Abstract
Circular RNA (circRNA) is a novel member of the noncoding cancer genome with distinct properties and diverse cellular functions, which is being explored at a steadily increasing pace. The list of endogenous circRNAs involved in cancer continues to grow; however, the functional relevance of the vast majority is yet to be discovered. In general, circRNAs are exceptionally stable molecules and some have been shown to function as efficient microRNA sponges with gene-regulatory potential. Many circRNAs are highly conserved and have tissue-specific expression patterns, which often do not correlate well with host gene expression. Here we review the current knowledge on circRNAs in relation to their implications in tumorigenesis as well as their potential as diagnostic and prognostic biomarkers and as possible therapeutic targets in future personalized medicine. Finally, we discuss future directions for circRNA cancer research and current caveats, which must be addressed to facilitate the translation of basic circRNA research into clinical use.
Similar content being viewed by others
Introduction
During the past decades, new cellular roles for RNA molecules have been continuously discovered leading to the current understanding that gene expression dynamics in cancer pathology are extremely complex. Recently, it has become apparent that thousands of genes produce highly conserved and stable covalently closed RNA circles with gene-regulatory potential. Although circular RNAs (circRNAs) were first discovered nearly 40 years ago,1 it was early on suggested that they may represent errors of the normal splicing process2 and were generally considered peculiarities of uncertain biological importance. In recent years, however, we have witnessed an explosion in published studies on all aspects of circRNA biology leading to the common understanding that these molecules are important players in normal cellular differentiation and tissue homeostasis as well as in disease development. Importantly, the expression of circRNAs does not often correlate well with the expression of host gene linear expression. This argues that circRNAs are not merely steady-state byproducts of mRNA splicing but rather the product of a new type of regulated alternative splicing. Another piece of evidence that circRNAs are biologically important has come from sequence conservation analyses.3, 4 Memczak et al.4 showed that 23 human circRNAs with orthologous exons being circularized in mice were more conserved in the third position of codons compared with exons not observed to be part of circRNAs, as expected if the circRNAs have important non-coding functions.
There are several reasons why these molecules lurked in the shadows for so many years. First, it is necessary to actively search for circRNAs to be able to detect them. Conventional reverse transcriptase-quantitative PCR (RT-qPCR) assays will not distinguish circular from linear RNA when using the linear genome as template for primer design. Likewise, sequencing reads spanning back-splicing junctions in deep sequencing data are discarded in standard bioinformatic pipelines as they do not map to the linear reference genome. Another feature of circRNAs that has facilitated their escape from the spotlight is the lack of poly(A) tails as most protocols for RNA sequencing library preparation include a poly(A) purification step to remove rRNA.
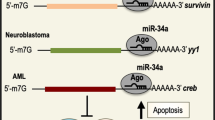
We were among the first to reveal a gene-regulatory function of a particular circRNA. We named it circRNA sponge for miR-7 (ciRS-7), as it contains >70 conserved binding sites for miR-7.4, 5 It is also known as CDR1 antisense (CDR1as), as it is located on the opposite strand of the gene CDR1. Owing to the relatively high expression and stability of ciRS-7 in many human tissues, it can increase the expression levels of miR-7 target genes by suppressing miR-7 activity. Because miR-7 directly targets several oncogenes and is involved in many different human cancers,6 ciRS-7 has now become one of the most studied circRNAs. Other circRNAs have subsequently been shown to have microRNA (miRNA) sponging properties, though most circRNAs are believed to possess other functions.7 Here we will discuss the various functions that have been proposed for circRNAs with an emphasis on their relevance in cancer.
Not much is known about how circRNA biogenesis is regulated. For some circRNAs, the characteristic back-splicing reaction that covalently joins the 3′-end of an exon to an upstream 5′-end, leading to the formation of a circRNA, is facilitated by reverse complementary Alu repeats flanking the circularized exon.8, 9, 10, 11, 12 Back splicing can also be regulated by splicing factors through their binding to cis-acting splicing-regulatory elements, although with regulatory rules distinct from those of canonical splicing.13, 14 Another mechanism that has been proposed for circRNA biogenesis involves an exon-containing lariat precursor created by an exon-skipping event. This lariat can be internally spliced leading to removal of the intronic sequence and production of a circRNA.15 Only a few transacting RNA-binding factors involved in circRNA biogenesis have been discovered. Most studied are ADAR, quaking (QKI), FUS, HNRNPL and DHX9, which regulate some but not all circRNAs.10, 16, 17, 18, 19 Interestingly, QKI has recently been shown to be a novel tumor suppressor in lung cancer and to be associated with prognosis,20, 21 and DHX9 has been suggested as a potential therapeutic target in several malignancies.22 A summary of circRNA biogenesis mechanisms is provided in Figure 1.
