Abstract
Laboratory rats and mice are representative experimental animals for models of human disease. The emergence of genome editing technologies has enabled us to produce a variety of genetically modified animals, including rats, as a means of elucidating the in vivo functions of the gene of interest and characterizing the molecular mechanisms of human disease. Several advanced techniques for knock-in methodologies in rats are currently in development, which permit researchers to introduce precise nucleotide modifications at target sites in the rat’s genome. Furthermore, recent studies with knock-out rats have revealed that observed disease phenotypes are often more similar than mouse models to those of humans. In this article, we introduce the methodologies for efficient gene manipulation in rats using genome editing technologies, and describe the advances made using rats for human disease models. We also discuss the importance of gene manipulation in animal models for the better understanding of fundamental processes among different species.
Similar content being viewed by others
Introduction
In vivo experiments using laboratory animals are indispensable for the elucidation of pathological and physiological mechanisms in human diseases, which can lead to the development of new treatments and prevention approaches. Altogether with the mouse, the laboratory rat (Rattus norvegicus) is a representative experimental mammal that has been widely used for the past century as a human disease model for hypertension [1], diabetes [2, 3], epilepsy [4, 5], inflammation [6], and cancer [7, 8]. Indeed, rats are often a better choice than mice because of their larger body size and ease of manipulation. They are useful not only for longitudinal drug efficiency tests and toxicity assessments but also for surgical manipulations in neurological research such as electrode insertions in the brain and optogenetic neural stimulation [9]. Moreover, comprehensive database platforms related to laboratory rats have been recently developed, for example, the National BioResource Project-Rat in Japan and The Rat Genome Database in the US [10, 11]. These platforms provide valuable resources and updated datasets of genomic elements including gene annotations, transcripts, sequence information, and variations associated with phenotypes and disease.
While rats have contributed greatly to the development of therapeutic agents and regimens as human disease models, they have lagged behind mice in the genetic research field. Mouse reproductive technologies such as in vitro fertilization and embryo manipulation have been established since the 1970s and gene knock-out (KO) mice using embryonic stem (ES) cells were developed before 1990 as critical tools for understanding gene functions [12,13,14]. Knock-in (KI) mice, including reporter tagging of endogenous genes and conditional knock-outs, are widely used for spatial or temporal observation to control gene activation, avoiding early lethal phenotypes [15, 16]. Despite these advances, ES cell-mediated gene targeting technologies remain critical tools for understanding gene functions in mice. Although fundamental reproductive technologies have also been developed in rats, it has proved more difficult to produce KO rats using gene targeting technologies because of the lack of optimal culture conditions for germline-competent ES cells until the mid-2000s.
Therefore, before the availability of ES cell-mediated gene targeting and genome editing technologies for rats, transgenic techniques involving random mutagenesis were used instead. Transgenic rats can be generated by the microinjection of donor DNA derived from another species into a male pronucleus of a fertilized egg. This enables the random integration of foreign genes into a rat genome which permits investigation of the in vivo function of the gene of interest. These methods contributed to the past acquisition of important biological knowledge about human diseases such as inflammation [17], neurological disorders [18], and cancer [19]. Random mutagenesis can induce mutations throughout the rat genome by introducing a transposon such as Sleeping Beauty or a chemical mutagen such as N-ethyl-N-nitrosourea (ENU) [20, 21]. However, these methods require large populations of rats and a high-throughput screening strategy to identify mutations in a targeted gene. Thus, several strategies have been developed to identify ENU-induced mutations in animals, including yeast-based screening [21], CelI-based enzymatic cleavage [22], sequencing-based screening [23], and Mu transposase-based heteroduplex identification [24].
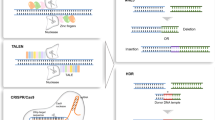
Mouse ES cells have facilitated the application of genome engineering technologies to create precise genetic modifications in vitro via homologous recombination (HR). Transplantation of these cells into a host embryo can then generate a chimeric mouse carrying genetic modifications. Although several methodologies have been attempted to establish authentic rat ES cells, there is no robust evidence of chimera generation because rat ES cell stability can be greatly influenced by culture conditions and the genetic background of rat strains. A new strategy to maintain rat-derived ES cells was finally developed in 2008 using the 3i/2i culture system [25]. This method lead to the generation of a p53 gene KO rat via HR in 2010, almost 20 per s after the emergence of KO mice [26]. Around this time, other genome editing milestones were developed such as zinc finger nucleases (ZFNs), transcription activator-like (TAL) effector nucleases (TALENs), and clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated nucleases (Cas). These groundbreaking technologies are major approaches for gene modification that avoid the need for germline-competent ES cells. Furthermore, their simplicity, efficiency, and reliability facilitate KO and KI production in a variety of animals including mice and rats [27]. Here, we introduce recent methods of these innovative technologies that have been used to manipulate a rat genome, and describe genetically modified rats that have been developed as models of human disease.
