Abstract
Hematopoietic stem cells (HSCs) are pluripotent cells that give rise to all of the circulating blood cell types. Their unique ability to self-renew while generating differentiated daughter cells permits HSCs to sustain blood cell production throughout life. In mammals, the pool of HSCs shifts from early sites in the aorta-gonad-mesonephros region and the placenta to the fetal liver and ultimately bone marrow. During the past decade, a map of transcriptional activators and repressors that regulate gene expression in HSCs, their precursors and their progeny, at distinct stages of development has been drafted. These factors control a program that first establishes the pool of HSCs in the fetus, and later guides decisions between quiescence, self-renewal, and lineage commitment with progressive differentiation to maintain homeostasis. Continuing studies of the regulatory mechanisms that control HSC gene expression followed by the identification of specific loci that are activated or silenced during the life of an HSC will help to further elucidate longstanding issues in HSC decisions to self-renew or to differentiate, and to define the origins of and connections between distinct HSC pools and their precursors.
Similar content being viewed by others
Main
All blood cells in the body derive from pluripotent hematopoietic stem cells (HSCs) through a progression of commitment and differentiation that begins with the generation of multi-potential and lineage-restricted progenitors. HSCs are unique compared with all other hematopoietic cells, as they have the ability to self-renew, and can thereby sustain blood cell production throughout a lifetime (1). To match self-renewal with the production of mature blood cells from physiologic demand, an HSC must monitor and continuously choose between quiescence, self-renewal, and lineage differentiation. Importantly, self-renewal of an HSC can only be maintained in an appropriate microenvironment, the stem cell niche, which in adults has been identified in distinct locations within the bone marrow (2,3).
Establishment of the pool of self-renewing HSCs during embryonic development is a complex process that involves multiple anatomic sites (Fig. 1) (4,5). The process that initiates the future blood-forming system starts in the primitive streak in gastrulating embryos, when clusters of mesodermal cells commit to becoming blood cells (6). Hematopoietic lineage specification proceeds through a bi-potential precursor cell, the hemangioblast, which in addition to blood cells gives rise to the endothelial cells that comprise the vascular system. The birth of nascent hematopoietic precursors is documented in multiple locations, first in the yolk sac, and slightly later in the mesenchyme that surrounds the large vessels in the embryo (i.e. the aorta-gonad-mesonephros (AGM) region and vitelline and umbilical arteries) and presumably also the placenta, which has recently been identified as a major source of HSCs during fetal life (7–9). After hematopoietic commitment nascent hematopoietic precursors, or pre-HSCs, must develop further into functional HSCs, circulate to the fetal liver, and expand in number, to establish a stockpile of HSCs for the future. The challenge of fetal hematopoiesis is to provide the differentiated blood cells that are needed during fetal life while establishing and conserving self-renewal capacity in HSCs that are on a journey to an ultimate destination in the bone marrow.
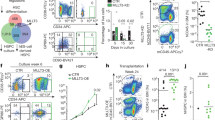
Transcription factors regulating the establishment and maintenance of HSCs. A) The pool of HSCs is generated during development through a complex process that involves several anatomical locations. The sites that are participating in hematopoiesis in mid-gestation mouse embryos are highlighted. B) Hematopoietic development starts in the primitive streak mesoderm and progresses to the yolk sac, AGM, and the placenta, where the first hematopoietic progenitors including HSC precursors are generated. HSC precursors mature into functional HSCs, seed the fetal liver where they expand and differentiate, an ultimately establish steady-state levels in the bone marrow, with a tightly regulated balance between self-renewal, quiescence, and differentiation. The critical transcription factors that have been identified in each of these processes have been highlighted.
HSC development, differentiation, and homeostasis are dictated by transcriptional regulators, which control gene expression under the influence of signals from the microenvironment. Each commitment step requires the activation of lineage-specific genes, while unnecessary and conflicting genes are concomitantly repressed (10). This balance involves the concerted actions of multiple transcriptional activators, repressors, and epigenetic modifiers. Many of the key hematopoietic transcription factors were first discovered in leukemic cells, after which knockout studies in mice revealed important roles for these proto-oncogenes in normal HSC development or lineage differentiation (11). However, as many of these regulators were essential for embryonic blood cell development and survival of the embryo, the requirement of these molecules at later stages of hematopoiesis remained unknown. Establishment of conditional gene targeting strategies made it possible to reassess these factors at later developmental stages and in distinct cellular contexts (12). These studies have greatly extended our understanding about how HSCs develop and function, while changing existing notions about how fate decisions in the hematopoietic system are established and maintained. This review will focus on the hematopoietic transcriptional regulatory mechanisms that play key roles during the development and function of HSCs in the embryo and adult.
COMMITMENT TO THE HEMATOPOIETIC FATE
The hematopoietic program is specified from the mesodermal germ layer shortly after gastrulation in a process that is critically dependent on the bHLH transcription factor SCL/tal1 (Fig. 1B). SCL/tal1 (stem cell leukemia gene) was discovered in multi-lineage and T cell leukemias. Subsequent studies with SCL knockout mice and ES cells demonstrated a complete lack of hematopoietic cells and failed expression of blood-specific genes (13,14).
