Abstract
The expression of epidermal growth factor receptor (EGFR/ERBB1/HER1) is implicated in the progress of numerous cancers, a feature that has been exploited in the development of EGFR antibodies and EGFR tyrosine kinase inhibitors as anti-cancer drugs. However, EGFR also has important normal cellular functions, leading to serious side effects when EGFR is inhibited. One damaging characteristic of many oncogenes is the ability to be expressed in the hypoxic conditions associated with the tumour interior. It has previously been demonstrated that expression of EGFR is maintained in hypoxic conditions via an unknown mechanism of translational control, despite global translation rates generally being attenuated under hypoxic conditions. In this report, we demonstrate that the human EGFR 5′ untranslated region (UTR) sequence can initiate the expression of a downstream open reading frame via an internal ribosome entry site (IRES). We show that this effect is not due to either cryptic promoter activity or splicing events. We have investigated the requirement of the EGFR IRES for eukaryotic initiation factor 4A (eIF4A), which is an RNA helicase responsible for processing RNA secondary structure as part of translation initiation. Treatment with hippuristanol (a potent inhibitor of eIF4A) caused a decrease in EGFR 5′ UTR-driven reporter activity and also a reduction in EGFR protein level. Importantly, we show that expression of a reporter gene under the control of the EGFR IRES is maintained under hypoxic conditions despite a fall in global translation rates.
Similar content being viewed by others
Introduction
Epidermal growth factor receptor (EGFR, also ErbB-1 or HER1) is an important drug target and prognostic indicator in many cancers. It is a transmembrane glycoprotein tyrosine kinase. Its primary function is to stimulate Akt-, MAPK- or JNK-mediated cellular proliferation in response to a range of ligands including transforming growth factor alpha and the family of EGFs (reviewed in the study by Oda et al.1).
Overexpression of EGFR has been strongly linked to poor prognosis in a large number of cancers including breast, head, neck, ovarian, cervical, bladder and oesophageal cancers (reviewed in the study by Nicholson et al.2). Mutations in EGFR have also been shown to be important, for example a subset of tumours possess a constitutively active truncated version of EGFR.3 Although there is a strong relationship between the frequency of this mutation and poor prognosis, possibly more importantly is overexpression of EGFR with nearly half of glioblastomas displaying significantly elevated levels of the wild-type receptor.3, 4, 5, 6, 7, 8, 9 A major mechanism by which EGFR is overexpressed is gene amplification, as demonstrated in colorectal, pulmonary, bile duct and soft tissue cancers.10, 11, 12, 13 In each of these instances however, amplification was not the only cause of the overexpression, with a marked discrepancy between gene dosage and EGFR protein levels.14 While a proportion of this overexpression may be attributable to mutations that cause the transcriptional upregulation of the EGFR gene,15, 16, 17, 18 it has also been shown that that EGFR protein levels are upregulated in response to both hypoxia and the activation of hypoxia-inducible factor 2α without observing either mutational events or changes in EGFR mRNA levels.19 Increased EGFR activity is also associated with Alzheimer’s disease, and inhibiting EGFR reverses amyloid beta-induced memory loss in mice.20 The untranslated regions (UTRs) of an mRNA have major roles in the translational control of its expression. While the 3′ UTR of EGFR has been investigated and shown to be a target for microRNA-induced suppression under certain conditions, the 5′ UTR remains largely unstudied.21
In general, cells respond to hypoxia by decreasing protein synthesis rates.22, 23 One mechanism used to accomplish this in the short term involves the phosphorylation of the translation initiation factor eIF2α. When hypo-phosphorylated, the function of eIF2α is to assist in the binding of the initiator transfer RNA to the 40S ribosomal subunit by forming a ternary complex with GTP. Phosphorylation of eIF2α inhibits this process, thereby acting as a brake on global translation rates.24, 25
Since internal ribosome entry site (IRES)-mediated translation initiation does not involve the binding of the 5′ cap, it is favoured under certain conditions, like hypoxia, that inhibit eukaryotic initiation factor 4E (eIF4E) function.26, 27, 28 Despite this reduced requirement for eIF4E, it has been demonstrated that the IRESs belonging to the human genes c-myc, N-myc and BiP have a strong requirement for eIF4A function for their expression.29, 30, 31 It has been suggested that this requirement indicates that the structure of these IRESs needs ‘remodelling’ by eIF4A before they are able to function, similar to the processing required by the encephalomyocarditis virus IRES.30, 32, 33, 34
We tested the EGFR 5′ UTR for IRES activity using a dicistronic reporter system and found that it was able to initiate the translation of the downstream cistron. Experiments on the EGFR 5′ UTR reporter using the eIF4A inhibitor hippuristanol found that the EGFR IRES has a high requirement for eIF4A. Finally, we show that the EGFR 5′ UTR allows hypoxic expression of a downstream cistron under conditions where global translation is compromised. These findings strongly support the idea that the induction of EGFR expression in response to hypoxic conditions is attributable to the presence of a previously unidentified IRES within its 5′ UTR.