Mechanisms of circRNA biogenesis. (a) Efficient back splicing and circRNA generation requires that the 3′-end of an exon be located in close proximity to an upstream 5′-end of the same or another exon. This may be facilitated by the formation of a looping structure held in place by complementary base pairing of the RNA, for example, due to the presence of flanking reverse complementary Alu repeats. Alternatively, the looping structure may be held in place by binding of dimerizing RNA binding proteins with corresponding binding motifs present in the flanking regions. The small arrows indicate the back-splicing junctions of the circular splicing products. (b) Alternatively, circRNAs can be generated from an exon-containing lariat precursor created by an exon-skipping event followed by internal splicing depending on the kinetics of splicing versus debranching and exonuclease-mediated degradation.
The regulation of circRNA turnover is also not well understood. High cellular stability is a common circRNA feature owing to the lack of free ends and resistance to exonucleolytic degradation, but circRNA may contain specific endonuclease sites that could be cleaved in a regulated manner. At least one example exists where a highly complementary site to miR-671 triggers specific cleavage of ciRS-7 by Ago2.23
Though only limited knowledge has been attained on the mechanisms governing the expression of these important biomolecules, hundreds of circRNAs have been shown to be deregulated in distinct human cancers, including liver, lung, colorectal, breast, prostate, bladder, ovarian, kidney and gastric cancer as well as hematological malignancies and tumors of the central nervous system. Thus it is becoming increasingly clear that circRNAs are important in cancer pathogenesis, and they are likely to affect several of the hallmarks of cancer.24
The expression of distinct circRNAs is often disease specific or predictive of outcome. Thus circRNAs have great potential as biomarkers in cancer.25 This potential is further substantiated by circRNAs being detectable in body fluids, such as blood and saliva, which is essential if they are to be used as non-invasive biomarkers.26, 27, 28, 29
Altogether, this review summarizes the current knowledge on circRNAs in relation to their biological implications in cancer development and progression as well as their potential as diagnostic and prognostic biomarkers and as therapeutic targets in future personalized medicine. Finally, we also discuss future directions for circRNA cancer research and current caveats, which need to be addressed to facilitate the translation of basic circRNA research into clinical use.
Potential functions of circRNAs in cancer
Though some endogenous circRNAs contain AUG sites and internal ribosome entry site (IRES) elements, there is currently limited evidence for their translation in vivo.3 However, it has been shown that some circRNAs become translated in some tissues under certain conditions,30, 31, 32 but the relevance of this has yet to be shown in cancer. In general, circRNAs are expected to have other functions independent of their host genes, which may be related to the much longer half-life compared with linear RNA transcripts. Recently, a web tool named CircInteractome (circRNA interactome) has been developed for mapping RNA-binding protein- and miRNA-binding sites on human circRNAs, which may be a helpful first step toward understanding the function(s) of a circRNA in question.33 Below we discuss potential circRNA functions, which may explain their involvement in cancer (Figure 2).
Putative functions of circRNAs in cancer. (a) Many circRNAs are likely to function as miRNA sponges or decoys. Binding of miRNAs to circRNAs may release target mRNAs from miRNA-dependent degradation resulting in more efficient translation. (b) Exon–intron circRNAs have been shown to associate with RNA pol II and enhance the transcription of their parental genes via interaction with U1 snRNP. Splicing and transcription of many genes may also be indirectly regulated through a competition between canonical splicing and back splicing. However, it is largely unknown which factors may impinge on the balance between circRNA and canonical linear splicing. (c) CircRNAs with binding motifs for an enzyme and its substrate may function as scaffolds facilitating co-localization and reaction kinetics. (d) CircRNAs with RNA-binding protein-binding motifs may function as sponges or decoys for proteins and thereby regulate their activity. (e) CircRNAs with IRES elements and AUG sites may, under certain circumstances, be used as templates for translation; however, it is currently unknown if this has any relevance in cancer.
