Abstract
Our recent determination of a microRNA (miRNA) expression signature in prostate cancer (PCa) revealed that miR-205-5p was significantly reduced in PCa tissues and that it acted as an antitumor miRNA. The aim of this study was to identify oncogenic genes and pathways in PCa cells that were regulated by antitumor miR-205-5p. Genome-wide gene expression analyses and in silico miRNA database searches showed that 37 genes were putative targets of miR-205-5p regulation. Among those genes, elevated expression levels of seven in particular (HMGB3, SPARC, MKI67, CENPF, CDK1, RHOU, and POLR2D) were associated with a shorter disease-free survival in a large number of patients in the The Cancer Genome Atlas (TCGA) database. We focused on high-mobility group box 3 (HMGB3) because it was the most downregulated by ectopic expression of miR-205-5p in PC3 cells and its expression was involved in PCa pathogenesis. Luciferase reporter assays showed that HMGB3 was directly regulated by miR-205-5p in PCa cells. Knockdown studies using si-HMGB3 showed that expression of HMGB3 enhanced PCa cell aggressiveness. Overexpression of HMGB3/HMGB3 was confirmed in naive PCa and castration-resistant PCa (CRPC) clinical specimens. Novel approaches to analysis of antitumor miRNA-regulated RNA networks in PCa cells may provide new insights into the pathogenic mechanisms of the disease.
Similar content being viewed by others
Introduction
Prostate cancer (PCa) is the most frequently diagnosed cancer in developed countries [1]. Most patients with naive PCa initially respond to androgen deprivation therapy; however, most patients eventually progress to treatment resistance, and distant metastasis, a state termed “castration-resistant prostate cancer” (CRPC) [2]. Currently approved androgen receptor (AR)-targeted agents have resulted in survival benefits in patients with CRPC; however, CRPC is not considered curative [3, 4]. One of the main challenges of CRPC treatment is controlling distant metastases [5, 6]. Therefore, for the control of CRPC, it is indispensable to elucidate the molecular mechanisms of PCa cell metastasis, preferably using genomic approaches.
MicroRNAs (miRNAs) consist of small-sized (19 to 22 nucleotides) noncoding RNAs. They act to fine-tune the expression of protein coding RNAs and noncoding RNAs by repressing translation or cleaving RNA transcripts in a sequence-dependent manner [7]. A large body of evidence indicates that aberrantly expressed miRNAs induce the breakdown of RNA networks in cells, leading to human diseases, including various cancers [8]. In cancer cells, overexpressed miRNAs can promote cancer through their targeting of tumor-suppressive genes. In contrast, downregulated antitumor miRNAs can lead to overexpression or activation of oncogenes. Discovery of microRNA (miRNA)-regulated genes improves our understanding RNA networks in cancer cells [9,10,11,12,13,14,15].
We have undertaken the identification of antitumor miRNAs, with the goal of determining how these miRNAs mediate gene expression networks in PCa cells [16,17,18]. The starting point of our strategy of miRNA studies is to identify aberrantly expressed miRNAs based on miRNA expression signatures [17,18,19,20,21]. Analyses of our miRNA signatures of PCa and CRPC have revealed that miR-205-5p is downregulated in cancer tissues [16,17,18, 22]. Our past study showed that ectopic expression of miR-205-5p significantly suppressed cancer cell migration and invasion in PCa cells [22]. We hypothesized that re-analysis of the target genes regulated by antitumor miR-205-5p might reveal more about PCa pathogenesis. In this study, we aimed to identify therapeutic targets of PCa through determining the gene targets of miR-205-5p regulation. Our present data showed that a total of seven genes (HMGB3, SPARC, MKI67, CENPF, CDK1, RHOU, and POLR2D) were involved in PCa pathogenesis based on TCGA database analyses. Moreover, we focused on the high-mobility group box 3 (HMGB3) gene and investigated its functional significance in PCa and CRPC.