Generation of genetically modified rats using ZFN, TALEN, and CRISPR/Cas9
ZFNs are artificial fusion proteins carrying DNA binding domains, which consist of tandem zinc finger motifs with customized specificity and a non-specific nuclease domain from the endonuclease FokI [28, 29]. FokI domains can introduce a double-strand break (DSB) at a precise genomic locus following recognition of the targeted DNA sequence by paired ZFNs. Inserted or deleted mutations often occur at the DSB site via the non-homologous end joining (NHEJ) repair process. Introduction of a DSB at the coding region can cause a frameshift mutation of the targeted gene, which creates a KO mutation. The process of generating KO rats using ZFNs is facilitated by the microinjection of DNA or RNA coding for ZFN components into fertilized rat embryos (Fig. 1). Moreover, KO founder rats with a KO efficiency >20% can be generated in the G0 generation within 3–4 months, which is quicker than the ES cell-based method for mice that usually takes 12–18 months. In addition, gene targeting with artificial nucleases is not strain-dependent and accordingly can be performed with any inbred strains. Therefore, numerous research articles about KO rats generated using ZFN technology have been published since the first report of GFP-KO rats in 2009 [30,31,32,33]. However, despite the great contribution of ZFNs to KO rats, several limitations remain associated with this technique. These include the difficulty of producing suitable ZFNs for the targeted sequence, the potential to produce off-target cleavage and mutations, and the high expense of custom-designed ZFNs [27, 34].
Experimental procedures for generating genetically modified rats with the CRISPR/Cas9 system. gRNA recognizing the target sequence in the genome is designed using web tools to avoid off-target effects. The target-specific gRNA and Cas9 mRNA or protein are then introduced into rat embryos. It is also possible to introduce an all-in-one plasmid expressing Cas9 and gRNA. Donor DNAs such as ssODN and plasmid DNA should be mixed and co-introduced into embryos. While microinjection is a popular choice for this, electroporation and lentiviral infection are suitable alternatives. About 3 weeks after transfer of the eggs to a pseudopregnant females, founder pups are obtained. Genetic mutations can be identified in founder individuals by DNA sequencing analysis. The off-target effect and germline transmission to the next generation can then be confirmed
TALENs are an alternative tool for genome engineering [35,36,37]. They are also fusion proteins of tandem repeats of a TAL effector protein derived from plant-pathogenic Xanthomonas bacteria and the FokI nuclease. While the targeted sequences are severely limited because of the recognition pattern of ZF domains, TAL effector motifs can recognize any sequence except when thymine is at the first position. TALENs cleave the targeted region in pairs, similar to ZFNs, which increases their efficiency as a tool for KO animals and cells [38, 39]. To provide a cost-effective targeted nuclease platform at the general laboratory level, several strategies have been established for the simple and rapid construction of TALENs [40,41,42,43,44,45]. For example, Sakuma et al. [42] showed that TALENs with periodically patterned repeat variants harboring non-repeat-variable di-residue (non-RVD) variations (Platinum TALENs) have higher activities than TALENs without non-RVD variations. We applied the Platinum TALEN to the rat Il2rg gene to produce KO rats at a higher efficiency compared with ZFNs and original TALENs.
Although ZFNs and TALENs have been applied to various species, CRISPR/Cas9, the newest genome editing tool, is widely used for generating KO and KI mutations in many cell types of different species including mice. The CRISPR/Cas9 system was first identified as a gene-targeting technology in mammalian cells [46,47,48]. Cas9 nucleases are combined with synthetic single-guide RNAs (gRNAs) that are complementary to the 20 bases of target sequence beside the NGG sequence. The complex then recognizes the target sequence and introduces DSBs. The system enables the navigation of Cas9 to any genomic locus through the design of synthetic gRNA, which can be readily generated. The precision of this mechanism has made the CRISPR/Cas system more renowned as a robust genome editing tool than ZFNs and TALENs [49,50,51,52]. Several studies have reported the generation of targeted KO in rats using CRISPR/Cas9 [53,54,55,56], and we also generated a targeted KO rat at the tyrosinase (Tyr) locus by microinjecting target-specific gRNA and Cas9 mRNA into fertilized eggs [57]. Crossing founders demonstrated that the CRISPR/Cas9-mediated mutations were faithfully transmitted to the next generation.
In addition to producing a gene KO rat, we used the target-specificity of CRISPR/Cas9 to perform allele-specific genome editing [57]. An albino allele-recognizing gRNA:Tyrc was introduced into F1 rat embryos from a DA strain×F344 strain cross containing wild-type Tyr and single-nucleotide polymorphisms (SNPs) for each albino type, leading to the occurrence of KO mutations only at the albino allele. Conversely, introduction of a wild allele-recognizing gRNA:Tyrc caused mutations to occur only at this allele. This suggested that the CRISPR/Cas9 system can be used for highly accurate allele-specific genome editing, which is applicable to the highly heterogenized human genome. In the near future, CRISPR/Cas9 is predicted to be used as gene therapy for destroying genetic mutations that cause human disease and restoring mutated regions to normal alleles [58].