Contrasting the vital role of SCL during initiation of the hematopoietic program in the fetus, studies using SCL-conditional knockout mice revealed that SCL/tal1 is dispensable for HSC development and function in the adult (15,16). Inactivation of SCL in adult hematopoietic cells by crossing mice that harbored loxP site flanked SCL loci with an interferon-inducible MxCre deleting strain showed that SCL deficient adult bone marrow HSCs are able to engraft, self-renew, and give rise to multi-lineage progeny in vivo. Furthermore, when a Tie2-Cre deleting strain cross was used to inactivate loxP site flanked SCL alleles during early stages of fetal hematopoiesis, establishment of the fetal liver HSC pool was largely unaffected (17). These data showed that HSC development and function is not dependent on SCL expression once specification of the hematopoietic fate in the early embryo has been achieved. Yet, SCL remains essential for maintaining proper differentiation of primitive and definitive erythroid and megakaryocyte lineage cells in the yolk sac, fetal liver, and bone marrow (15,17,18). These findings contradict existing paradigms of lineage specification, in which continuous expression of the transcription factor that specifies a lineage is required to maintain the lineage, and suggests that SCL-induced hematopoietic specification is a stable cell fate that is not challenged with loss of SCL later on in development. The critical target genes that SCL activates or represses during hematopoietic commitment are yet to be defined. Identification of these downstream molecules and the mechanisms by which their expression is subsequently maintained or repressed will be vital for understanding how the hematopoietic program is established and maintained.
EMERGENCE OF DEFINITIVE HSCS IN FETAL HEMOGENIC SITES
After the specification of hematopoietic fate, generation of adult-type HSCs is contingent on the emergence of definitive hematopoietic precursor cells in hemogenic sites in the AGM, vitelline and umbilical vessels, and placenta. This process is critically dependent on the transcription factor Runx1/AML1 (Fig. 1B), a core-binding factor that is also commonly involved in chromosomal translocations in leukemia (19). Primitive erythroid cells develop in the yolk sacs of Runx1/AML1 knockout embryos, indicating that hematopoietic development extends beyond commitment to hematopoietic fate. In contrast, definitive hematopoiesis in Runx1 null embryos is completely disturbed, as was shown by the absence of fetal liver hematopoietic progenitors and hematopoietic clusters in the lumen of the major arteries (19,20). In Runx1 knockout embryos in which the LacZ gene has been introduced into the Runx1 locus, β-galactocidase stained cells are found beneath the endothelium in sites where definitive hematopoietic cells emerge, defining the developmental stage and anatomic sites where Runx1 function is initially required (20). Runx1 continues to be expressed in HSCs during subsequent development and into adult life (21,22). Surprisingly, crossing mice that harbored loxP site flanked Runx1 loci with an interferon-inducible MxCre deleting strain showed that Runx1 is not critical for the function of definitive HSCs in the adult, yet continues to be essential in the lymphoid and megakaryocyte lineages (23,24). These results suggest that Runx1/AML1, like SCL/tal1, is critical for HSC development during a restricted period of time. Runx1 also remains essential at a later stage in HSC development than does SCL, as Tie2-Cre crossed loxP site flanked Runx1 embryos essentially recapitulated the original Runx1 knockout phenotype (25), whereas a comparable cross with loxP site flanked SCL mice showed that an SCL-dependent phase in HSC development had ended before Tie2-Cre mediated inactivation of the SCL loci was complete (17).
HSC MATURATION AND EXPANSION IN THE FETUS
Since the description of the HSC and its central role in producing all other hematopoietic cells in the body, investigators have remained puzzled by the finding that the self-renewing, adult-type HSC is a much later product of fetal hematopoiesis than the short-lived, lineage-restricted or multi-potential progenitors. Indeed, adult-type HSCs that can robustly reconstitute hematopoiesis in irradiated adult recipients appear only around E11, although definitive hematopoietic precursors appear in the same hemogenic sites a few days earlier. The discordant development of various hematopoietic progenitors with in vivo engraftment and self-renewal ability could be explained in part by different hematopoietic programs that separately produce the early progenitors and the self-renewing HSCs. Indeed, this hypothesis seems to apply for yolk sac-derived primitive erythroid precursors. However, a non-exclusive alternative is that a maturation process occurs in which nascent HSC precursors acquire competence to function as stem cells in the adult environment. Such a maturation process is supported by the findings that HSC activity can be detected earlier if more permissive hosts, such as newborn mice or NK-cell deficient mice, are used as transplant recipients (26,27). However, due to a lack of markers and tools for lineage tracing, the relationship between definitive HSCs and various progenitor pools remains difficult to decipher, especially since developing HSCs shift locations multiple times during embryonic development.
Whatever the origin of distinct HSC pools, a molecule that is crucial for establishing a functional HSC is the Mll (mixed lineage leukemia) gene (Fig. 1B), a chromatin modifier from the trithorax group (trxG). Although phenotypically distinct CD41+c-Kit+ definitive hematopoietic precursors develop in vivo in Mll knockout embryos and in vitro in Mll-deficient embryoid bodies (EBs), they proliferate poorly in culture and fail to seed the fetal liver or reconstitute adult bone marrow upon transplantation (28,29). These defects might be explained by a proliferative disadvantage of Mll-deficient hematopoietic cells, or a lack of other qualities conferred by Mll that are essential for HSC trafficking and function. Interestingly, Mll activates the expression of various Hox genes (28), many of which have been independently shown to play a role in the proliferation of normal hematopoietic and leukemic cells. Strikingly, transient over-expression of Hoxb4 in yolk sac and EB-derived hematopoietic precursors conferred these cells with the ability to engraft adult recipients (30) by establishing as yet poorly-defined programs that are essential for HSC function. These programs are otherwise insufficient in EB-derived hematopoietic progenitors, which lack robust in vivo reconstitution and self-renewal ability despite the potential to differentiate into multiple lineages in vitro. Another regulator of Hox-gene expression, Cdx4, was recently shown to cooperate with Hoxb4 to enhance the in vivo repopulation ability and lymphoid reconstitution potential of ES cell-derived hematopoietic precursors (31). These data suggest that Hoxb4 and Cdx4 together facilitate the critical maturation program in early HSC precursors that confers an ability to function as HSCs in vivo (Fig. 1B). The expression of posterior Hoxa6, a7, a9, a10, b9, and c6 genes was increased due to Cdx4/Hoxb4 expression, suggesting that these genes play a role in establishing self-renewal ability in developing HSCs.