Materials and methods
Clinical prostate specimens, cell lines, and cell culture
In this study, expression analyses of miR-205-5p and HMGB3 were performed using a total of 27 needle biopsy specimens and metastatic CRPC specimens (17 cancer samples and 10 non-cancer tissues). Immunochemical staining of HMGB3 was performed using a total of six specimens (three radical prostatectomy specimens and three metastatic CRPC specimens). These specimens were obtained from patients admitted to Chiba University Hospital and Teikyo University Chiba Medical Center Hospital from 2008 to 2013. Clinical features of patients are summarized in Supplemental Table 1. All patients in this study provided informed consent, and the study protocol was approved by the Institutional Review Board of Chiba University and Teikyo University Chiba Medical Center.
We also used human prostate cancer cells PC3 and DU145 obtained from the American Type Culture Collection (Manassas, VA, USA), as described previously [19,20,21,22,23].
Mature miRNA and small-interfering RNA (siRNA) transfection into PCa cell
We used the following RNA species in this study: mature miRNAs, Pre-miR miRNA Precursors (hsa-miR-205-5p, Product ID: PM11015; Applied Biosystems, Foster City, CA, USA), negative control miRNA (assay ID: AM17111; Applied Biosystems), and siRNA (SR302142; OriGene, Rockville, USA). The transfection procedures were described previously [19,20,21,22,23].
Cell proliferation, migration, and invasion assays
Cell proliferation, migration, and invasion assays were described previously [19,20,21,22,23].
Quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR)
The procedure for PCR quantification was described previously [19,20,21,22,23]. TaqMan probes and primers for HMGB3 ((P/N: Hs00801334_m1; Applied Biosystems) were assay-on-demand gene expression products. Stem-loop RT-PCR for miR-205-5p ((Assay ID: 000509; Applied Biosystems) was used to quantify the expression levels of miRNAs according to the manufacturer’s protocol. To normalize the data for quantification of mRNA and miRNAs, we used human GUSB (P/N: Hs99999908_m1; Applied Biosystems), GAPDH (P/N: Hs02758991_m1; Applied Biosystems) and RNU48 (assay ID: 001006; Applied Biosystems).
Western blot analysis
Immunoblotting was performed with monoclonal anti-HMGB3 antibodies (1:250 dilution; HPA062583; Sigma-Aldrich, St. Louis, MO, USA). Anti-glyceraldehyde 3-phosphate dehydrogenase (GAPDH) antibodies (1:10,000 dilution; ab8245; Abcam, Cambridge, UK) were used as an internal control. The procedures were described in our previous studies [19,20,21,22,23].
Genome-wide gene expression and in silico analysis for the identification of genes regulated by miR-205-5p
We searched putative genes regulated by miR-205-5p by a combination of in silico and genome-wide gene expression analyses. Genes regulated by miR-205-5p were listed using the TargetScan database (release 7.0). Oligo microarrays (Human GE 60 K; Agilent Technologies) were used for gene expression analyses. The microarray data were deposited into GEO (http://www.ncbi.nlm.nih.gov/geo/), with accession number GSE66498. Upregulated genes in PCa were obtained from publicly available data sets in GEO (accession number: GSE29079).
TCGA-PCa data analysis
To explore the clinical significance of miR-205-5p target genes and HMGB3-mediated genes, we used OncoLnc database (http://www.oncolnc.org/) and the RNA sequencing database in TCGA (The Cancer Genome Atlas: https://tcga-data.nci.nih.gov/tcga/) [24]. Gene expression and clinical data were retrieved from cBioportal (http://www.cbioportal.org/, the provisional data downloaded on 27 August 2017) [25, 26].
Regulation of targets downstream of HMGB3 in PCa
We investigated pathways regulated by HMGB3 in PCa cells. We analyzed gene expression using si-HMGB3-transfected PC3 cells. Microarray were used for expression profiling of si-HMGB3 transfectants. The microarray data were deposited into GEO (accession number: GSE66498). To identify signaling pathways regulated in silico, gene expression data were analyzed using the KEGG pathway categories with the GeneCodis program.
Plasmid construction and dual-luciferase reporter assay
The partial wild-type sequences of the HMGB3 3'-untranslated region (UTR) or those with deleted miR-205-5p target sites (position 922-999 or position 1747-1754) were inserted in the 3′-UTR of the hRluc gene in the psiCHECK-2 vector (C8021; Promega, Madison, WI, USA). The procedure for dual-luciferase reporter assay was described previously [19,20,21,22,23].