Cas9 and gRNA expression vectors have been used for genome editing in rodent embryos [59, 60]. These vectors can be readily prepared at low cost and express high levels of Cas9 and gRNA over several days, leading to more complete embryonic modifications. However, the potential for off-target mutations and random integration of plasmids into genomic DNA may also be increased. We used Cas9 mRNA for genome editing in rodents to avoid these limitations. Although it is necessary to transcribe them in vitro, the toxicity level is very low and the mRNA in a fertilized egg can express Cas9 protein rapidly and transiently. Recently, highly active purified Cas9 protein has been made commercially available, and is ready for use in genome editing of embryos as soon as it is mixed with gRNAs. The Cas9 and gRNA complex can efficiently modify the rodent genome with a high turnover that induces fewer off-target effects. Several researchers have also reported rodent genome editing using Cas9 protein introduced by microinjection and electroporation [61,62,63].
Methodologies to introduce CRISPR components such as Cas9 mRNA and gRNA into embryos have also improved the efficiency of gene modification leading to time and cost savings (Fig. 1). Microinjection by manipulating a glass needle under a microscope into fertilized eggs is the main strategy for producing genetically modified rats, as well as the conventional method for producing transgenic animals. However, laboratories routinely performing this technique are limited because of the requirement for advanced skills and expensive equipment. Recently, several electroporation methods for mouse and rat embryos have been developed to overcome these limitations [64,65,66,67]. During electroporation, DNA or RNA can be introduced through fine holes in the egg membrane that have been generated by a series of electric pulses. We successfully optimized electric pulse conditions for fertilized rat and mouse eggs, which led to the highly efficient generation of KO rats [64, 65]. This method can simultaneously introduce DNA and RNA into about 100 cells in one experiment by arranging fertilized eggs in line into the medium. It requires no complicated skills, and can also be applied to fertilized eggs from various species. In addition, not only Cas9 mRNA but also Cas9 protein can be available for gene modification by electroporation [68]. This technique is expected to become a popular method for the introduction of CRISPR components into fertilized eggs because of its high level of convenience.
Efficient methodologies for precise KI
The development of effective genome editing technologies to generate KI models is one of the major research subjects in laboratory animal science. KI animals may carry insertions in their genomes such as additional genes, amino acid tags, fluorescent reporters, or conditional knockout alleles, which aid our study of human disease and provide more sophisticated information about targeted gene functions. The KI strategy using the CRISPR/Cas9 platform is based on the co-introduction of CRISPR/Cas9 components with double-stranded DNA or single-stranded oligodeoxynucleotides (ssODNs). CRISPR/Cas9 induction of a DSB at the target site is restored via homology directed repair using a DNA template homologous to both ends of the DSB site. Utilization of this mechanism enables researchers to insert, delete, or replace genetic material of choice into the genome [55, 69, 70]. However, the KI efficiency depends on the frequency of the HR pathway. Several small molecules, such as RS-1 or Scr7, that enhance the HR pathway or inhibit NHEJ pathways can increase the HR efficiency in cell lines [71,72,73]. However, researchers still face difficulties with KI in cell lines and animals with low HR frequencies. Furthermore, it is necessary to construct targeting vectors by adding two homology arms of appropriate length to both sides of the inserted DNA sequence.
Several laboratories including our own have reported on targeted KI such as an SNP or small size integration with ssODN donors in mice and rats (Fig. 2) [51, 57]. Although high-grade ssODNs are readily designed and synthesized by several companies, their maximum length is only ~200 bases. Therefore, it is difficult to use this system for integrating long sequences such as green fluorescent protein (GFP) gene cassettes into the genome. To overcome this, laboratory-based strategies have generated long single-strand DNAs (lssDNAs) by PCR-mediated methods including magnetic separation by streptavidin-coated beads [74], lambda exonuclease enzymatic digestion [75], and asymmetric PCR using unequal concentrations of primers [76]. Recently, we demonstrated a novel method to generate highly purified lssDNAs using double nicking endonucleases [77] that digest DNA plasmids including lssDNA sequences between two nickase-specific sites; lssDNAs are then purified by electrophoresis and gel extraction. We microinjected lssDNAs with CRISPR/Cas9 components to generate several types of KI, such as GFP cassette insertions and conditional KO alleles in mice and rats (Fig. 2). Another research group generated KI mice with lssDNAs synthesized by reverse transcription and the digestion of an RNA template [61, 78]. LssDNAs can also be used in electroporation-mediated KI in embryos as a simpler, more rapid, and efficient technique than microinjection-mediated KIs (Miyasaka, unpublished data). In the near future, nucleotide synthesis technology is expected to allow us to synthesize even longer lssDNAs of several kilobases which will extend the possibilities of generating more productive animal KIs.