A significant challenge for stem cell biologists is to develop methods that facilitate HSC expansion in vitro without the loss of self-renewal ability. Interestingly, over-expression of the Hox genes has beneficial effects on HSC expansion. Over-expression of Hoxb4 in the bone marrow enabled HSC expansion in vitro and increased the in vivo pool size of HSCs after transplantation, without inducing leukemia (32–34). Considering these findings, it was surprising that Hoxb4 deficient HSCs exhibited only subtle hematopoietic defects in vivo (35), although it remains possible that important Hox gene functions were masked in knockout models by redundancies amongst Hox family members. Nevertheless, deletion of the entire Hoxb cluster (Hoxb1 to Hoxb9) was recently shown to be dispensable for hematopoietic reconstitution (36). In contrast, inactivation and over-expression of the myeloid leukemia-related Hoxa9 gene was accompanied by HSC dysfunction and expansion, respectively (37–39).
REGULATION OF SELF-RENEWAL VERSUS DIFFERENTIATION OF HSCS
By the end of gestation in mice, the skeletal system and bone marrow hematopoietic microenvironment have developed sufficiently and the main location of hematopoiesis shifts from the fetal liver to the marrow. At this time, the major expansion phase that establishes the HSC pool during fetal life ends, and the next challenge is to establish a steady state in which the number of HSCs and mature blood cells match and respond to current and future physiologic demands. Regulation of HSC numbers requires both the correct microenvironmental cues and intact signaling and nuclear machinery to recruit mainly quiescent cells from the HSC pool into executing programs for HSC self-renewal versus differentiation (2,3,40).
A transcriptional regulator that has been implicated in regulating the balance between adult HSC self-renewal and differentiation by controlling interactions with the stem cell niche is the proto-oncogene c-Myc (Fig. 1B). Crossing mice that harbored loxP site flanked c-Myc loci with an interferon-inducible MxCre deleting strain caused a severe pancytopenia with HSC accumulation in the bone marrow and increased expression of stromal adhesion molecules (41). The increased number of c-Myc deficient HSCs was not the result of enhanced proliferation or reduced apoptosis but rather from severely impaired hematopoietic differentiation in all lineages tested. Furthermore, c-Myc deficient HSCs failed to reconstitute hematopoiesis when transplanted alone or in combination with wild type HSCs, as c-Myc deficient HSCs accumulated in recipient bone marrow without the generation of lineage committed hematopoietic precursors. In contrast, c-Myc deficient HSC failed to expand but could be induced to differentiate in vitro, indicating distinct c-Myc requirements for HSC proliferation versus differentiation that depended on noncell-autonomous microenvironment conditions. Enforced c-Myc over-expression conversely showed an increased propensity for HSC differentiation, decreased levels of stem cell niche surface adhesion molecules, and a reduced capacity for long-term self-renewal without affecting the overall rate of apoptosis.
Another transcription factor that regulates the commitment of HSCs along distinct differentiation pathways is the ETS family member Pu.1. Disruption of Pu.1 during development impairs the genesis of B-cells, monocytes, neutrophils, and eosinophils that manifest in the fetal liver. This developmental defect occurs during the generation of CLPs (common lymphoid progenitors) and CMPs (common myeloid progenitors) from HSCs, demonstrating that HSCs lacking Pu.1 fail to initiate a commitment to both myeloid and lymphoid lineages (Fig. 1B). Interestingly, inactivation of the loxP site flanked Pu.1 locus after myeloid and lymphoid commitment demonstrated that Pu.1 continues to be essential for myeloid maturation, whereas its requirement for lymphoid development ends after a commitment to becoming a CLP.
Given the known roles of Pu.1 in lineage differentiation, the low level expression of Pu.1 in HSCs was thought to reflect lineage priming, a form of sterile transcription from selective open chromatin that could characterize early oligopotent stages in development without functional impact of the expressed genes (42). However, Pu.1 is also important for maintaining HSC pools (Fig. 1B). Crossing mice that harbored loxP site flanked Pu.1 loci with an interferon-inducible MxCre deleting strain resulted in Pu.1 deficient bone marrow HSCs that could not compete with wild type HSCs in steady-state hematopoiesis or in bone marrow reconstitution assays (42). Furthermore, transplantation of a high dose of Pu.1 deficient fetal liver or bone marrow HSCs showed that Pu.1 deficient HSCs could home into the bone marrow as transient hematopoietic reconstitution was established. However, marrow failure ensued by 6 mo, demonstrating that Pu.1 is required for maintaining the pool of self-renewing HSCs in the bone marrow. Although multiple downstream target genes for Pu.1 during lineage differentiation have been described, the downstream pathways that control HSC maintenance are largely unknown. The versatility of Pu.1 in multiple stages of the hematopoietic hierarchy is likely to be facilitated by the preexisting status of the target cell, where availability of cooperating transcription factors, accessibility of specific genetic loci, and levels of Pu.1 probably dictate the outcome of Pu.1 regulated gene expression.