Immunohistochemistry
Formalin-fixed, paraffin-embedded (FFPE) tissues were used. Tissue sections were incubated overnight at 4 °C with anti-HMGB3 antibodies diluted 1:1000 (HPA062583; Sigma-Aldrich). The procedure for immunohistochemistry was described previously [19,20,21,22,23].
Statistical analysis
Relationships between two or three variables and numerical values were analyzed using Mann–Whitney U-tests or Bonferroni-adjusted Mann–Whitney U-tests. Spearman’s rank tests were used to evaluate the correlations between the expression of miR-205-5p and target genes. Expert StatView software (version 5.0, SAS Institute Inc., Cary, NC, USA) was used for these analyses.
Results
Identification of putative targets of miR-205-5p regulation in PCa cells
Our previous study showed that miR-205-5p was significantly downregulated in naive PCa clinical specimens and ectopic expression of the miRNA inhibited cancer cell migration and invasion [22]. This finding indicated that miR-205-5p acted as an antitumor miRNA in PCa cells.
Extending that study, we performed in silico and gene expression analyses to identify those genes that were targeted by miR-205-5p for regulation. Our strategy is shown in Supplemental Fig. 1. First, we selected putative miR-205-5p target genes using the TargetScan database and identified 4413 genes (release 7.0). Next, we performed comprehensive gene expression analysis using miR-205-5p transfectants of PC3 cells, with negative control miRNA transfectants serving as controls (GEO accession number: GSE66498). A total of 178 genes were downregulated (Log2 ratio < −1.5) in miR-205-5p transfectants. The gene set was then analyzed with a publicly available gene expression data set in GEO (accession number: GSE29079), and genes upregulated in PCa clinical specimens were chosen (fold change > 1.0).
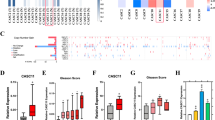
A total of 37 genes were identified as candidate targets of miR-205-5p regulation. We investigated the expression status of these genes and PCa clinicopathological features by using TCGA database (Table 1). Among 37 putative targets, high expression of HMGB3, SPARC, MKI67, CENPF, CDK1, RHOU, and POLR2D was significantly associated with patients’ shorter disease-free survival compared to those with low expression (Fig. 1).
In this study, we focused on HMGB3 because this gene was the most downregulated gene in miR-205-5p transfectants and was associated with a significant difference in disease-free survival. We investigated the functional significance of the gene in PCa cells.
Expression levels of miR-205-5p and HMGB3 in PCa clinical specimens and cell lines
We validated the expression levels of miR-205-5p in PCa clinical specimens and cell lines. Figure 2a shows that the expression levels of miR-205-5p were significantly reduced in cancer tissues compared with noncancerous epithelium (P = 0.0068). Next, we used qRT-PCR to investigate the mRNA expression levels of HMGB3 in 27 clinical specimens. HMGB3 was significantly upregulated in PCa tissues compared to non-PCa tissues (Fig. 2b, P = 0.0031). Spearman’s rank test showed a negative correlation between the expression of HMGB3 and miR-205-5p (P = 0.0043, R = −0.56, Fig. 2c).
Expression of HMGB3 in naive PCa and CRPC clinical specimens. a Expression of miR-205-5p in naive PCa and CRPC clinical specimens. RNU48 was used as an internal control. b Expression levels of mRNA of HMGB3 in PCa clinical specimens. GUSB was used as an internal control. c Spearman’s rank test showed a negative correlation between HMGB3 expression and miR-205-5p (R = −0.56 and P = 0.0043). d, e Immunostaining showed that HMGB3 was strongly expressed in PCa and metastatic castration-resistant prostate cancer (mCRPC) lesions (100 × and 400 × magnification field)
We also examined the expression levels of HMGB3 in naive PCa and metastatic castration-resistant prostate cancer (mCRPC) clinical specimens by immunostaining. HMGB3 was strongly expressed in cancer lesions of PCa and mCRPC (Fig. 2d, e; the patients’ backgrounds are summarized in Supplemental Table 2).