miRNA sponges or decoys
miRNAs are important players in the pathogenesis of virtually all human cancers.34, 35 Consequently, circRNAs that function as regulators of miRNA activity are likely to be involved in cancer. To date only a few circRNAs with multiple binding sites for a single miRNA have been discovered and it has been noted that most circRNAs probably have other functions than regulating miRNAs.7 The existence of circRNAs in organisms lacking RNA interference pathways also points toward other functions. However, it has been noted that miRNA-binding sites in circRNAs are virtually devoid of single-nucleotide polymorphisms, indicating that there is a strong selective pressure in conserving them,36 and multiple miRNA-binding sites may not be necessary in order to regulate miRNAs, as exemplified by transcribed pseudogenes37 and linear long noncoding RNAs38 functioning as miRNA decoys. Indeed, many circRNAs harbor single or a few miRNA-binding sites, and some harbor binding sites for multiple miRNAs or even regulate entire miRNA families. For instance, a circRNA derived from the CCDC66 gene has been shown to harbor many more binding sites for miRNAs that target oncogenes than tumor-suppressor genes,39 and circFoxo3 harbor several miRNA-binding sites for miRNAs targeting the linear host transcript, thus releasing it from miRNA-dependent degradation.40 circHIPK3 is another example of a circRNA, which is likely to function as a miRNA sponge in cancer,41 and predicted miRNA-binding sites have been verified in Argonaute HITS-CLIP data for this circRNA.42 Finally, circPVT1 is an example of a circRNA sponging several tumor-suppressor miRNAs, including let-7b.43, 44
It should also be mentioned that binding of a miRNA to a circRNA might not always result in suppression of the miRNA. This is perhaps best exemplified by our work on ciRS-7 where we have observed it to be cleaved upon miR-671 binding in an AGO2-dependent manner and then releasing bound miR-7.23 Thus one could imagine that circRNAs with miRNA-binding sites may function as a reservoir of miRNAs or facilitate miRNA transportation. Nevertheless, more and more circRNAs are being designated miRNA-inhibitory functions in cancer (Table 1).
Protein sponges or decoys
Some circRNAs with a high density of binding sites for a single or multiple RNA-binding proteins may function as protein sponges or decoys. The best experimentally supported example of a circRNA protein sponge is derived from the mbl locus, which harbors binding sites for the MBL protein itself.9 Thus MBL is prevented from binding to other targets when tethered to the circRNA. The MBL protein can bind to the introns flanking the circularized exon and facilitate its biogenesis and may therefore be part of an auto-regulatory circuit: when the protein is in excess, it may decrease the production of its own mRNA by promoting circRNA biogenesis. Another recent example is binding of HuR to a circRNA derived from the PABPN1 gene.45 HuR also binds to PABPN1 mRNA and enhances its translation. Thus extensive binding of circPABPN1 to HuR prevents HuR binding to PABPN1 mRNA and lowers its translation. At this time, it is unknown whether circPABPN1 has a role in cancer, but it could be interesting to study as HuR is also targeting several mRNAs from tumor-suppressor and cancer-related genes, such as TP53, VHL, MYC, HIF1A and BCL2.45 Furthermore, HuR has been shown to regulate miRNAs, including the tumor-suppressor miRNA miR-7.46 In cancer, one could also imagine that circRNAs may bind tumor-suppressor proteins preventing them from carrying out their normal cellular tasks. However, strong evidence for this hypothesis is currently lacking.
Protein scaffolding
circRNAs that harbor binding sites for enzymes and their substrates are likely to function as scaffolds facilitating contact between two or more proteins. This is perhaps best exemplified by circFoxo3, which has binding sites for mouse double-minute 2 (MDM2) and p53. Mutation of these binding sites or circRNA knockdown resulted in less pull-down of p53 with an MDM2 antibody and vice versa supporting the notion that circFoxo3 can function as a protein scaffold. Further, it was shown that circFoxo3 facilitated MDM2-mediated ubiquitination of p53, which subsequently becomes degraded by the proteasome.47 The long half-lives of circRNAs probably enhance their ability to function as scaffolds and more circRNAs functioning as such are likely to be discovered in the future.