PATHWAYS REGULATING SURVIVAL OF ADULT HSCS
A transcription factor that is essential for maintaining survival of adult HSCs is Tel/Etv6 (Fig. 1B) (43). Tel/Etv6 is also an ETS family transcriptional repressor that is a frequent target of chromosomal translocations in human leukemias (44). Conventional Tel/Etv6 knockout embryos died by E11 from vascular abnormalities, while blood cell formation in chimeric fetuses was largely unaffected (45,46). However, crossing mice that harbored loxP site flanked Tel/Etv6 loci with an interferon-inducible MxCre deleting strain showed that adult HSCs require continuous Tel/Etv6 expression for survival, which indirectly controls self-renewal and reconstitution capacity. Indeed, loss of Tel/Etv6 in HSCs rapidly resulted in depletion of Tel-deficient bone marrow. The differential requirement of Tel/Etv6 during embryonic and adult hematopoiesis may be a reflection of the young age of fetal HSCs, the development of which does not require a longterm survival program, or unique requirements for survival of HSCs in the bone marrow microenvironment. Strikingly, the requirement for Tel/Etv6 for survival was shown to be a unique property of HSCs. Crossing a Tel/Etv6 loxP site flanked strain with hematopoietic lineage-specific Cre deleting mice demonstrated that Tel was dispensable for the differentiation of most hematopoietic lineages, as the megakaryocyte lineage was the only lineage affected upon loss of Tel/Etv6 in lineage committed progeny of HSCs. Identification of the target genes that are repressed by Tel/Etv6 in HSCs is likely to reveal essential pathways that control the survival of HSCs during adult life.
REGULATION OF SELF-RENEWAL MACHINERY IN NORMAL AND LEUKEMIC HSCS
The self-renewal ability of normal bone marrow is limited to HSCs; however, leukemic cells also acquire a capacity to self-renew, allowing the production of unlimited progeny while differentiation into mature blood cells is inhibited. A key difference between normal and malignant stem cells is the ability for a normal stem cell to respond to instructive signals from its niche, whereas the option to regulate self-renewal is missing from malignant stem cells. Despite this difference, leukemic stem cells and normal HSCs share critical molecular programs to sustain self-renewal (40,47).
The self-renewal of both normal HSCs and leukemic stem cells is dependent on Bmi-1 (Fig. 1B), a zinc finger transcriptional repressor from the polycomb group complex 2 (PRC2). Polycomb complexes are nuclear protein aggregates that control gene silencing by epigenetic modifications that include histone methylation, deacetylation, and possibly histone 2A ubiquitination. PRC2 proteins initiate gene silencing, while a related group of PRC1 proteins are required for the maintenance of gene repression. Bmi-1 was first identified as a cooperating oncogene that collaborates with c-Myc to promote lymphomagenesis (48). Bmi-1 knockout mice generated a normal number of fetal HSCs, whereas the pool of postnatal bone marrow HSCs became exhausted in young adulthood (47,49). Upon transplantation, Bmi-1 deficient fetal liver or bone marrow HSCs re-established a normal pattern of multi-lineage blood cell generation; however, hematopoietic reconstitution was only transient as the transplanted HSC pool was progressively depleted. Interestingly, functional integrity of Bmi-1 null HSCs in the fetal liver could be rescued upon reintroduction of Bmi-1 by retroviral infection, confirming that the original pool of HSCs was generated in the absence of Bmi-1, while Bmi-1 remained essential for HSC maintenance in the bone marrow. These results showed that the molecular mechanisms that control HSC self-renewal and expansion during fetal life are in part different from those that control HSC self-renewal and maintenance in the adult bone marrow. Gene expression profiling of Bmi-1 deficient HSCs showed that silencing of the cell cycle inhibitors p16Ink4a and p19Arf was relieved in the absence of Bmi-1, introducing the hypothesis of premature senescence as a cause for the depletion of Bmi-1 deficient HSCs (49). However, as the cell cycle status and overall apoptosis of Bmi-1 null HSCs was not markedly different from control HSCs, it is likely that other, yet undefined pathways contribute to the self-renewal defect (50). Conversely, over-expression of Bmi-1 enhanced symmetrical division of HSCs, leading to a higher probability of inheriting stemness through cell division and to a striking ex vivo expansion of HSCs (50). Interestingly, Bmi-1 was also required for the maintenance of self-renewal in leukemic HSCs that were transformed by retroviral infection of collaborating oncogenes Hoxa9 and Meis1. Although acute myeloid leukemia formed in Bmi-1 null fetal HSCs, Bmi-1 deficient leukemic cells were unable to transfer disease to syngeneic recipients (47) due to replicative exhaustion and increased apoptosis. Rare high proliferation leukemic stem cell escapees also showed loss of G1 cyclin-dependent kinase inhibitors, suggesting clonal evolution of a cancer stem cell with genetic or epigenetic loss of growth control (47).
COMBATING HSC EXHAUSTION
Replicative stress on bone marrow HSCs from processes like serial transplantation or aging causes a gradual decline in self-renewal capacity (51,52). This can be overcome by over-expression of Ezh2 (enhancer of zeste homolog 2) (Fig. 1B), which is a polycomb group (PcG) protein and member of PRC2 (53). Indeed, serial transplantation of retrovirally-transduced Ezh2 over-expressing HSCs supported long-term re-populating capacity and an increase in the HSC pool size without malignant transformation. Consistent with the notion that Ezh2 helps to maintain HSC renewal capacity, Ezh2 was abundantly expressed in isolated HSCs and then rapidly down-regulated with in vitro HSC differentiation. Furthermore, Ezh2 down-regulation was detected in aged HSCs relative to young HSCs (51). However, the absolute dependence of HSC renewal on Ezh2 expression has not yet been determined with gene ablation or knockdown, an approach that provided the surprising result that Hoxb4 was not essential for HSC renewal despite its ability to promote HSC self-renewal when over-expressed (35).