Direct regulation of HMGB3 by miR-205-5p in PCa cells
Next, we investigated whether the expression of HMGB3 decreased in miR-205-5p-transfected PCa cells. Figure 3a shows that HMGB3 mRNA levels were significantly reduced by miR-205-5p transfection compared with the mock or miR-control transfectants in both PC3 and DU145 cells. Furthermore, HMGB3 protein levels were also reduced by miR-205-5p transfection compared with mock or miR-control transfectants (Fig. 3b).
Regulation of HMGB3 expression by miR-205-5p in PCa cells. a Expression levels of HMGB3 mRNA 48 h after transfection with 10 nM miR-205-5p into PCa cell lines. GUSB was used as an internal control. *P < 0.0001. b Protein expression of HMGB3 72 h after transfection with miR-205-5p. GAPDH was used as a loading control. c Diagrammatic presentation of two putative miR-205-5p-binding sites in the 3′-UTR of HMGB3 mRNA. Dual-luciferase reporter assays using vectors encoding putative miR-205-5p target sites (positions 992–999 or 1747–1754) in the HMGB3 3′-UTR for both wild-type and deleted regions. Normalized data were calculated as the ratio of Renilla/firefly luciferase activities. *P < 0.001
We then carried out luciferase reporter assays with a vector that included the 3′-UTR of HMGB3 to confirm that miR-205-5p directly regulated HMGB3 in a sequence-dependent manner. The TargetScan Human database predicted that there were two binding sites for miR-205-5p in the 3′-UTR of HMGB3 (positions 992–999 and 1747–1754). Cotransfection with miR-205-5p and vectors significantly reduced luciferase activity in comparison with those in mock and miR-control transfectants (Fig. 3c).
Effects of silencing HMGB3 on cell proliferation, migration, and invasion in PCa cell lines
A loss-of-function assay using siRNA was performed to examine the function of HMGB3 in 2 PCa cell lines. The HMGB3 expression levels of mRNA and protein were significantly suppressed by si-HMGB3 in PCa cell lines (Fig. 4a, b).
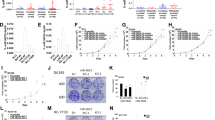
Effects of si-HMGB3 transfection on PCa cell lines. a HMGB3 mRNA expression 72 h after transfection with 10 nM si-HMGB3 into PCa cell lines. GUSB was used as an internal control. b HMGB3 protein expression 72 h after transfection with si-HMGB3. GAPDH was used as a loading control. c Cell proliferation was determined with XTT assays 72 h after transfection with 10 nM si-HMGB3-1 or si-HMGB3-2. d Cell migration activity was determined by migration assays. e Cell invasion activity was determined using Matrigel invasion assays. *P < 0.001; **P < 0.0001
Furthermore, we investigated the effects of HMGB3 knockdown on cell proliferation, migration, and invasion in PCa cell lines. Cancer cell proliferation was significantly reduced in si-HMGB3 transfectants in comparison with that in mock- or miR control-transfected cell lines (Fig. 4c). Next, migration activities were significantly suppressed in si-HMGB3 transfectants in comparison with that in mock- or miR control-transfected cell lines (Fig. 4d). Furthermore, invasion activity was also significantly inhibited in si-HMGB3 transfectants in comparison with that in mock- or miR control-transfected cell lines (Fig. 4e).
TCGA-PCa data analysis
We examined the relationship between stage and pathological diagnosis and expression level of HMGB3 in PCa using the TCGA database. As shown in Supplemental Fig. 2, high expression of HMGB3 was significantly associated with high T stage, positive lymph node metastasis and high Gleason score.
Downstream genes affected by silencing of HMGB3 in PC3 cells
Finally, we performed genome-wide gene expression analysis using si-HMGB3 in PC3 cells to investigate which genes were modulated by HMGB3 signaling. We submitted the raw data to the GEO database (accession number: GSE66498). In this study, we focused on genes that were significantly downregulated by both si-HMGB3-1 and si-HMGB3-2 transfection (Log2 [si-HMGB3/mock] < −0.5). HMGB3 was the most significantly downregulated gene, indicating that the array data were worthy of evaluation. Genes significantly downregulated by silencing of HMGB3 are listed in Table 2. We also listed the upregulated genes by si-HMGB3 in PC3 cells (Supplemental Table 3). These genes are candidate of tumor suppressors in PCa cells and future functional analysis is necessary.