Strategies for single strand DNA-mediated KI with CRISPR/Cas9. a Single-stranded oligodeoxynucleotide (ssODN)-mediated KI for SNP substitution. The introduction of ssODNs with short homology arms can generate SNP substitutions or short fragment insertions at the targeted site of genome editing. b Long single-stranded DNA (lssDNA)-mediated KI for large genetic insertions. LssDNAs including exogenous gene sequences with homologous arms can produce genetic insertions at the targeted site, as well as ssODN-mediated KI. The length of homology is typically 50–300 bases, although this should be optimized fort each targeted region. c LssDNA-mediated KI for floxed alleles. The co-introduction of two gRNAs and lssDNAs carrying an exon between two loxP sites can induce the genetic replacement of targeted exons with the floxed sequence. DSB double strand break, HA homology arm
Alternatively, NHEJ and the microhomology-mediated end-joining (MMEJ) pathway can assist the targeted integration of donor DNAs into DSB sites (Fig. 3). Improved KI strategies using these systems have been reported as ObLiGaRe and CRIS-PITCh methods [79, 80]. When microhomologies from several bases to several tens of bases in length exist at both DNA ends, they can be combined and ligate together via the MMEJ pathway. This enables the directional integration of linearized donor DNA into the DSB site. MMEJ-assisted genome editing strategies have also been reported for the generation of cassette KI in mammalian cells and zygotes [81, 82].
Strategies for plasmid KI with CRISPR/Cas9. a Homologous recombination (HR)-mediated KI. Exogenous genes can be inserted via HR without backbone integration, as well as into ES cells. The KI efficiency of this technique is low and requires the inclusion of 1–2 kb homology arms in the plasmid. b Non-homologous end joining (NHEJ)-mediated KI. This technique integrates the plasmid into the cleaved region with a higher efficiency than random integration, but there are several risks including bidirectional and multiple copy insertion. c Microhomology-mediated end-joining (MMEJ)-mediated KI. The design of several bases of microhomology allows the generation of directional plasmid integration via the MMEJ repair system. Design and development of the plasmid carrying microhomology arms of cleaved genomic regions must be done with precision. d ssODN-mediated KI that we developed as a 2H2OP method. Plasmids can be integrated directionally via ssODN-mediated ligation with no plasmid modification, though the plasmid backbone is also integrated. e CAG-GFP knock-in rat (arrow) prepared by the 2H2OP method, and its sequencing analysis. The CAG-GFP plasmid was inserted into an intron of the rat Rosa26 locus. DSB double strand break, HA homology arm, MA microhomology arm
We have developed another strategy for efficient KI in zygotes using the NHEJ system assisted by two ssODNs; this is known as the two-hit by gRNAs and two oligos with the targeting plasmid (2H2OP) method (Fig. 3) [77]. In this technique, the co-introduction of two target-specific gRNAs excises target sites in genomic and plasmid DNA, producing cutting edges without homology arms between the genome and the plasmid. Two short ssODNs carrying homology arms between the genome and the plasmid then assist in ligating the ends cut by CRISPR/Cas9, resulting in the integration of an extrinsic targeting vector into the endogenous targeted region. This method can modify both small and large genomic regions, resulting in the generation of bacterial artificial chromosome KI including the replacement of whole genes, gene clusters, and endogenous promoters [77]. This enables evaluation of the intact in vivo function of targeted genes.
Genetically modified rats elucidate the mechanisms of human disease
While genome editing technologies are progressing rapidly, gene functional analysis using laboratory mice is still at a fundamental level. The characterization of gene function using multiple species is nevertheless vital to clarify universal in vivo functions and species-specific functions of the targeted gene. However, some experiments with genetically modified mice found no significant difference in predicted disease phenotype compared with wild-type. Recent studies with genetically modified rats reported closer phenotypes to human disease than mice; for example, rat models have contributed more to colorectal cancer research. Adenomatous polyposis coli (Apc) mutant mouse strains such as the Min mouse demonstrate tumorigenesis in their intestinal tract, so have been used as a human colorectal cancer model [83]. However, their main site of polyp formation is in the small intestine rather than the large intestine where tumors develop in humans with familial adenomatous polyposis. Therefore it is difficult to observe the real-time sequential changes of these tumors in live animals. On the other hand, the Apc KO rat developed by ENU mutagenesis spontaneously developed colorectal tumors in the large intestine, similar to humans [84]. It was also possible to observe and manipulate specific tumors using an endoscope, providing us with a useful model of colon cancer development [7].
In immunology research, a KO mouse with severe combined immunodeficiency (SCID) was generated by interrupting the Prkdc gene which is crucial for T cell and B cell function. However, homozygous KO mice showed neither fetal lethality nor the decrease in cell proliferation seen in humans with SCID. On the other hand, SCID rats that are deficient in the Prkdc gene following the use of TALEN demonstrated significant weight loss and decreased cell proliferation, as well as immunodeficiency [33, 85]. Furthermore, SCID mice with the “leaky” phenotype sometimes recovered their defect immune system through the production of immunoglobulins such as IgG in the blood, whereas SCID rats showed no immune system recovery [86]. This suggests that the rat model could be used for the xenotransplantation of human cancer cells, stem cells, and tissues [87].