A transcriptional regulator that appears essential for combating exhaustion of HSCs is Gfi-1 (growth factor independent-1) (Fig. 1B), a SNAG-domain-containing zinc finger transcription repressor that promotes growth factor-independent expansion and malignant transformation of lymphoid cells (54). Gfi-1 null mice are viable at birth, but lack mature neutrophils, while morphologically atypical myeloid cells accumulate in the bone marrow (55). A detailed inspection of the HSC pool demonstrated marked defects in Gfi-1 null HSCs. A severe competitive disadvantage of Gfi-1 null HSCs in transplantation assays and loss of Gfi-1 null hematopoietic cells in chimeric mice shortly after birth was observed (56). Surprisingly, the frequency of phenotypic HSCs in Gfi-1 null bone marrow was increased compared with controls, although the functionality in competitive assays was severely reduced. It can be concluded that in contrast to Bmi-1 and Hoxb4, whose expression promotes HSC proliferation and self-renewal, Gfi-1 acts to suppress HSC proliferation, combating HSC exhaustion by replicative stress, and preserving HSC functional integrity for sustained hematopoiesis. A possible mechanism contributing to Gfi-1 growth restraint in HSCs is through the down-regulation of the cyclin-dependent kinase inhibitor and G1 checkpoint regulator p21Cip1/Waf1. p21 knockout mice exhibit loss of the HSC pool through serial transplantation; however, as the HSC defect in Gfi-1 null mice is much more severe than in p21 null mice, additional pathways are likely to be selectively regulated by Gfi-1. Furthermore, since Gfi-1 regulates proliferation of HSCs and myeloid cells differentially compared with lymphoid cells, the transcriptional targets for Gfi-1 are likely to be cell type and context dependent.
REGULATION OF HSCS BY EPIGENETIC MODIFICATIONS
Pluripotency as well as lineage differentiation depend upon specific chromatin organization, which is required for establishing and maintaining gene expression programs. Epigenetic modifications mainly target the protein or DNA components of chromatin to heritably modify patterns of gene expression. Post-translational modifications of exposed histone tail motifs create an intricate histone code, which, when combined with DNA methylation patterns form the major types of epigenetic regulation that contribute to environmentally responsive gene expression programs to generate HSCs and regulate their quiescence, self-renewal, or differentiation. The control of HSCs by trxG and PcG chromatin modifying proteins in the fetal liver and bone marrow confirms a critical role for epigenetic modifications in regulating HSC development and function. Consistent with this notion is the recent finding that the PcG protein Ezh2, which regulates adult HSC self-renewal capacity, binds DNA methyltransferases (DNMTs) and is required for DNA methylation and gene silencing (57). The transcription factor Pu.1 also can participate in a co-repressor complex with histone deacetylase (HDAC) activity and directly binds DNMTs 3a and 3b to direct DNA methylation (58,59). Furthermore, repression of gene expression by c-Myc is mediated by both passive interference with transcriptional activator binding and by direct recruitment of DNMT3a to specific gene loci (60). Combined, an emerging link between histone and DNA modifying enzymes and key transcription factors that control HSC development and function strongly suggest a critical role for chromatin configuration in establishing and maintaining HSC gene expression programs. To more clearly understand these processes, global patterns for DNA methylation and histone modifications must be established for key genes at different stages and locations of HSC development. Elucidation of genome-wide and site-specific epigenetic markings may even provide new approaches to trace connections between distinct pools of HSCs and their precursors throughout early embryonic development into adulthood.
Interestingly, the effects of compounds that alter chromatin further implicate epigenetic modifications in the control of HSC development and function. Human CD34+ bone marrow cells exposed in vitro to a combination of 5-aza-2′-deoxycytidine (5aza), a DNMT inhibiting nucleotide analogue, and trichostatin A (TSA), a HDAC inhibitor, showed an increased proliferative potential and enhanced xenograft reconstitution capacity in immunodeficient NOD/SCID mice (61). Valproic acid, a distinct type of HDAC inhibitor used clinically to treat specific neurologic disorders, augmented the expansion of cytokine-treated human CD34+ HSC isolated from cord blood, from bone marrow, or from mobilized peripheral blood in vitro, which could provide increased cell numbers for transplantation or gene and stem cell therapies (62). Valproate exposure increased the proliferation and self-renewal of human and mouse bone marrow HSCs and augmented the histone acetylation and expression of the HOXB4 gene, which may have a causative role in expanding the bone marrow HSC pool in vivo and in vitro (62,63). Valproate has an opposite effect on leukemic stem cells, causing reduced proliferation and enhancing differentiation, suggesting the intriguing possibility of simultaneously combating leukemia while enhancing the repopulation capacity of HSCs in vivo. Although the use of chromatin modifying compounds does not yet allow manipulation of specific loci, these findings encourage further studies to identify the target genes and their regulators that are involved in epigenetic control of HSC biology.
CONCLUDING REMARKS
While rapid progress in identifying the transcriptional regulators that control HSCs at various stages of development has been made, much remains to be done to identify the critical target genes that ultimately facilitate HSC generation and function, and to understand how these transcription factors form an interactive network that activate and silence the correct target genes at the right time. Furthermore, very little is known about the relative roles epigenetic modifiers and patterns of epigenetic modifications that control the establishment and maintenance of hematopoietic fate decisions, and facilitate maintenance of the perfect balance between states of HSC quiescence, self-renewal and differentiation to sustain blood cell homeostasis. Although global epigenetic regulatory mechanisms are the least well-defined controllers of hematopoiesis, their importance is clearly indicated by the functional outcomes that result from exposure to chromatin modifying compounds. Expanding our knowledge of the interrelationships between HSC transcriptional and epigenetic programs will provide an increasingly integrated model that can be used to address key unresolved issues in HSC physiology, which include the lineage tracing of precursors to adult-type HSCs during development, and defining the mechanism for establishing and maintaining stemness in the hematopoietic system.