Discussion
An effective strategy for the discovery of novel regulatory pathways in cancer is to analyze antitumor miRNAs and the genes they modulate. Based on this strategy, we have addressed antitumor miRNAs and associated cancer pathways in naive PCa and CRPC cells [16, 17]. For example, miR-26a, miR-26b, miR-218, the miR-29-family, and miR-223 were downregulated in naive PCa tissues and these miRNAs inhibited cancer cell migration and invasion through targeting of genes involved in the extracellular matrix [27,28,29,30]. More recently, we showed that passenger strands of miRNAs, e.g., miR-150-3p and miR-145-3p acted as antitumor miRNAs in naive PCa and CRPC [18, 31]. Interestingly, genes targeted by these miRNA passenger strands (SPOCK1, MELK, NCAPG, BUB1, and CDK1) were overexpressed in naive PCa and CRPC specimens and high expression of these genes predicted poor survival in patients with PCa [18].
Analyses of our signatures revealed that miR-205-5p was significantly downregulated in naive PCa and CRPC specimens [16, 17]. Our previous study showed that restoration of miR-205-5p inhibited cancer cell aggressiveness through its targeting of centromere protein F (CENPF). Overexpression of CENPF was observed in naive PCa specimens and expression of CENPF acted as an oncogenic gene in PCa cells [22]. Initially, overexpression of miR-205-5p was observed in endoderm and ectoderm in embryonic germ cells and its function was involved in the differentiation and maintenance of epithelium [32]. Expression of miR-205-5p might play a pivotal role in the prevention of the epithelial-mesenchymal transition (EMT) process in epithelial cells. There is accumulating evidence that downregulation of miR-205-5p occurs in several cancers, e.g., breast, lung, esophageal and prostate cancers, and that it acts as an antitumor miRNA [33].
Downregulation of miR-205-5p in PCa tissues and its antitumor roles in PCa cells have been observed in past studies, including data from our laboratory [22, 34]. Androgen signaling through AR is an essential pathway in PCa progression and aggressiveness [35]. AR was directly regulated by miR-205-5p and overexpression of miR-205-5p was inversely correlated with the occurrence of metastases and shortened overall survival of patients [36]. These findings indicate that downregulation of miR-205-5p is significantly involved in naive PCa and CRPC pathogenesis. Discovering novel molecular pathways mediated by miR-205-5p should provide new approaches to treating naive PCa and CRPC.
In our present study, a total of 37 putative miR-205-5p targets were identified. Moreover, analysis of a large number of patients in the TCGA database demonstrated that patients with high expression of HMGB3, SPARC, MKI67, CENPF, CDK1, RHOU, and POLR2D had significantly shorter disease-free survival than did those with low expression. These genes are deeply involved in naive PCa and CRPC pathogenesis and are strong candidates for therapeutic targets of the disease.
In this study, we focused on HMGB3 and investigated its functional significance in PCa cells. HMGB3 is a member of the high-mobility group box family that currently includes four members (HMGB1-HMGB4) [37]. Overexpression of HMGB3 was reported in several cancers, e.g., breast cancer, gastric adenocarcinoma, bladder cancer, and esophageal cancer, and its high expression was associated with poor prognosis of the patients [38,39,40,41]. Our present data showed that overexpression of HMGB3 was observed in naive PCa and CRPC specimens and its high expression was significantly associated with shorter disease-free survival, high T stage, positive lymph node status and high Gleason score. HMGB3 could be a useful prognostic marker in PCa pathogenesis and other cancers. Our present analyses and previous studies showed that aberrantly expressed HMGB3 acted as an oncogene in human cancers [42].
Our efforts focused on the identification of molecular targets and pathways controlled by HMGB3 in PCa cells. Towards that end, we used gene expression analysis of si-HMGB3 transfectants to investigate how HMGB3 affected other genes. The expression levels of eight genes were suppressed by the knockdown of HMGB3 in PCa cells. Interestingly, three genes, MCM5, MCM7, and MCM4, members of the minichromosome maintenance protein (MCM) family were downregulated by HMGB3 knockdown. The MCM family consists of six homologs (MCM2-MCM7), and these members act as eukaryotic DNA replicative helicases [43]. Overexpression of MCM members was reported in several cancers and aberrantly expressed MCMs have been shown to be reliable markers of malignancy [43, 44]. Previous studies reported that amplification and overexpression of MCM7 was associated with PCa clinicopathological features, such as relapse, local invasion, and tumor grade [45]. Furthermore, constitutive expression of MCM7 in a PCa cell line markedly increased DNA synthesis and cell proliferation [46]. A large cohort study in the TCGA database showed that high expression of MCM7 was significantly associated with disease-free survival of patients with PCa (Supplemental Fig. 3). Our present data indicated that HMGB3 and its modulated genes might promote PCa malignancies.