Results and Discussion
It has previously been shown that EGFR protein levels are maintained under hypoxic conditions despite a global decrease in protein synthesis rates, but the mechanism responsible is not known.19 Moreover, EGFR has recently been shown to be a regulator of hypoxic microRNA maturation through phosphorylation of AGO2.35
The EGFR 5′ UTR was cloned between the Renilla and firefly luciferase open reading frames of pRF and the resultant construct (pR-EGFR-F) was transfected into human neuroblastoma-derived SH-SY5Y cells. Parallel control transfections of pRF (a negative control lacking any IRES element36, 37), pRMF (containing the c-myc IRES36) and pR-Tub-F (containing the β tubulin 5′ UTR, which lacks IRES activity). The EGFR 5′ UTR permitted the expression of the downstream cistron in SH-SY5Y cells to a similar extent as the well-validated c-myc 5′ UTR, whereas the 5′ UTR of β tubulin did not (Figure 1a.). This effect was also observed in HeLa, Huh7 and MCF7 cells (data not shown). To determine whether cryptic splicing was occuring (which could lead to functional firefly luciferase transcripts in the absence of IRES activity) Northern analysis was performed using a radio-labelled probe against the firefly luciferase coding region. This confirmed the presence of a single luciferase containing transcript of the appropriate size in the transfected cells (Figure 1b). To confirm that the observed activity was not due to transcription of the firefly luciferase open reading frame driven by a cryptic promoter in the EGFR 5′ UTR sequence, the cytomegalovirus promoter was removed from the dicistronic constructs by restriction digestion, and the resultant promoter-less constructs were transfected into SH-SY5Y cells to confirm the absence of endogenous cryptic promoter activity in any of the 5′ UTR sequences (Figure 1c). We therefore conclude that the expression of the downstream cistron is directed by an IRES in the EGFR 5′ UTR.
The EGFR 5′ UTR sequence permits the expression of the downstream open reading frame of a dicistronic reporter construct without exhibiting cryptic promoter or splicing activity. The UTRs of β tubulin and EGFR were synthesised by GenScript (Piscataway, NJ, USA) to match the nucleotide sequences with accession numbers NM_178014.2 and NM_201283.1, respectively. These sequences were amplified by PCR using primers containing SpeI restriction sites upstream and NcoI sites downstream, the UTRs were then cloned into pRF between these sites and the resulting plasmid was termed pR-EGFR-F. The dicistronic reporter containing the c-myc 5′ UTR (pRMF) was as described in a previous paper.36 The promoter-less version of the dicistronic reporter was created by cloning the two luciferase cistrons, flanking the EGFR 5′ UTR into a plasmid, which did not contain a promoter sequence. The promoter-less version of the monocistronic reporter was made by cloning the EGFR 5′ UTR into p15hp in place of the hairpin39 and then disabling the cytomegalovirus promoter by AseI digest. (a, c) Twenty-four-well plates were seeded with SH-SY5Y cells at a density of 50 000 cells/well. The following day, cells were transfected with 200 ng/well of the plasmids described above using FuGene 6 (Roche, Mannheim, Germany). The growth medium was changed after 6 h and the cells were cultured for the following 24 h. Luciferase expression within the cells was then quantified using a Dual Luciferase Assay Kit (Promega, Madison, WI, USA) following manufacturer’s instructions. Mean and s.d. of at least three replicates are shown. (b) A Northern blot was performed on lysates from HeLa cells previously transfected with pRF, pR-EGFR-F or pRMF using a probe complementary to the firefly luciferase open reading frame. The dicistronic EGFR reporter plasmid generates a single mRNA.