Splicing and transcription
Most circRNAs are located to the cytoplasm where they may function as miRNA or protein decoys or as transporters or scaffolds. However, some circRNAs are retained in the nucleus, where they may interfere with transcription or promote alternative splicing. Exon–intron circRNAs have been shown to associate with RNA pol II and promote the transcription of their parental genes via interaction with U1 snRNP.48 However, at this stage it is unknown whether exon–intron circRNAs become deregulated in cancer. On the other hand, there is a passive competition between circular and linear splicing for most host genes as circRNA generation is generally not compatible with formation of a functional mRNA. Thus perturbation of the balance between circular and linear splicing could promote aberrant transcription of oncogenes and tumor-suppressor genes. Finally, it could be imagined that some circRNAs become relocated to the nucleus as part of the malignant transformation as we have shown for circZEB1 in porcine brain development.8
circRNAs and the hallmarks of cancer
At this time, circRNAs have been implicated in several of the hallmarks of cancer (Figure 3). One of the more studied circRNAs is derived from the tumor-suppressor gene Foxo3 and has been found to promote cancer cell apoptosis through various mechanisms.40, 47 Du et al.47 showed that circFoxo3 may indirectly upregulate its linear host gene by interacting with p53 and MDM2. Overexpression of circFoxo3 decreased the interaction between Foxo3 and MDM2, releasing Foxo3 from MDM2-dependent degradation. This led to increased Foxo3 activity and, in turn, downstream Puma and Bax-mediated apoptosis.47 circFoxo3 has also been implicated in inhibition of angiogenesis40 and cell cycle progression,49 the latter by forming a circFoxo3–p21–CDK2 (cyclin-dependent kinase 2) ternary complex, which prevents CDK2 from promoting cell cycle progression.49 Unlike circFoxo3, circRNAs derived from TTBK2 and UBAP2 have been shown to inhibit apoptosis,50, 51 and similar to circFoxo3, circZNF292 has been shown to have tumor-suppressor properties by negatively regulating cell cycle progression.52 However, another study has suggested that circZNF292 is induced by hypoxia and exhibits proangiogenic activities.42 The molecular functions underlying these effects are unknown and it is unlikely to function as a miRNA sponge as Boeckel et al.42 found no evidence for Argonaute binding to circZNF292 from HITS-CLIP data. circMYLK is another example of a circRNA that may promote angiogenesis through the vascular endothelial growth factor A (VEGFA)/VEGFR2 signaling pathway.53
Another well-characterized circRNA is circITCH, which negatively affects the cell cycle and proliferation. These effects are mediated through inhibition of the Wnt/β-Catenin pathway.54, 55 On the other hand, ciRS-7 has a positive effect on the cell cycle by enhancing the epidermal growth factor receptor/RAF1/mitogen-activated protein kinase pathway through inhibition of miR-7 activity,56 and circSLC30A7 promotes cell cycle progression by enhancing CDK6 expression through sponging of miR-29b family members.57 circMTO1 was shown to negatively affect the cell cycle and proliferation by releasing p21 from miR-9 mediated downregulation.58 Finally, aberrant expression of several circRNAs have been associated with a late-stage diagnosis and metastases,58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68 and a few (circZKSCAN1, circCCDC66, circKCNH1 and circHIAT1) have been shown to affect metastasis both in vitro and in vivo.39, 61, 65, 69
It is likely that many more circRNAs are involved in the hallmarks of cancer as many studies have shown an effect of circRNA knockdown or overexpression on proliferation rates of cancer cells in culture without exploring the molecular mechanisms behind.
Aberrant expression of circRNAs in diverse cancer types and tissues
Early on, it was shown that circRNAs are abundantly and differentially expressed in various cancer cell lines using publicly available high-throughput RNA sequencing (RNA-Seq) data from the ENCODE consortium.70 This, and other groundbreaking studies, has facilitated an explosion in published studies of circRNAs in cancer. Below we discuss the most important studies of circRNAs in solid tumors and hematological malignancies. The circRNAs most likely to be involved in cancer and their putative functions are shown in Table 1.