KO rats are also useful models for human neurological disorders. For example, Atm KO rats showed clear signs of neurodegeneration in the spinal cord and developed hind limb paralysis, as early as 4 months of age, which is likely caused by the loss of motor neurons in the lumbar region of the spinal cord [88]. Moreover, although there are discrepancies between rats and humans, the KO rat can be useful for providing insights into the disease mechanisms of ataxia telangiectasia. In another example, Bscl2 KO rats showed several abnormalities associated with nervous systems that were not seen in mice [89]. For instance, Bscl2 associated with fat atrophy was strongly expressed in mouse adipose tissue and testes, whereas rats also expressed it in the brain, similar to humans. This likely explains the reduction of spatial task memory seen in the KO rat model.
In addition to the physiological similarities between KO rat models and human systems, recent studies have demonstrated that the gut bacterial communities of humanized models achieved by fecal microbiota transplantation in rats more closely reflect the gut microbiota of human donors than those of mice. Hence, rats can also be used as representative models of digestive system disorders [90]. These examples show that the choice of animal model depends on the purpose and aim of the study. Indeed, the intensity and distribution of gene expression can be dependent on the species. Therefore, the influence of the species should be carefully considered to obtain reliable and reproducible findings.
Conclusion
In this review, we have described the history and the recent developments in cutting-edge genetic modification technologies. We have also explained the utility of genetically modified rats as models for human diseases. Such rats can be developed quickly with a high level of efficiency compared with ES cell-mediated modification. Moreover, new techniques such as nucleic acid introduction are rapidly progressing alongside in vivo genome editing methods using fertilized eggs. KI generation is one of the most informative approaches for gene functional assessment. It can be used to establish humanized animals as models by introducing SNP or mutations identified in human diseases, and replacing the genomic region of animals with human genomic sequences. These humanized animals are expected to contribute to the elucidation of novel gene functions and to be beneficial in translational medicine.
Taken together, rats are powerful animal models because their biological background and systems are more similar than mice to those of humans. The use of genetically engineered rats in biomedical research can therefore provide fundamental knowledge to understand human physiological and pathological systems and to help develop therapeutic strategies for human disease.
References
Yamori Y. Overview: studies on spontaneous hypertension-development from animal models toward man. Clin Exp Hypertens A. 1991;13:631–44.
Chappel CI, Chappel WR. The discovery and development of the BB rat colony: an animal model of spontaneous diabetes mellitus. Metabolism. 1983;32:8–10.
Kawano K, Hirashima T, Mori S, Natori T. OLETF (Otsuka Long-Evans Tokushima Fatty) rat: a new NIDDM rat strain. Diab Res Clin Pract. 1994;24:S317–320.
Mashimo T, Ohmori I, Ouchida M, Ohno Y, Tsurumi T, Miki T, et al. A missense mutation of the gene encoding voltage-dependent sodium channel (Nav1.1) confers susceptibility to febrile seizures in rats. J Neurosci. 2010;30:5744–53.
Serikawa T, Yamada J. Epileptic seizures in rats homozygous for two mutations, zitter and tremor. J Hered. 1986;77:441–4.
Kleinau S, Erlandsson H, Klareskog L. Percutaneous exposure of adjuvant oil causes arthritis in DA rats. Clin Exp Immunol. 1994;96:281–4.
Irving AA, Yoshimi K, Hart ML, Parker T, Clipson L, Ford MR, et al. The utility of Apc-mutant rats in modeling human colon cancer. Dis Model Mech. 2014;7:1215–25.
Mori M, Hattori A, Sawaki M, Tsuzuki N, Sawada N, Oyamada M, et al. The LEC rat: a model for human hepatitis, liver cancer, and much more. Am J Pathol. 1994;144:200–4.
Diester I, Kaufman MT, Mogri M, Pashaie R, Goo W, Yizhar O, et al. An optogenetic toolbox designed for primates. Nat Neurosci. 2011;14:387–97.
Serikawa T, Mashimo T, Takizawa A, Okajima R, Maedomari N, Kumafuji K, et al. National BioResource project-rat and related activities. Exp Anim. 2009;58:333–41.
Shimoyama M, De Pons J, Hayman GT, Laulederkind SJ, Liu W, Nigam R, et al. The Rat genome database 2015: genomic, phenotypic and environmental variations and disease. Nucleic Acids Res. 2015;43:D743–750.
Koller BH, Hagemann LJ, Doetschman T, Hagaman JR, Huang S, Williams PJ, et al. Germ-line transmission of a planned alteration made in a hypoxanthine phosphoribosyltransferase gene by homologous recombination in embryonic stem cells. Proc Natl Acad Sci USA. 1989;86:8927–31.
Mansour SL, Thomas KR, Capecchi MR. Disruption of the proto-oncogene int-2 in mouse embryo-derived stem cells: a general strategy for targeting mutations to non-selectable genes. Nature. 1988;336:348–52.