Having identified an IRES in the EGFR 5′ UTR, we wished to determine whether this could further explain previous observations that increases in EGFR protein can occur without parallel increases in mRNA.19 Hypoxic conditions (1% O2) caused a reduction in control Renilla luciferase expression to ~50% of its control value while the expression of the firefly open reading frames preceded by the EGFR and c-myc 5′ UTR sequences were maintained (Figure 2). The response of the c-myc IRES to hypoxia has been documented previously and it is included here as a positive control.38
Expression of a reporter gene preceded by the 5′ UTR of EGFR is maintained under hypoxic conditions despite a fall in control reporter expression. (a) Twenty-four-well plates were seeded with SH-SY5Y cells at a density of 50 000 cells/well. The following day, cells were transfected with 200 ng/well of the plasmids described in the legend of Figure 1 using FuGene 6. The growth medium was changed after 6 h, and for the following 24 h one of the plates was kept in a control incubator set to atmospheric O2 levels. The other plate was incubated in a ProOx 110 (BioSpherix Ltd., Lacona, NY, USA) hypoxic chamber which O2 restricted to 1%. Luciferase expression within the cells was then quantified using a Dual Luciferase Assay Kit following manufacturer’s instructions. Mean and s.d. of at least three replicates are shown. (b) Six-well plates were seeded with SH-SY5Y cells at a density of 3 × 105 cells/well. The following day, cells were incubated for 24 h at either atmospheric or 1% O2 levels. EGFR and tubulin protein levels were determined by western blotting using primary antibodies ab6012 (Abcam, Cambridge, UK) and sc-7396 (Santa Cruz, Dallas, TX, USA). (c) Four replicates of the blot described in Figure 2b were quantified using ImageJ (US National Institutes of Health, Bethesda, Maryland, USA). The data are presented as EGFR expression normalised to tubulin expression. No significant differences in EGFR expression were seen between normoxic (atmospheric) and hypoxic (1% O2) cells.
To begin to characterise the requirements of the EGFR IRES, we added 10 μM hippuristanol (a potent and specific small molecule inhibitor of eIF4A31) to HeLa cells transfected with the monocistronic and dicistronic reporters. eIF4A is a DEAD-box helicase involved in unwinding secondary structure in 5′ UTRs and is required for efficient cap-independent translation of a number of transcripts.31, 39 This treatment with hippuristanol revealed a significant reduction in EGFR 5′ UTR-mediated reporter expression (P=0.007; Figure 3a). The dependency of the EGFR IRES on eIF4A function is also demonstrated by western blot, with hippuristanol causing a dramatic reduction in the protein level of EGFR, relative to a β tubulin loading control in HeLa (P=0.01; Figure 3b).
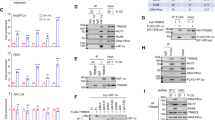
(a) EGFR expression is reduced by hippuristanol treatment and iron treatment via its 5′ UTR. Ten micromolar Hippuristanol (or DMSO) was added to HeLa cells, which were transfected with reporter plasmids 4 h previously. The creation of the reporter plasmid is described previously (Figure 1.). Luciferase expression was quantified 24 h later and firefly values were normalised to control Renilla values (pGL4.80cmv was co-transfected with the monocistronic reporter). Mean and s.d. of at least three replicates are shown. (b) Cellular lysate was also western blotted for EGFR and β tubulin using Abcam antibodies ab2430 and ab6046, respectively; representative blots are pictured. Quantification of three replicates of this experiment was performed using ImageJ and the mean expression levels of EGFR normalised to tubulin were calculated. (c, d) The experiment was repeated as above substituting hippuristanol with 250 μM of ammonium iron citrate.