Solid tumors
Studies of circRNA have been conducted in most common types of solid cancer. However, the majority were designed to investigate one or more predefined circRNAs without the possibility to discover novel circRNA species. Common for most studies is that circRNA expression is compared between cancer tissues and adjacent noncancerous tissues often followed by additional functional studies. Most of the studies were performed using microarray analysis followed by validation of selected circRNAs in larger cohorts by RT-qPCR.
In basal cell carcinoma (BCC)
A common cancer of the skin, Sand et al.71 discovered 23 upregulated and 48 downregulated circRNAs in BCC tissue samples relative to noncancerous epithelial skin samples. The two most downregulated circRNAs were hsa_circ_0022383 and hsa_circ_0022392, which are both derived from the FADS2 gene. Likewise, the two most upregulated circRNAs were derived from the same host gene, LINC00340, encoding a long noncoding RNA likely to be involved in BCC.72
In bladder cancer
Zheng et al.41 found an overall upregulation of circRNAs, which is in contrast to what has been found in most other cancers. Similarly, a study by Zhong et al.73 found more upregulated than downregulated circRNAs in bladder cancer tissues relative to adjacent noncancerous tissue samples. Most interestingly, a circRNA derived from TCF25 was observed to promote proliferation and migration of bladder cancer cells by sponging miR-103a-3p and miR-107.73 More recently, a circRNA from MYLK was shown to have oncogenic properties in bladder cancer by binding miR-29a and abolishing the endogenous suppressive effect on its target gene VEGFA, in turn, leading to epithelial–mesenchymal transition, angiogenesis and metastasis.53
In breast cancer
Nair et al.74 used RNA-seq data from 885 samples, including tumors and normal adjacent tissue, provided from The Cancer Genome Atlas (TCGA). Limitations of this study are that the RNA-seq libraries from TCGA were prepared using a poly(A) purification step to remove rRNA and most tumor samples did not have a paired normal adjacent tissue sample. However, 256, 288 and 411 tumor-specific circRNAs were found in triple negative (TN), estrogen receptor positive (ER+) and HER2-positive breast cancer subtypes, respectively.74 Another study found the putative tumor suppressor, circFOXO3, to be significantly downregulated in breast tumor samples relative to benign samples47 and this circRNA is likely to have a role in breast cancer progression.75 Finally, a circRNA from ABCB10 was shown to be upregulated in breast cancer and knockdown of this circRNA in vitro suppressed proliferation of breast cancer cells and enhanced apoptosis.76
In colorectal carcinoma (CRC)
Bachmayr-Heyda et al.77 showed a global reduction of circRNAs in cancer cell lines and CRC tissues compared with adjacent noncancerous tissues. This reduction was negatively correlated to proliferation and the authors explain this by circRNAs accumulating in non-proliferating cells due to their high stability, while being distributed between daughter cells of proliferating cells.77 In contrast to most circRNAs, ciRS-7 was found to be upregulated in CRC, while a circRNA derived from the m6A methyltransferase gene METTL3 was the most downregulated.77 Upregulation of ciRS-7 in CRC was later also shown by Weng et al.56 in a large study comprising a training cohort of 153 primary CRC tissues and a validation cohort of 165. High expression of ciRS-7 was associated with poor patient survival in both cohorts. In another study, using RNA-seq data, circRNAs were also found to be significantly downregulated in CRC.41 In contrast, Hsiao et al.39 found several upregulated circRNAs in CRC, among them a circRNA derived from the CCDC66 gene, which may function as an oncogenic decoy by binding miRNAs that mainly target oncogenes. Zhang et al.66 also found more downregulated than upregulated circRNAs in CRC tissues relative to adjacent noncancerous tissues, and the most downregulated was derived from the PTK2 tumor-suppressor gene. Finally, downregulation of a circRNA derived from ITCH78 and upregulation of circRNAs derived from STIL60 and BANP79 in 45, 30 and 35 CRC samples compared with adjacent noncancerous tissue samples, respectively, have been observed.