Thompson S, Clarke AR, Pow AM, Hooper ML, Melton DW. Germ line transmission and expression of a corrected HPRT gene produced by gene targeting in embryonic stem cells. Cell. 1989;56:313–21.
Jackson EL, Willis N, Mercer K, Bronson RT, Crowley D, Montoya R, et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Gene Dev. 2001;15:3243–8.
Lewandoski M. Conditional control of gene expression in the mouse. Nat Rev Genet. 2001;2:743–55.
Hammer RE, Maika SD, Richardson JA, Tang JP, Taurog JD. Spontaneous inflammatory disease in transgenic rats expressing HLA-B27 and human beta 2m: an animal model of HLA-B27-associated human disorders. Cell. 1990;63:1099–112.
von Horsten S, Schmitt I, Nguyen HP, Holzmann C, Schmidt T, Walther T, et al. Transgenic rat model of Huntington’s disease. Hum Mol Genet. 2003;12:617–24.
Ota T, Asamoto M, Toriyama-Baba H, Yamamoto F, Matsuoka Y, Ochiya T, et al. Transgenic rats carrying copies of the human c-Ha-ras proto-oncogene exhibit enhanced susceptibility to N-butyl-N-(4-hydroxybutyl)nitrosamine bladder carcinogenesis. Carcinogenesis. 2000;21:1391–6.
Izsvak Z, Frohlich J, Grabundzija I, Shirley JR, Powell HM, Chapman KM, et al. Generating knockout rats by transposon mutagenesis in spermatogonial stem cells. Nat Methods. 2010;7:443–5.
Zan Y, Haag JD, Chen KS, Shepel LA, Wigington D, Wang YR, et al. Production of knockout rats using ENU mutagenesis and a yeast-based screening assay. Nat Biotechnol. 2003;21:645–51.
Smits BM, Mudde J, Plasterk RH, Cuppen E. Target-selected mutagenesis of the rat. Genomics. 2004;83:332–4.
Smits BM, Mudde JB, van de Belt J, Verheul M, Olivier J, Homberg J, et al. Generation of gene knockouts and mutant models in the laboratory rat by ENU-driven target-selected mutagenesis. Pharmacogenet Genomics. 2006;16:159–69.
Mashimo T, Yanagihara K, Tokuda S, Voigt B, Takizawa A, Nakajima R, et al. An ENU-induced mutant archive for gene targeting in rats. Nat Genet. 2008;40:514–5.
Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B, Woodgett J, et al. The ground state of embryonic stem cell self-renewal. Nature. 2008;453:519–23.
Tong C, Li P, Wu NL, Yan Y, Ying QL. Production of p53 gene knockout rats by homologous recombination in embryonic stem cells. Nature. 2010;467:211–3.
Mashimo T. Gene targeting technologies in rats: zinc finger nucleases, transcription activator-like effector nucleases, and clustered regularly interspaced short palindromic repeats. Dev Growth Differ. 2014;56:46–52.
Bibikova M, Beumer K, Trautman JK, Carroll D. Enhancing gene targeting with designed zinc finger nucleases. Science. 2003;300:764.
Porteus MH, Carroll D. Gene targeting using zinc finger nucleases. Nat Biotechnol. 2005;23:967–73.
Geurts AM, Cost GJ, Freyvert Y, Zeitler B, Miller JC, Choi VM, et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science. 2009;325:433.
Cui X, Ji D, Fisher DA, Wu Y, Briner DM, Weinstein EJ. Targeted integration in rat and mouse embryos with zinc-finger nucleases. Nat Biotechnol. 2011;29:64–7.
Remy S, Tesson L, Menoret S, Usal C, Scharenberg AM, Anegon I. Zinc-finger nucleases: a powerful tool for genetic engineering of animals. Transgenic Res. 2010;19:363–71.
Mashimo T, Takizawa A, Voigt B, Yoshimi K, Hiai H, Kuramoto T, et al. Generation of knockout rats with X-linked severe combined immunodeficiency (X-SCID) using zinc-finger nucleases. PLoS ONE. 2010;5:e8870.
Jacob HJ, Lazar J, Dwinell MR, Moreno C, Geurts AM. Gene targeting in the rat: advances and opportunities. Trends Genet. 2010;26:510–8.
Mussolino C, Cathomen T. TALE nucleases: tailored genome engineering made easy. Curr Opin Biotechnol. 2012;23:644–50.
Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol. 2013;14:49–55.
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics. 2010;186:757–61.
Sung YH, Baek IJ, Kim DH, Jeon J, Lee J, Lee K, et al. Knockout mice created by TALEN-mediated gene targeting. Nat Biotechnol. 2013;31:23–4.
Hockemeyer D, Wang H, Kiani S, Lai CS, Gao Q, Cassady JP, et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol. 2011;29:731–4.
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 2011;39:e82.