Iron has been implicated in the function of several IRESes including those in APP,40 HCV41 and cytoplasmic serine hydroxymethyltransferase.42 Iron stress is also known to induce the unfolded protein response,37 which leads to a global reduction in protein synthesis caused by eIF2alpha phosphorylation.23 EGFR protein levels specifically have also been shown to respond to iron, however, the mechanisms responsible have not previously been identified.43, 44 We therefore wished to test whether iron has a specific effect on EGFR IRES function beyond the general downregulation of cap-dependent translation caused by eIF2alpha phosphorylation. SH-SY5Y, HeLa and Huh7 cells were treated with 250 μM ammonium iron citrate or control, and a marked reduction in expression of firefly luciferase expression was observed from reporters driven by the EGFR 5′ UTR (Figure 3c) compared with controls. This inhibition was paralleled by a significant reduction in endogenous EGFR expression following treatment of HeLa cells with 250 μM ammonium iron citrate (P=0.02; Figures 3d and e) compared with expression of beta-tubulin. We have not been able to identify a recognisable iron response element in the EGFR 5′ UTR primary sequence, however, it is apparent that the EGFR 5′ UTR confers susceptibility to iron stress beyond that explained by the modest global decrease in translation or proliferation exemplified by beta-tubulin protein levels (Figure 3d). There are precedents for iron levels influencing IRES activity40, 42 and further work is required to confirm whether similar mechanisms are at work in EGFR translational control.
By introducing a series of upstream out-of-frame (relative to luciferase) AUG start codons into the EGFR 5′ UTR sequence we have identified that the 40S ribosomal subunit enters the IRES between 23 and 56 nucleotide (nt) upstream of the authentic AUG start codon (Figures 3a and b). Introduction of out-of-frame AUGs further than 56 nt from the luciferase start codon have no effect on luciferase expression, indicating that the 40S ribosomal subunit enters at a position downstream; introduction of an out-of-frame AUG at position −20 completely abolishes luciferase expression, suggesting that the ribosome enters before this position. An alignment of refseq45 primate EGFR 5′ UTR sequences using LocARNA46, 47, 48 suggests that the ribosome entry region, between positions −56 and −20, may be located within a conserved stem loop close to the start codon (Figure 4b).
The ribosome enters the EGFR 5′ UTR near the start codon. Five mutant versions of the plasmid pR-EGFR-F were created that introduced AUG start codons at different positions within the EGFR 5′ UTR (mutants are named according to the position of the introduced AUG relative to the wild-type AUG). The introduced AUGs are out-of-frame with the downstream luciferase gene, therefore, if the ribosome bound to the sequence upstream of one of these, a non-functional protein would result and quantified luciferase levels would reduce to background. (a) A 24-well plate was seeded with SH-SY5Y cells. These were allowed to recover overnight before being transfected with 200 ng/well of the mutant or wild-type reporter plasmids. After 24 h, luciferase levels were assayed. Mean and s.d of at least three replicates are shown. (b) Alignment of primate EGFR 5′ UTR refseq sequences using Locarna (Will et al.47, 48 and Smith et al.46) suggests that the ribosome entry site (blue bar) may lie within a conserved stem loop structure near the start codon.
These findings suggest an opportunity to exploit the hypoxic control of EGFR translation to develop new targeted therapeutics. Current therapies such as tyrosine kinase inhibitors or antibodies, which indiscriminately target EGFR, are associated with a number of unpleasant and potentially serious side effects.49 Since hypoxia in otherwise healthy patients is generally restricted to tumour masses,50 targeting only the hypoxic expression of EGFR offers a way of restricting EGFR inhibition to tumour masses, allowing normal expression of EGFR elsewhere in the body and a consequent reduction in systemic side effects. Although hypoxia targeting drug delivery systems have been under development for a number of years,50, 51 understanding the mechanisms that allow hypoxic expression of drug targets is particularly important in allowing us to directly target gene expression specifically in cancer cells.
References
Oda K, Matsuoka Y, Funahashi A, Kitano H . A comprehensive pathway map of epidermal growth factor receptor signaling. Mol Syst Biol 2005; 1: 2005.0010.
