutation, c.1191+2insT (IVS5+2insT). The mutation encoding the missense mutation, p.Arg77Pro, was inherited from the patient’s father and the novel consensus splice-site mutation was inherited from his mother. b Patient 2 had the previously reported duplication mutation, c.845_851dup encoding p.Gln284fs*128, and a novel consensus splice-site mutation, c.1191+1G>T (IVS5+1G>T). The mutation encoding the duplication mutation was inherited from the patient’s father and the novel consensus splice-site mutation was inherited from her mother
In Patient 2, sequencing CYP26C1 revealed compound heterozygosity for the previously reported seven-base-pair duplication, c.845_851dup encoding p.Gln284fs*128 [5], and a novel consensus splice-site mutation, c.1191+1G>T (IVS5+1G>T), located at the donor splice site of the exon 5/intron 5 junction (Fig. 2b). The c.845_851dup was inherited from her father and c.1191+1G>T was from her mother. Her unaffected sibling was heterozygous for c.845_851dup.
Discussion
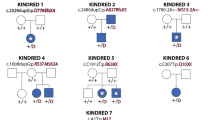
The FFDDs are a group of inherited developmental facial defects that occur at the site of the dermal fusion along the frontonasal and maxillary prominences or along the maxillary and mandibular prominences. The former defects are seen in FFDD1, FFDD2, and FFDD3, and the latter occur in FFDD4 [1,2,3,4]. FFDD1 (MIM136500) is characterized by bitemporal lesions without other dysmorphic features and is a clear autosomal dominant trait. FFDD2 (MIM614973) has the characteristic bitemporal lesions as well as other facial and systemic abnormalities and is inherited as an autosomal dominant trait with variable expressivity and lack of penetrance. To date, the genes responsible for FFDD1 and FFDD2 have not been identified. FFDD3 (MIM227260) is characterized by bitemporal lesions and other characteristic dysmorphic facial features. Some FFDD3 patients have recessive mutations in the TWIST2 gene, which encodes a basic helix-loop-helix transcription factor involved in craniofacial dermal and bone development [3, 4, 12, 13], whereas some have a genomic duplication at 1p36.22p36.21 and normal TWIST2 alleles [14, 15].
FFDD4 is inherited as an autosomal recessive trait in most patients. Previously, only two CYP26C1 mutations, c.844_851dup encoding p.Gln284fs*128 and c.1433G>A encoding p.Arg478His, were identified in four of five unrelated probands with FFDD4 [5]. There have been no other confirmatory reports documenting the CYP26C1 gene as the genetic defect in FFDD4. Here, we report two additional unrelated patients with three novel CYP26C1 mutations: one missense mutation, c.230G>C encoding p.Arg77Pro, and two consensus splice-site mutations, c.1191+1G>T (IVS5+1G>T) and c.1191+2insT (IVS5+2insT). Notably, of the seven unrelated FFDD4 probands who were CYP26C1-genotyped (including the two from this report), three were heterozygotes and three were homozygotes for the p.Gln284fs*128 mutation, which occurs in Caucasians, according to the gnomAD database (http://gnomad.broadinstitute.org) [16], at a minor allele frequency of 0.0037, so homozygosity occurs in ~1 in 286,000 Caucasians. Mutations c.230G>C and c.1191+2insT have not been found in over 138,000 individuals in the gnomAD. However, c.1191+1G>T has been reported with an allele frequency of 0.0002 in the Caucasian population. The CYP26C1 mutation encoding p.Arg77Pro is located in exon 2 and is predicted as pathogenic by in silico programs Polyphen-2 (http://genetics.bwh.harvard.edu/pph2/) [17] and MutationTester (http://www.mutationtaster.org) [18]. The two splice-site mutations are located at the donor consensus site of intron 5 and predicted to abolish the donor site by the in silico splicing program, Neural Network (http://www.fruitfly.org/seq_tools/splice.html) [19], resulting in defective CYP26C1 pre-mRNA processing and non-functional CYP26C1 protein products. Leukocyte RNAs to confirm the aberrant splicing of exon 5 were not available.
CYP26C1 maps to human chromosome 10q23.33. In mouse embryos, CYP26C1 was found to be expressed along the fusion line for the maxillary and mandibular prominences [20], the site of the preauricular skin lesions in FFDD4 patients. CYP26C1 belongs to a family of three mammalian P450 enzymes involved in retinoic acid metabolism (Fig. 3). Although the precise mechanisms responsible for the atrophic skin lesions caused by the CYP26C1 mutations remain to be elucidated, altered retinoic acid metabolism during embryogenesis is presumably the causative factor.