Li T, Huang S, Zhao X, Wright DA, Carpenter S, Spalding MH, et al. Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res. 2011;39:6315–25.
Sakuma T, Ochiai H, Kaneko T, Mashimo T, Tokumasu D, Sakane Y, et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity. Sci Rep. 2013;3:3379.
Wood AJ, Lo TW, Zeitler B, Pickle CS, Ralston EJ, Lee AH, et al. Targeted genome editing across species using ZFNs and TALENs. Science. 2011;333:307.
Zhang F, Cong L, Lodato S, Kosuri S, Church GM, Arlotta P. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat Biotechnol. 2011;29:149–53.
Schmid-Burgk JL, Schmidt T, Kaiser V, Honing K, Hornung V. A ligation-independent cloning technique for high-throughput assembly of transcription activator-like effector genes. Nat Biotechnol. 2013;31:76–81.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23.
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339:823–6.
Cho SW, Kim S, Kim JM, Kim JS. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol. 2013;31:230–2.
Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD, et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol. 2013;31:227–9.
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F, et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell. 2013;153:910–8.
Yang H, Wang H, Shivalila CS, Cheng AW, Shi L, Jaenisch R. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell. 2013;154:1370–9.
Li D, Qiu Z, Shao Y, Chen Y, Guan Y, Liu M, et al. Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat Biotechnol. 2013;31:681–3.
Li W, Teng F, Li T, Zhou Q. Simultaneous generation and germline transmission of multiple gene mutations in rat using CRISPR-Cas systems. Nat Biotechnol. 2013;31:684–6.
Ma Y, Ma J, Zhang X, Chen W, Yu L, Lu Y, et al. Generation of eGFP and Cre knockin rats by CRISPR/Cas9. FEBS J. 2014;281:3779–90.
Ma Y, Zhang X, Shen B, Lu Y, Chen W, Ma J, et al. Generating rats with conditional alleles using CRISPR/Cas9. Cell Res. 2014;24:122–5.
Yoshimi K, Kaneko T, Voigt B, Mashimo T. Allele-specific genome editing and correction of disease-associated phenotypes in rats using the CRISPR-Cas platform. Nat Commun. 2014;5:4240.
Courtney DG, Moore JE, Atkinson SD, Maurizi E, Allen EH, Pedrioli DM, et al. CRISPR/Cas9 DNA cleavage at SNP-derived PAM enables both in vitro and in vivo KRT12 mutation-specific targeting. Gene Ther. 2016;23:108–12.
Mashiko D, Fujihara Y, Satouh Y, Miyata H, Isotani A, Ikawa M. Generation of mutant mice by pronuclear injection of circular plasmid expressing Cas9 and single guided RNA. Sci Rep. 2013;3:3355.
Mizuno S, Dinh TT, Kato K, Mizuno-Iijima S, Tanimoto Y, Daitoku Y, et al. Simple generation of albino C57BL/6 J mice with G291T mutation in the tyrosinase gene by the CRISPR/Cas9 system. Mamm Genome. 2014;25:327–34.
Quadros RM, Miura H, Harms DW, Akatsuka H, Sato T, Aida T, et al. Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol. 2017;18:92.
Hashimoto M, Yamashita Y, Takemoto T. Electroporation of Cas9 protein/sgRNA into early pronuclear zygotes generates non-mosaic mutants in the mouse. Dev Biol. 2016;418:1–9.
Aida T, Chiyo K, Usami T, Ishikubo H, Imahashi R, Wada Y, et al. Cloning-free CRISPR/Cas system facilitates functional cassette knock-in in mice. Genome Biol. 2015;16:87.
Kaneko T, Sakuma T, Yamamoto T, Mashimo T. Simple knockout by electroporation of engineered endonucleases into intact rat embryos. Sci Rep. 2014;4:6382.
Kaneko T, Mashimo T. Simple genome editing of rodent intact embryos by electroporation. PLoS ONE. 2015;10:e0142755.
Qin W, Dion SL, Kutny PM, Zhang Y, Cheng AW, Jillette NL, et al. Efficient CRISPR/Cas9-Mediated Genome Editing in Mice by Zygote Electroporation of Nuclease. Genetics. 2015;200:423–30.
Hashimoto M, Takemoto T. Electroporation enables the efficient mRNA delivery into the mouse zygotes and facilitates CRISPR/Cas9-based genome editing. Sci Rep. 2015;5:11315.
Chen S, Lee B, Lee AY, Modzelewski AJ, He L. Highly efficient mouse genome editing by CRISPR ribonucleoprotein electroporation of zygotes. J Biol Chem. 2016;291:14457–67.
Shao Y, Guan Y, Wang L, Qiu Z, Liu M, Chen Y, et al. CRISPR/Cas-mediated genome editing in the rat via direct injection of one-cell embryos. Nat Protoc. 2014;9:2493–512.
Remy S, Tesson L, Menoret S, Usal C, De Cian A, Thepenier V, et al. Efficient gene targeting by homology-directed repair in rat zygotes using TALE nucleases. Genome Res. 2014;24:1371–83.