Nicholson RI, Gee JMW, Harper ME . EGFR and cancer prognosis. Eur J Cancer 2001; 37: 9–15.
Wikstrand C, Reist CJ, Archer GE, Zalutsky MR, Bigner DD . The class III variant of the epidermal growth factor receptor (EGFRvIII): characterization and utilization as an immunotherapeutic target. J Neurovirol 1998; 4: 148–158.
Bigner SH, Humphrey PA, Wong AJ, Vogelstein B, Mark J, Friedman HS et al. Characterization of the epidermal growth factor receptor in human glioma cell lines and xenografts. Cancer Res 1990; 50: 8017–8022.
Ekstrand AJ, James CD, Cavenee WK, Seliger B, Pettersson RF, Collins VP . Genes for epidermal growth factor receptor, transforming growth factor α, and epidermal growth factor and their expression in human gliomas in vivo. Cancer Res 1991; 51: 2164–2172.
Humphrey PA, Wong AJ, Vogelstein B, Zalutsky MR, Fuller GN, Archer GE et al. Anti-synthetic peptide antibody reacting at the fusion junction of deletion-mutant epidermal growth factor receptors in human glioblastoma. Proc Natl Acad Sci USA 1990; 87: 4207–4211.
Schlegel J, Merdes A, Stumm G, Albert FK, Forsting M, Hynes N et al. Amplification of the epidermal-growth-factor-receptor gene correlates with different growth behaviour in human glioblastoma. Int J Cancer 1994; 56: 72–77.
Schwechheimer K, Huang S, Cavenee WK . EGFR gene amplification—rearrangement in human glioblastomas. Int J Cancer 1995; 62: 145–148.
Yamazaki H, Ohba Y, Tamaoki N, Shibuya M . A deletion mutation within the ligand binding domain is responsible for activation of epidermal growth factor receptor gene in human brain tumors. Cancer Sci 1990; 81: 773–779.
Ooi A, Takehana T, Li X, Suzuki S, Kunitomo K, Iino H et al. Protein overexpression and gene amplification of HER-2 and EGFR in colorectal cancers: an immunohistochemical and fluorescent in situ hybridization study. Mod Pathol 2004; 17: 895–904.
Dacic S, Flanagan M, Cieply K, Ramalingam S, Luketich J, Belani C et al. Significance of EGFR protein expression and gene amplification in non–small cell lung carcinoma. Am J Clin Pathol 2006; 125: 860–865.
Nakazawa K, Dobashi Y, Suzuki S, Fujii H, Takeda Y, Ooi A . Amplification and overexpression of c-erbB-2, epidermal growth factor receptor, and c-met in biliary tract cancers. J Pathol 2005; 206: 356–365.
Dobashi Y, Takei N, Suzuki S, Yoneyama H, Hanawa M, Ooi A . Aberration of epidermal growth factor receptor expression in bone and soft-tissue tumors: protein overexpression, gene amplification and activation of downstream molecules. Mod Pathol 2004; 17: 1497–1505.
Kersting C, Tidow N, Schmidt H, Liedtke C, Neumann J, Boecker W et al. Gene dosage PCR and fluorescence in situ hybridization reveal low frequency of egfr amplifications despite protein overexpression in invasive breast carcinoma. Lab Invest 2004; 84: 582–587.
Haley JD, Waterfield MD . Contributory effects of de novo transcription and premature transcript termination in the regulation of human epidermal growth factor receptor proto-oncogene RNA synthesis. J Biol Chem 1991; 266: 1746–1753.
Maekawa T, Imamoto F, Merlino GT, Pastan I, Ishii S . Cooperative function of two separate enhancers of the human epidermal growth factor receptor proto–oncogene. J Biol Chem 1989; 264: 5488–5494.
Chi DD, Hing AV, Helms C, Steinbrueck T, Mishra SK, Donis-Keller H . Two chromosome 7 dinucleotide repeat polymorphisms at gene loci epidermal growth factor receptor (EGFR) and proα2 (1) collagen (COL1A2). Hum Mol Genet 1992; 1: 135.