Abbreviations
- AGM:
-
aorta-gonad-mesonephros
- DNMT:
-
DNA methyltransferase
- EB:
-
embryoid body
- HDAC:
-
histone deacetylase
- HSC:
-
hematopoietic stem cell
- PcG:
-
polycomb group
- PRC:
-
polycomb group complex
- trxG:
-
trithorax group
References
Morrison SJ, Weissman IL 1994 The long-term repopulating subset of hematopoietic stem cells is deterministic and isolatable by phenotype. Immunity 1: 661–673
Arai F, Hirao A, Suda T 2005 Regulation of hematopoietic stem cells by the niche. Trends Cardiovasc Med 15: 75–79
Suda T, Arai F, Hirao A 2005 Hematopoietic stem cells and their niche. Trends Immunol 26: 426–433
Tavian M, Peault B 2005 The changing cellular environments of hematopoiesis in human development in utero. Exp Hematol 33: 1062–1069
Jaffredo T, Nottingham W, Liddiard K, Bollerot K, Pouget C, de Bruijn M 2005 From hemangioblast to hematopoietic stem cell: an endothelial connection?. Exp Hematol 33: 1029–1040
Huber TL, Kouskoff V, Fehling HJ, Palis J, Keller G 2004 Haemangioblast commitment is initiated in the primitive streak of the mouse embryo. Nature 432: 625–630
Alvarez-Silva M, Belo-Diabangouaya P, Salaun J, Dieterlen-Lievre F 2003 Mouse placenta is a major hematopoietic organ. Development 130: 5437–5444
Gekas C, Dieterlen-Lievre F, Orkin SH, Mikkola HK 2005 The placenta is a niche for hematopoietic stem cells. Dev Cell 8: 365–375
Ottersbach K, Dzierzak E 2005 The murine placenta contains hematopoietic stem cells within the vascular labyrinth region. Dev Cell 8: 377–387
Nerlov C, Querfurth E, Kulessa H, Graf T 2000 GATA-1 interacts with the myeloid PU. 1 transcription factor and represses PU. 1-dependent transcription. Blood 95: 2543–2551
Orkin SH, Porcher C, Fujiwara Y, Visvader J, Wang LC 1999 Intersections between blood cell development and leukemia genes. Cancer Res 59: 1784s–1787s; discussion 1788s
Mikkola HK, Orkin SH 2005 Gene targeting and transgenic strategies for the analysis of hematopoietic development in the mouse. Methods Mol Med 105: 3–22
Shivdasani RA, Mayer EL, Orkin SH 1995 Absence of blood formation in mice lacking the T-cell leukaemia oncoprotein tal-1/SCL. Nature 373: 432–434
Robertson SM, Kennedy M, Shannon JM, Keller G 2000 A transitional stage in the commitment of mesoderm to hematopoiesis requiring the transcription factor SCL/tal-1. Development 127: 2447–2459
Mikkola HK, Klintman J, Yang H, Hock H, Schlaeger TM, Fujiwara Y, Orkin SH 2003 Haematopoietic stem cells retain long-term repopulating activity and multipotency in the absence of stem-cell leukaemia SCL/tal-1 gene. Nature 421: 547–551
Hall MA, Curtis DJ, Metcalf D, Elefanty AG, Sourris K, Robb L, Gothert JR, Jane SM, Begley CG 2003 The critical regulator of embryonic hematopoiesis, SCL, is vital in the adult for megakaryopoiesis, erythropoiesis, and lineage choice in CFU-S12. Proc Natl Acad Sci U S A 100: 992–997
Schlaeger TM, Mikkola HK, Gekas C, Helgadottir HB, Orkin SH 2005 Tie2Cre-mediated gene ablation defines the stem-cell leukemia gene (SCL/tal1)-dependent window during hematopoietic stem-cell development. Blood 105: 3871–3874
Hall MA, Slater NJ, Begley CG, Salmon JM, Van Stekelenburg LJ, McCormack MP, Jane SM, Curtis DJ 2005 Functional but abnormal adult erythropoiesis in the absence of the stem cell leukemia gene. Mol Cell Biol 25: 6355–6362
Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR 1996 AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84: 321–330
North T, Gu TL, Stacy T, Wang Q, Howard L, Binder M, Marin-Padilla M, Speck NA 1999 Cbfa2 is required for the formation of intra-aortic hematopoietic clusters. Development 126: 2563–2575
North TE, Stacy T, Matheny CJ, Speck NA, de Bruijn MF 2004 Runx1 is expressed in adult mouse hematopoietic stem cells and differentiating myeloid and lymphoid cells, but not in maturing erythroid cells. Stem Cells 22: 158–168
North TE, de Bruijn MF, Stacy T, Talebian L, Lind E, Robin C, Binder M, Dzierzak E, Speck NA 2002 Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo. Immunity 16: 661–672
Ichikawa M, Asai T, Saito T, Seo S, Yamazaki I, Yamagata T, Mitani K, Chiba S, Ogawa S, Kurokawa M, Hirai H 2004 AML-1 is required for megakaryocytic maturation and lymphocytic differentiation, but not for maintenance of hematopoietic stem cells in adult hematopoiesis. Nat Med 10: 299–304
Growney JD, Shigematsu H, Li Z, Lee BH, Adelsperger J, Rowan R, Curley DP, Kutok JL, Akashi K, Williams IR, Speck NA, Gilliland DG 2005 Loss of Runx1 perturbs adult hematopoiesis and is associated with a myeloproliferative phenotype. Blood 106: 494–504