In conclusion, antitumor miR-205-5p directly targets oncogenic HMGB3 in naive PCa and CRPC cells. High expression of HMBG3 was significantly associated with shorter disease-free survival of PCa patients. HMGB3-controlled genes were deeply involved in PCa pathogenesis. Our strategy for identification of genes regulated by antitumor miRNAs is effective and this novel approach may contribute to the development of new therapeutic strategies.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67:7–30.
Labrie F. Hormonal therapy of prostate cancer. Prog Brain Res. 2010;182:321–41.
Loriot Y, Bianchini D, Ileana E, Sandhu S, Patrikidou A, Pezaro C, et al. Antitumour activity of abiraterone acetate against metastatic castration-resistant prostate cancer progressing after docetaxel and enzalutamide (MDV3100). Ann Oncol. 2013;24:1807–12.
Sartor AO, Fitzpatrick JM. Urologists and oncologists: adapting to a new treatment paradigm in castration-resistant prostate cancer (CRPC). BJU Int. 2012;110:328–35.
Sridhar SS, Freedland SJ, Gleave ME, Higano C, Mulders P, Parker C, et al. Castration-resistant prostate cancer: from new pathophysiology to new treatment. Eur Urol. 2014;65:289–99.
Sturge J, Caley MP, Waxman J. Bone metastasis in prostate cancer: emerging therapeutic strategies. Nat Rev Clin Oncol. 2011;8:357–68.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Adams BD, Kasinski AL, Slack FJ. Aberrant regulation and function of microRNAs in cancer. Curr Biol. 2014;24:R762–776.
Kojima S, Goto Y, Naya Y. The roles of microRNAs in the progression of castration-resistant prostate cancer. J Hum Genet. 2017;62:25–31.
Goto Y, Kurozumi A, Enokida H, Ichikawa T, Seki N. Functional significance of aberrantly expressed microRNAs in prostate cancer. Int J Urol. 2015;22:242–52.
Kurozumi A, Goto Y, Okato A, Ichikawa T, Seki N. Aberrantly expressed microRNAs in bladder cancer and renal cell carcinoma. J Hum Genet. 2017;62:49–56.
Koshizuka K, Hanazawa T, Fukumoto I, Kikkawa N, Okamoto Y, Seki N. The microRNA signatures: aberrantly expressed microRNAs in head and neck squamous cell carcinoma. J Hum Genet. 2017;62:3–13.
Koshizuka K, Hanazawa T, Arai T, Okato A, Kikkawa N, Seki N. Involvement of aberrantly expressed microRNAs in the pathogenesis of head and neck squamous cell carcinoma. Cancer Metastasis Rev. 2017;36:525–45.
Nohata N, Hanazawa T, Kinoshita T, Okamoto Y, Seki N. MicroRNAs function as tumor suppressors or oncogenes: aberrant expression of microRNAs in head and neck squamous cell carcinoma. Auris Nasus Larynx. 2013;40:143–9.
Mizuno K, Mataki H, Arai T, Okato A, Kamikawaji K, Kumamoto T et al. The microRNA expression signature of small cell lung cancer: tumor suppressors of miR-27a-5p and miR-34b-3p and their targeted oncogenes. J Hum Genet. 2017;62:671–8.
Fuse M, Kojima S, Enokida H, Chiyomaru T, Yoshino H, Nohata N, et al. Tumor suppressive microRNAs (miR-222 and miR-31) regulate molecular pathways based on microRNA expression signature in prostate cancer. J Hum Genet. 2012;57:691–9.