Gebhardt F, Zänker KS, Brandt B . Modulation of epidermal growth factor receptor gene transcription by a polymorphic dinucleotide repeat in intron 1. J Biol Chem 1999; 274: 13176–13180.
Franovic A, Gunaratnam L, Smith K, Robert I, Patten D, Lee S . Translational up-regulation of the EGFR by tumor hypoxia provides a nonmutational explanation for its overexpression in human cancer. Proc Natl Acad Sci USA 2007; 104: 13092–13097.
Wang L, Chiang HC, Wu W, Liang B, Xie Z, Yao X et al. Epidermal growth factor receptor is a preferred target for treating amyloid-beta-induced memory loss. Proc Natl Acad Sci USA 2012; 109: 16743–16748.
Weiss GJ, Bemis LT, Nakajima E, Sugita M, Birks DK, Robinson WA et al. EGFR regulation by microRNA in lung cancer: correlation with clinical response and survival to gefitinib and EGFR expression in cell lines. Ann Oncol 2008; 19: 1053–1059.
Pettersen EO, Juul NO, Rønning ØW . Regulation of protein metabolism of human cells during and after acute hypoxia. Cancer Res 1986; 46: 4346–4351.
Koritzinsky M, Rouschop KM, van den Beucken T, Magagnin MG, Savelkouls K, Lambin P et al. Phosphorylation of eIF2alpha is required for mRNA translation inhibition and survival during moderate hypoxia. Radiother Oncol 2007; 83: 353–361.
Koumenis C, Naczki C, Koritzinsky M, Rastani S, Diehl A, Sonenberg N et al. Regulation of protein synthesis by hypoxia via activation of the endoplasmic reticulum kinase PERK and phosphorylation of the translation initiation factor eIF2{alpha}. Mol Cell Biol 2002; 22: 7405–7416.
Koritzinsky M, Magagnin MG, van den Beucken T, Seigneuric R, Savelkouls K, Dostie J et al. Gene expression during acute and prolonged hypoxia is regulated by distinct mechanisms of translational control. EMBO J 2006; 25: 1114–1125.
Hellen CU, Sarnow P . Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev 2001; 15: 1593–1612.
Stoneley M, Willis AE . Cellular internal ribosome entry segments: structures, trans-acting factors and regulation of gene expression. Oncogene 2004; 23: 3200–3207.
Prevot D, Darlix JL, Ohlmann T . Conducting the initiation of protein synthesis: the role of eIF4G. Biol Cell 2003; 95: 141–156.
Thoma C, Bergamini G, Galy B, Hundsdoerfer P, Hentze MW . Enhancement of IRES-mediated translation of the c-myc and BiP mRNAs by the poly(A) tail is independent of intact eIF4G and PABP. Mol Cell 2004; 15: 925–935.
Spriggs KA, Cobbold LC, Jopling CL, Cooper RE, Wilson LA, Stoneley M et al. Canonical initiation factor requirements of the Myc family of internal ribosome entry segments. Mol Cell Biol 2009; 29: 1565–1574.
Bordeleau ME, Mori A, Oberer M, Lindqvist L, Chard LS, Higa T et al. Functional characterization of IRESes by an inhibitor of the RNA helicase eIF4A. Nat Chem Biol 2006; 2: 213–220.
Komar AA, Hatzoglou M . Cellular IRES-mediated translation: The war of ITAFs in pathophysiological states. Cell Cycle 2011; 10: 229–240.
Pause A, Methot N, Svitkin Y, Merrick WC, Sonenberg N . Dominant negative mutants of mammalian translation initiation factor eIF-4A define a critical role for eIF-4F in cap-dependent and cap-independent initiation of translation. EMBO J 1994; 13: 1205–1215.
Kolupaeva VG, Lomakin IB, Pestova TV, Hellen CUT . Eukaryotic initiation factors 4G and 4A mediate conformational changes downstream of the initiation codon of the encephalomyocarditis virus internal ribosomal entry site. Mol Cell Biol 2003; 23: 687–698.