Li Z, Chen MJ, Stacy T, Speck NA 2005 Runx1 function in hematopoiesis is required in cells that express Tek. Blood 107: 106–110
Yoder MC, Hiatt K, Mukherjee P 1997 In vivo repopulating hematopoietic stem cells are present in the murine yolk sac at day 9.0 postcoitus. Proc Natl Acad Sci U S A 94: 6776–6780
Cumano A, Ferraz JC, Klaine M, Di Santo JP, Godin I 2001 Intraembryonic, but not yolk sac hematopoietic precursors, isolated before circulation, provide long-term multi-lineage reconstitution. Immunity 15: 477–485
Ernst P, Mabon M, Davidson AJ, Zon LI, Korsmeyer SJ 2004 An Mll-dependent Hox program drives hematopoietic progenitor expansion. Curr Biol 14: 2063–2069
Ernst P, Fisher JK, Avery W, Wade S, Foy D, Korsmeyer SJ 2004 Definitive hematopoiesis requires the mixed-lineage leukemia gene. Dev Cell 6: 437–443
Kyba M, Perlingeiro RC, Daley GQ 2002 HoxB4 confers definitive lymphoid-myeloid engraftment potential on embryonic stem cell and yolk sac hematopoietic progenitors. Cell 109: 29–37
Wang Y, Yates F, Naveiras O, Ernst P, Daley GQ 2005 Embryonic stem cell-derived hematopoietic stem cells. Proc Natl Acad Sci U S A 102: 19081–19086
Antonchuk J, Sauvageau G, Humphries RK 2002 HOXB4-induced expansion of adult hematopoietic stem cells ex vivo. Cell 109: 39–45
Antonchuk J, Sauvageau G, Humphries RK 2001 HOXB4 over-expression mediates very rapid stem cell regeneration and competitive hematopoietic repopulation. Exp Hematol 29: 1125–1134
Thorsteinsdottir U, Sauvageau G, Humphries RK 1999 Enhanced in vivo regenerative potential of HOXB4-transduced hematopoietic stem cells with regulation of their pool size. Blood 94: 2605–2612
Brun AC, Bjornsson JM, Magnusson M, Larsson N, Leveen P, Ehinger M, Nilsson E, Karlsson S 2004 Hoxb4-deficient mice undergo normal hematopoietic development but exhibit a mild proliferation defect in hematopoietic stem cells. Blood 103: 4126–4133
Bijl J, Thompson A, Ramirez-Solis R, Krosl J, Grier DG, Lawrence HJ, Sauvageau G 2005 Analysis of HSC activity and compensatory Hox gene expression profile in Hoxb cluster mutant fetal liver cells. Blood Dec 8, [Epub ahead of print]
Lawrence HJ, Christensen J, Fong S, Hu YL, Weissman I, Sauvageau G, Humphries RK, Largman C 2005 Loss of expression of the Hoxa-9 homeobox gene impairs the proliferation and repopulating ability of hematopoietic stem cells. Blood 106: 3988–3994
Lawrence HJ, Helgason CD, Sauvageau G, Fong S, Izon DJ, Humphries RK, Largman C 1997 Mice bearing a targeted interruption of the homeobox gene HOXA9 have defects in myeloid, erythroid, and lymphoid hematopoiesis. Blood 89: 1922–1930
Thorsteinsdottir U, Mamo A, Kroon E, Jerome L, Bijl J, Lawrence HJ, Humphries K, Sauvageau G 2002 Over-expression of the myeloid leukemia-associated Hoxa9 gene in bone marrow cells induces stem cell expansion. Blood 99: 121–129
Lessard J, Faubert A, Sauvageau G 2004 Genetic programs regulating HSC specification, maintenance and expansion. Oncogene 23: 7199–7209
Wilson A, Murphy MJ, Oskarsson T, Kaloulis K, Bettess MD, Oser GM, Pasche AC, Knabenhans C, Macdonald HR, Trumpp A 2004 c-Myc controls the balance between hematopoietic stem cell self-renewal and differentiation. Genes Dev 18: 2747–2763
Iwasaki H, Somoza C, Shigematsu H, Duprez EA, Iwasaki-Arai J, Mizuno S, Arinobu Y, Geary K, Zhang P, Dayaram T, Fenyus ML, Elf S, Chan S, Kastner P, Huettner CS, Murray R, Tenen DG, Akashi K 2005 Distinctive and indispensable roles of PU. 1 in maintenance of hematopoietic stem cells and their differentiation. Blood 106: 1590–1600
Hock H, Meade E, Medeiros S, Schindler JW, Valk PJ, Fujiwara Y, Orkin SH 2004 Tel/Etv6 is an essential and selective regulator of adult hematopoietic stem cell survival. Genes Dev 18: 2336–2341
Golub TR, McLean T, Stegmaier K, Carroll M, Tomasson M, Gilliland DG 1996 The TEL gene and human leukemia. Biochim Biophys Acta 1288: M7–M10
Wang LC, Swat W, Fujiwara Y, Davidson L, Visvader J, Kuo F, Alt FW, Gilliland DG, Golub TR, Orkin SH 1998 The TEL/ETV6 gene is required specifically for hematopoiesis in the bone marrow. Genes Dev 12: 2392–2402
Wang LC, Kuo F, Fujiwara Y, Gilliland DG, Golub TR, Orkin SH 1997 Yolk sac angiogenic defect and intra-embryonic apoptosis in mice lacking the Ets-related factor TEL. EMBO J 16: 4374–4383