In conclusion, this current study provides further evidence that CYP26C1 mutations are responsible for FFDD4. Although FFDD4 is rare, it has a distinct phenotype that should be familiar to clinicians. The identification of additional cases will likely expand the clinical spectrum and the molecular genetic heterogeneity of FFDD4, as one patient with FFDD4 had screened negative for a CYP26C1 mutation [5]. It is likely that other genes involved in the embryonic fusion of the mandibular and maxillary prominences will be identified.
References
Lee BH, Aggarwal A, Slavotinek AM, Edelmann L, Chen B, Desnick RJ. The focal facial dermal dysplasias: phenotypic spectrum and molecular genetic heterogeneity. J Med Genet. 2017;54:585–90.
Slavotinek AM, Lee BH, Desnick RJ. Focal facial dermal dysplasias. epstein’s inborn errors of development: the molecular basis of clinical disorder of morphogenesis. New York, NY, USA: Oxford University Press; 2016. doi: 10 1093/med/97801999345220030067.
Tukel T, Sosic D, Al-Gazali LI, Erazo M, Casasnovas J, Franco HL, et al. Homozygous nonsense mutations in TWIST2 cause Setleis syndrome. Am J Hum Genet 2010;87:289–96.
Cervantes-Barragan DE, Villarroel CE, Medrano-Hernandez A, Duran-McKinster C, Bosch-Canto V, Del-Castillo V, et al. Setleis syndrome in Mexican-Nahua sibs due to a homozygous TWIST2 frameshift mutation and partial expression in heterozygotes: review of the focal facial dermal dysplasias and subtype reclassification. J Med Genet 2011;48:716–20.
Slavotinek AM, Mehrotra P, Nazarenko I, Tang PL, Lao R, Cameron D, et al. Focal facial dermal dysplasia, type IV, is caused by mutations in CYP26C1. Hum Mol Genet 2013;22:696–703.
Kowalski DC, Fenske NA. The focal facial dermal dysplasias: report of a kindred and a proposed new classification. J Am Acad Dermatol 1992;27:575–82.
Drolet BA, Baselga E, Gosain AK, Levy ML, Esterly NB. Preauricular skin defects. A consequence of a persistent ectodermal groove. Arch Dermatol 1997;133:1551–4.
Stone N, Burge S. Focal facial dermal dysplasia with a hair collar. Br J Dermatol. 1998;139:1136–7.
Wells JM, Weedon D. Focal facial dermal dysplasia or aplasia cutis congenita: a case with a hair collar. Australas J Dermatol 2001;42:129–31.
Prescott T, Devriendt K, Hamel B, Pasch MC, Peeters H, Vander Poorten V, et al. Focal preauricular dermal dysplasia: distinctive congenital lesions with a bilateral and symmetric distribution. Eur J Med Genet 2006;49:135–9.
Krathen MS, Rosenbach M, Yan AC, Crawford GH. Focal preauricular dermal dysplasia: report of two cases and a review of literature. Pediatr Dermatol 2008;25:344–8.
Rosti RO, Uyguner ZO, Nazarenko I, Bekerecioglu M, Cadilla CL, Ozgur H, et al. Setleis syndrome: clinical, molecular and structural studies of the first TWIST2 missense mutation. Clin Genet 2015;88:489–93.
Girisha KM, Bidchol AM, Sarpangala MK, Satyamoorthy K. A novel frameshift mutation in TWIST2 gene causing Setleis syndrome. Indian J Pediatr 2014;81:302–4.
Weaver DD, Norby AR, Rosenfeld JA, Proud VK, Spangler BE, Ming JE, et al. Chromosome 1p36.22p36.21 duplications/triplication causes Setleis syndrome (focal facial dermal dysplasia type III). Am J Med Genet A 2015;167A:1061–70.
Lee BH, Kasparis C, Chen B, Mei H, Edelmann L, Moss C, et al. Setleis syndrome due to inheritance of the 1p36.22p36.21 duplication: evidence for lack of penetrance. J Hum Genet 2015;60:717–22.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016;536:285–91.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods 2010;7:248–9.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 2014;11:361–2.
Reese MG, Eeckman FH, Kulp D, Haussler D. Improved splice site detection in Genie. J Comput Biol 1997;4:311–23.
Tahayato A, Dolle P, Petkovich M. Cyp26C1 encodes a novel retinoic acid-metabolizing enzyme expressed in the hindbrain, inner ear, first branchial arch and tooth buds during murine development. Gene Express. Patterns 2003;3:449–54.
Acknowledgments
We thank the family members for their participation in this study, which was supported in part by a grant from the National Research Foundation of Korea, funded by the Ministry of Education, Science, and Technology (NRF-2016M3A9B4915706).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Lee, B., Morice-Picard, F., Boralevi, F. et al. Focal facial dermal dysplasia type 4: identification of novel CYP26C1 mutations in unrelated patients. J Hum Genet 63, 257–261 (2018). https://doi.org/10.1038/s10038-017-0375-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0375-x