Goto Y, Kojima S, Nishikawa R, Kurozumi A, Kato M, Enokida H, et al. MicroRNA expression signature of castration-resistant prostate cancer: the microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br J Cancer. 2015;113:1055–65.
Goto Y, Kurozumi A, Arai T, Nohata N, Kojima S, Okato A, et al. Impact of novel miR-145-3p regulatory networks on survival in patients with castration-resistant prostate cancer. Br J Cancer. 2017;117:409–20.
Goto Y, Kurozumi A, Nohata N, Kojima S, Matsushita R, Yoshino H, et al. The microRNA signature of patients with sunitinib failure: regulation of UHRF1 pathways by microRNA-101 in renal cell carcinoma. Oncotarget. 2016;7:59070–86.
Koshizuka K, Nohata N, Hanazawa T, Kikkawa N, Arai T, Okato A, et al. Deep sequencing-based microRNA expression signatures in head and neck squamous cell carcinoma: dual strands of pre-miR-150 as antitumor miRNAs. Oncotarget. 2017;8:30288–304.
Fukumoto I, Kinoshita T, Hanazawa T, Kikkawa N, Chiyomaru T, Enokida H, et al. Identification of tumour suppressive microRNA-451a in hypopharyngeal squamous cell carcinoma based on microRNA expression signature. Br J Cancer. 2014;111:386–94.
Nishikawa R, Goto Y, Kurozumi A, Matsushita R, Enokida H, Kojima S, et al. MicroRNA-205 inhibits cancer cell migration and invasion via modulation of centromere protein F regulating pathways in prostate cancer. Int J Urol. 2015;22:867–77.
Arai T, Okato A, Kojima S, Idichi T, Koshizuka K, Kurozumi A et al. Regulation of spindle and kinetochore-associated protein 1 by antitumor miR-10a-5p in renal cell carcinoma. Cancer Sci. 2017;108:2088–101.
Anaya J. OncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. Peer J Comp Sci. 2016;2:e 67.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Kato M, Goto Y, Matsushita R, Kurozumi A, Fukumoto I, Nishikawa R, et al. MicroRNA-26a/b directly regulate La-related protein 1 and inhibit cancer cell invasion in prostate cancer. Int J Oncol. 2015;47:710–8.
Kurozumi A, Goto Y, Matsushita R, Fukumoto I, Kato M, Nishikawa R, et al. Tumor-suppressive microRNA-223 inhibits cancer cell migration and invasion by targeting ITGA3/ITGB1 signaling in prostate cancer. Cancer Sci. 2016;107:84–94.
Nishikawa R, Goto Y, Sakamoto S, Chiyomaru T, Enokida H, Kojima S, et al. Tumor-suppressive microRNA-218 inhibits cancer cell migration and invasion via targeting of LASP1 in prostate cancer. Cancer Sci. 2014;105:802–11.
Nishikawa R, Goto Y, Kojima S, Enokida H, Chiyomaru T, Kinoshita T, et al. Tumor-suppressive microRNA-29s inhibit cancer cell migration and invasion via targeting LAMC1 in prostate cancer. Int J Oncol. 2014;45:401–10.
Okato A, Arai T, Kojima S, Koshizuka K, Osako Y, Idichi T, et al. Dual strands of pre-miR150 (miR1505p and miR1503p) act as antitumor miRNAs targeting SPOCK1 in naive and castration-resistant prostate cancer. Int J Oncol. 2017;51:245–56.
Li C, Finkelstein D, Sherr CJ. Arf tumor suppressor and miR-205 regulate cell adhesion and formation of extraembryonic endoderm from pluripotent stem cells. Proc Natl Acad Sci USA. 2013;110:E1112–1121.
Zhang JY, Sun MY, Song NH, Deng ZL, Xue CY, Yang J. Prognostic role of microRNA-205 in multiple human malignant neoplasms: a meta-analysis of 17 studies. BMJ Open. 2015;5:e006244.
Tucci P, Agostini M, Grespi F, Markert EK, Terrinoni A, Vousden KH, et al. Loss of p63 and its microRNA-205 target results in enhanced cell migration and metastasis in prostate cancer. Proc Natl Acad Sci USA. 2012;109:15312–7.
Griffiths K, Morton MS, Nicholson RI. Androgens, androgen receptors, antiandrogens and the treatment of prostate cancer. Eur Urol. 1997;32(Suppl 3):24–40.