Lessard J, Sauvageau G 2003 Bmi-1 determines the proliferative capacity of normal and leukaemic stem cells. Nature 423: 255–260
Haupt Y, Alexander WS, Barri G, Klinken SP, Adams JM 1991 Novel zinc finger gene implicated as myc collaborator by retrovirally accelerated lymphomagenesis in E mu-myc transgenic mice. Cell 65: 753–763
Park IK, Qian D, Kiel M, Becker MW, Pihalja M, Weissman IL, Morrison SJ, Clarke MF 2003 Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature 423: 302–305
Iwama A, Oguro H, Negishi M, Kato Y, Morita Y, Tsukui H, Ema H, Kamijo T, Katoh-Fukui Y, Koseki H, van Lohuizen M, Nakauchi H 2004 Enhanced self-renewal of hematopoietic stem cells mediated by the polycomb gene product Bmi-1. Immunity 21: 843–851
Rossi DJ, Bryder D, Zahn JM, Ahlenius H, Sonu R, Wagers AJ, Weissman IL 2005 Cell intrinsic alterations underlie hematopoietic stem cell aging. Proc Natl Acad Sci U S A 102: 9194–9199
Kamminga LM, van Os R, Ausema A, Noach EJ, Weersing E, Dontje B, Vellenga E, de Haan G 2005 Impaired hematopoietic stem cell functioning after serial transplantation and during normal aging. Stem Cells 23: 82–92
Kamminga LM, Bystrykh LV, de Boer A, Houwer S, Douma J, Weersing E, Dontje B, de Haan G 2005 The polycomb group gene Ezh2 prevents hematopoietic stem cell exhaustion. Blood Nov 17, [Epub ahead of print]
Gilks CB, Bear SE, Grimes HL, Tsichlis PN 1993 Progression of interleukin-2 (IL-2)-dependent rat T cell lymphoma lines to IL-2-independent growth following activation of a gene (GFI-1) encoding a novel zinc finger protein. Mol Cell Biol 13: 1759–1768
Hock H, Hamblen MJ, Rooke HM, Traver D, Bronson RT, Cameron S, Orkin SH 2003 Intrinsic requirement for zinc finger transcription factor GFI-1 in neutrophil differentiation. Immunity 18: 109–120
Hock H, Hamblen MJ, Rooke HM, Schindler JW, Saleque S, Fujiwara Y, Orkin SH 2004 GFI-1 restricts proliferation and preserves functional integrity of haematopoietic stem cells. Nature 431: 1002–1007
Vire, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden JM, Bollen M, Esteller M, Di Croce L, de Launoit Y, Fuks F 2005 The Polycomb group protein EZH2 directly controls DNA methylation. Nature Dec 14, [Epub ahead of print]
Suzuki M, Yamada T, Kihara Negishi F, Sakurai T, Hara E, Tenen DG, Hozumi N, Oikawa T 2005 Site-specific DNA methylation by a complex of PU. 1 and Dnmt3a/b. Oncogene Dec 5, [Epub ahead of print]
Kihara-Negishi F, Yamamoto H, Suzuki M, Yamada T, Sakurai T, Tamura T, Oikawa T 2001 In vivo complex formation of PU. 1 with HDAC1 associated with PU. 1-mediated transcriptional repression. Oncogene 20: 6039–6047
Brenner C, Deplus R, Didelot C, Loriot A, Vire E, De Smet C, Gutierrez A, Danovi D, Bernard D, Boon T, Pelicci PG, Amati B, Kouzarides T, de Launoit Y, Di Croce L, Fuks F 2005 Myc represses transcription through recruitment of DNA methyltransferase corepressor. EMBO J 24: 336–346
Milhem M, Mahmud N, Lavelle D, Araki H, DeSimone J, Saunthararajah Y, Hoffman R 2004 Modification of hematopoietic stem cell fate by 5aza 2′deoxycytidine and trichostatin A. Blood 103: 4102–4110
De Felice L, Tatarelli C, Mascolo MG, Gregorj C, Agostini F, Fiorini R, Gelmetti V, Pascale S, Padula F, Petrucci MT, Arcese W, Nervi C 2005 Histone deacetylase inhibitor valproic acid enhances the cytokine-induced expansion of human hematopoietic stem cells. Cancer Res 65: 1505–1513
Bug G, Gul H, Schwarz K, Pfeifer H, Kampfmann M, Zheng X, Beissert T, Boehrer S, Hoelzer D, Ottmann OG, Ruthardt M 2005 Valproic acid stimulates proliferation and self-renewal of hematopoietic stem cells. Cancer Res 65: 2537–2541
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by: NIH grants GM073981, CA90571, CA107300; the Margaret E. Early Medical Research Trust; and CMISE, a NASA URETI Institute (NCC 2-1364) (M.A.T.). M.A.T. is a Scholar of the Leukemia and Lymphoma Society. Also supported by NIH grant DK069659 and a Harvard Stem Cell Institute seed grant [H.K.A.M.].
Rights and permissions
About this article
Cite this article
Teitell, M., Mikkola, H. Transcriptional Activators, Repressors, and Epigenetic Modifiers Controlling Hematopoietic Stem Cell Development. Pediatr Res 59 (Suppl 4), 33–39 (2006). https://doi.org/10.1203/01.pdr.0000205155.26315.c7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/01.pdr.0000205155.26315.c7
This article is cited by
-
Direct induction of haematoendothelial programs in human pluripotent stem cells by transcriptional regulators
Nature Communications (2014)