Song J, Yang D, Xu J, Zhu T, Chen YE, Zhang J. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat Commun. 2016;7:10548.
Vartak SV, Raghavan SC. Inhibition of nonhomologous end joining to increase the specificity of CRISPR/Cas9 genome editing. FEBS J. 2015;282:4289–94.
Chu VT, Weber T, Wefers B, Wurst W, Sander S, Rajewsky K, et al. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat Biotechnol. 2015;33:543–8.
Hultman T, Stahl S, Hornes E, Uhlen M. Direct solid phase sequencing of genomic and plasmid DNA using magnetic beads as solid support. Nucleic Acids Res. 1989;17:4937–46.
Higuchi RG, Ochman H. Production of single-stranded DNA templates by exonuclease digestion following the polymerase chain reaction. Nucleic Acids Res. 1989;17:5865.
Gyllensten UB, Erlich HA. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc Natl Acad Sci USA. 1988;85:7652–6.
Yoshimi K, Kunihiro Y, Kaneko T, Nagahora H, Voigt B, Mashimo T. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat Commun. 2016;7:10431.
Miura H, Gurumurthy CB, Sato T, Sato M, Ohtsuka M. CRISPR/Cas9-based generation of knockdown mice by intronic insertion of artificial microRNA using longer single-stranded DNA. Sci Rep. 2015;5:12799.
Maresca M, Lin VG, Guo N, Yang Y. Obligate ligation-gated recombination (ObLiGaRe): custom-designed nuclease-mediated targeted integration through nonhomologous end joining. Genome Res. 2013;23:539–46.
Nakade S, Tsubota T, Sakane Y, Kume S, Sakamoto N, Obara M, et al. Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9. Nat Commun. 2014;5:5560.
Aida T, Nakade S, Sakuma T, Izu Y, Oishi A, Mochida K, et al. Gene cassette knock-in in mammalian cells and zygotes by enhanced MMEJ. BMC Genomics. 2016;17:979.
Renaud JB, Boix C, Charpentier M, De Cian A, Cochennec J, Duvernois-Berthet E, et al. Improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR-Cas9 Nucleases. Cell Rep. 2016;14:2263–72.
Fodde R, Edelmann W, Yang K, van Leeuwen C, Carlson C, Renault B, et al. A targeted chain-termination mutation in the mouse Apc gene results in multiple intestinal tumors. Proc Natl Acad Sci USA. 1994;91:8969–73.
Amos-Landgraf JM, Kwong LN, Kendziorski CM, Reichelderfer M, Torrealba J, Weichert J, et al. A target-selected Apc-mutant rat kindred enhances the modeling of familial human colon cancer. Proc Natl Acad Sci USA. 2007;104:4036–41.
Mashimo T, Takizawa A, Kobayashi J, Kunihiro Y, Yoshimi K, Ishida S, et al. Generation and characterization of severe combined immunodeficiency rats. Cell Rep. 2012;2:685–94.
Bosma GC, Fried M, Custer RP, Carroll A, Gibson DM, Bosma MJ. Evidence of functional lymphocytes in some (leaky) scid mice. J Exp Med. 1988;167:1016–33.
Shultz LD, Brehm MA, Garcia-Martinez JV, Greiner DL. Humanized mice for immune system investigation: progress, promise and challenges. Nat Rev Immunol. 2012;12:786–98.
Quek H, Luff J, Cheung K, Kozlov S, Gatei M, Lee CS, et al. A rat model of ataxia-telangiectasia: evidence for a neurodegenerative phenotype. Hum Mol Genet. 2017;26:109–23.
Ebihara C, Ebihara K, Aizawa-Abe M, Mashimo T, Tomita T, Zhao M, et al. Seipin is necessary for normal brain development and spermatogenesis in addition to adipogenesis. Hum Mol Genet. 2015;24:4238–49.
Nguyen TL, Vieira-Silva S, Liston A, Raes J. How informative is the mouse for human gut microbiota research? Dis Model Mech. 2015;8:1–16.
Acknowledgements
We are supported by a Grant-in-aid for Scientific Research from the Japan Society for the Promotion of Science (25890011 and 16K18402 to KY, 16H06276 and 26290033 to MT). We would like to thank our staff in the animal facility of Osaka University for discussions regarding this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yoshimi, K., Mashimo, T. Application of genome editing technologies in rats for human disease models. J Hum Genet 63, 115–123 (2018). https://doi.org/10.1038/s10038-017-0346-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0346-2
This article is cited by
-
Stimuli-responsive nanoformulations for CRISPR-Cas9 genome editing
Journal of Nanobiotechnology (2022)
-
Creation of X-linked Alport syndrome rat model with Col4a5 deficiency
Scientific Reports (2021)
-
Applications and considerations for the use of genetically engineered mouse models in drug development
Cell and Tissue Research (2020)
-
Successful production of genome-edited rats by the rGONAD method
BMC Biotechnology (2018)