Hagman Z, Haflidadottir BS, Ceder JA, Larne O, Bjartell A, Lilja H, et al. miR-205 negatively regulates the androgen receptor and is associated with adverse outcome of prostate cancer patients. Br J Cancer. 2013;108:1668–76.
Yang F, Li R, Hong A, Duan F, Li Y. Generation and characterization of a polyclonal antibody against human high mobility group box 4. Mol Med Rep.. 2013;8:1460–4.
Sun S, Zhang W, Cui Z, Chen Q, Xie P, Zhou C, et al. High mobility group box-1 and its clinical value in breast cancer. Onco Targets Ther. 2015;8:413–9.
Chung HW, Lim JB. High-mobility group box-1 contributes tumor angiogenesis under interleukin-8 mediation during gastric cancer progression. Cancer Sci. 2017;108:1594–601.
Yang GL, Zhang LH, Bo JJ, Huo XJ, Chen HG, Cao M, et al. Increased expression of HMGB1 is associated with poor prognosis in human bladder cancer. J Surg Oncol. 2012;106:57–61.
Sato N, Koinuma J, Fujita M, Hosokawa M, Ito T, Tsuchiya E, et al. Activation of WD repeat and high-mobility group box DNA binding protein 1 in pulmonary and esophageal carcinogenesis. Clin Cancer Res. 2010;16:226–39.
Xia Y, Papalopulu N, Vogt PK, Li J. The oncogenic potential of the high mobility group box protein Sox3. Cancer Res. 2000;60:6303–6.
Tsai FL, Vijayraghavan S, Prinz J, MacAlpine HK, MacAlpine DM, Schwacha A. Mcm2-7 is an active player in the DNA replication checkpoint signaling cascade via proposed modulation of Its DNA Gate. Mol Cell Biol. 2015;35:2131–43.
Kwok HF, Zhang SD, McCrudden CM, Yuen HF, Ting KP, Wen Q, et al. Prognostic significance of minichromosome maintenance proteins in breast cancer. Am J Cancer Res. 2015;5:52–71.
Ren B, Yu G, Tseng GC, Cieply K, Gavel T, Nelson J, et al. MCM7 amplification and overexpression are associated with prostate cancer progression. Oncogene. 2006;25:1090–8.
Padmanabhan V, Callas P, Philips G, Trainer TD, Beatty BG. DNA replication regulation protein Mcm7 as a marker of proliferation in prostate cancer. J Clin Pathol. 2004;57:1057–62.
Acknowledgements
This study was supported by JSPS KAKENHI (grant numbers; 16H05462, 17K16777, 16K20125, 17K11160, and 15K10801).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
10038_2017_371_MOESM3_ESM.pptx
Kaplan-Meier survival curves showing disease-free survival rates for minichromosome maintenance complex component (MCM) family, genes that are downstream from HMGB3
Clinical features of patients with naive PCa and CRPC in expression analyses of miR-205-5p and HBGB3
Rights and permissions
About this article
Cite this article
Yamada, Y., Nishikawa, R., Kato, M. et al. Regulation of HMGB3 by antitumor miR-205-5p inhibits cancer cell aggressiveness and is involved in prostate cancer pathogenesis. J Hum Genet 63, 195–205 (2018). https://doi.org/10.1038/s10038-017-0371-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0371-1
This article is cited by
-
A pathway activity-based proteomic classifier stratifies prostate tumors into two subtypes
Clinical Proteomics (2023)
-
Circ-IGF1R Affects the Progression of Colorectal Cancer by Activating the miR-362-5p/HMGB3-Mediated Wnt/β-Catenin Signal Pathway
Biochemical Genetics (2023)
-
Hsa_circ_0000078 Regulates miR-205-5p/EREG Pathway to Inhibit Cervical Cancer Progression
Molecular Biotechnology (2023)
-
Down-regulation of SNHG16 alleviates the acute lung injury in sepsis rats through miR-128-3p/HMGB3 axis
BMC Pulmonary Medicine (2021)
-
The role of high mobility group protein B3 (HMGB3) in tumor proliferation and drug resistance
Molecular and Cellular Biochemistry (2021)