Shen J, Xia W, Khotskaya YB, Huo L, Nakanishi K, Lim SO et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature 2013; 497: 383–387.
Stoneley M, Paulin FE, Le Quesne JP, Chappell SA, Willis AE . C-Myc 5' untranslated region contains an internal ribosome entry segment. Oncogene 1998; 16: 423–428.
Tan TC, Crawford DH, Jaskowski LA, Subramaniam VN, Clouston AD, Crane DI et al. Excess iron modulates endoplasmic reticulum stress-associated pathways in a mouse model of alcohol and high-fat diet-induced liver injury. Lab Invest 2013; 93: 1295–1312.
Lang KJ, Kappel A, Goodall GJ . Hypoxia-inducible factor-1alpha mRNA contains an internal ribosome entry site that allows efficient translation during normoxia and hypoxia. Mol Biol Cell 2002; 13: 1792–1801.
Bottley A, Phillips NM, Webb TE, Willis AE, Spriggs KA . eIF4A inhibition allows translational regulation of mRNAs encoding proteins involved in Alzheimer's disease. PLoS ONE 2010; 5: e13030.
Rogers JT, Randall JD, Cahill CM, Eder PS, Huang X, Gunshin H et al. An iron-responsive element type II in the 5'-untranslated region of the Alzheimer's amyloid precursor protein transcript. J Biol Chem 2002; 277: 45518–45528.
Cho H, Lee HC, Jang SK, Kim YK . Iron increases translation initiation directed by internal ribosome entry site of hepatitis C virus. Virus Genes 2008; 37: 154–160.
Woeller CF, Fox JT, Perry C, Stover PJ . A ferritin-responsive internal ribosome entry site regulates folate metabolism. J Biol Chem 2007; 282: 29927–29935.
Chung TH, Hsiao JK, Hsu SC, Yao M, Chen YC, Wang SW et al. Iron oxide nanoparticle-induced epidermal growth factor receptor expression in human stem cells for tumor therapy. ACS Nano 2011; 5: 9807–9816.
Baldys A, Aust AE . Role of iron in inactivation of epidermal growth factor receptor after asbestos treatment of human lung and pleural target cells. Am J Respir Cell Mol Biol 2005; 32: 436–442.
Pruitt KD, Brown GR, Hiatt SM, Thibaud-Nissen F, Astashyn A, Ermolaeva O et al. RefSeq: an update on mammalian reference sequences. Nucleic Acids Res 2014; 42: D756–D763.
Smith C, Heyne S, Richter AS, Will S, Backofen R . Freiburg RNA Tools: a web server integrating INTARNA, EXPARNA and LOCARNA. Nucleic Acids Res 2010; 38: W373–W377.
Will S, Joshi T, Hofacker IL, Stadler PF, Backofen R . LocARNA-P: accurate boundary prediction and improved detection of structural RNAs. RNA 2012; 18: 900–914.
Will S, Reiche K, Hofacker IL, Stadler PF, Backofen R . Inferring noncoding RNA families and classes by means of genome-scale structure-based clustering. PLoS Comput Biol 2007; 3: e65.
Lucchini E, Pilotto S, Spada E, Melisi D, Bria E, Tortora G . Targeting the epidermal growth factor receptor in solid tumors: focus on safety. Expert Opin Drug Saf 2014; 13: 535–549.
Wilson WR, Hay MP . Targeting hypoxia in cancer therapy. Nat Rev Cancer 2011; 11: 393–410.
Ward C, Langdon SP, Mullen P, Harris AL, Harrison DJ, Supuran CT et al. New strategies for targeting the hypoxic tumour microenvironment in breast cancer. Cancer Treat Rev 2013; 39: 171–179.
Acknowledgements
We thank Drs Anna Grabowska and Tyson Sharp for use of their hypoxic incubators. We also thank Dr Hilary Collins for assistance preparing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
Oncogenesis is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Webb, T., Hughes, A., Smalley, D. et al. An internal ribosome entry site in the 5′ untranslated region of epidermal growth factor receptor allows hypoxic expression. Oncogenesis 4, e134 (2015). https://doi.org/10.1038/oncsis.2014.43
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2014.43