In cutaneous squamous cell carcinoma
Sand et al.80 discovered 143 upregulated and 179 downregulated circRNAs in cutaneous squamous cell carcinoma tissue samples relative to noncancerous skin samples. The two most downregulated circRNAs were the same two FADS2 gene-originating circRNAs found to be most downregulated in BCC, in another study by Sand et al.,71 while the two most upregulated circRNAs were derived from the LARP1B gene.
In esophageal squamous cell carcinoma (ESCC)
circITCH has been studied54 in 684 ESCC cases and paired adjacent noncancerous tissue samples. Similar to the studies of this circRNA in CRC and lung cancer, it was observed to be downregulated. Another study indicated that circRNAs may be involved in the development of radiation resistance in ESCC. circRNA expression was compared between an ESCC cell line and a self-established radioresistant counterpart. Several circRNAs were observed to be upregulated and downregulated; however, circITCH was not among those.81 Finally, Xia et al.63 investigated the expression of a circRNA derived from exons 15 and 16 of the PRKCI gene in 51 cases of ESCC, finding it to be upregulated compared with adjacent noncancerous tissue samples and related to poor tumor differentiation and late Tumor, Node, Metastasis stage.
In gastric cancer
Both Zheng et al.41 and Chen et al.44 found more downregulated than upregulated circRNAs using RNA-seq. An interesting circRNA emerging from the latter study is circPVT1, which was upregulated in gastric cancer. Surprisingly though, high expression of circPVT1 was an independent predictor of better overall survival and disease-free survival. Thus it is unclear at the moment whether circPVT1 acts as a tumor suppressor or an oncogene. Another interesting finding is that circRNA expression could be a promising tool for predicting early recurrence after radical surgery of stage III gastric cancer.82 However, this study was limited by a short patient follow-up time. Finally, three independent studies from the same group found three different circRNAs from the CNIH4, HIAT1 and KIAA0907 genes to be downregulated gastric cancer tissues relative to adjacent noncancerous tissues.59, 61, 62
In glioma
Song et al.83 analyzed RNA-seq data from 46 gliomas and noncancerous brain samples and discovered thousands of circRNA candidates of which 476 were differentially expressed. Interestingly, a circRNA derived from the VCAN gene, which is implicated in gliomagenesis, was found to be upregulated in both oligodendroglioma and glioblastoma.83 Barbagallo et al.84 analyzed the expression of miR-671-5p in glioblastoma cell lines and primary patient samples and found it to be overexpressed and correlated with downregulation of ciRS-7. Finally, circZNF292 has been found to be downregulated in glioma52 and a circRNA derived from the TTBK2 gene to be upregulated.50
In hepatocellular carcinoma (HCC)
Overexpression of ciRS-7 was shown to be a risk factor for hepatic microvascular invasion and was inversely correlated with miR-7 expression in a study including 108 patients.85 However, there was no significant difference in disease-free survival between patients with high and low ciRS-7 expression, respectively. Another study of a single candidate circRNA derived from the SHPRH gene found it to be significantly downregulated in 89 HCC samples relative to adjacent noncancerous tissues.86 Likewise, Yao et al.65 found a circRNA from the ZKSCAN1 gene to be significantly downregulated in 102 HCC samples compared with adjacent noncancerous tissues. Shang et al.87 also discovered differentially expressed circRNAs in HCC samples relative to adjacent noncancerous tissues. They were subsequently able to confirm a significant differential expression of a circRNA derived from the EIF4G3 gene in a validation cohort of 30 patients. Fu et al.67, 88 identified 527 differentially expressed circRNAs of which most were downregulated in HCC. Two of the most downregulated were associated with HCC clinicopathological characteristics and derived from SMYD4 and FAM53B, respectively. Using RNA-seq, Zheng et al.41 discovered that a circRNA derived from exon 2 of HIPK3 was differentially expressed in HCC. Recently, Han et al.58 identified a circRNA from the MTO1 gene to be downregulated in HCC and associated with poor prognosis. Finally, Guo et al.89 found circITCH to be downregulated with a concomitant inferior overall survival in a large cohort of HCC patients.
In lung cancer
Wan et al.55 studied 78 patients and focused on a circRNA derived from the ITCH gene. This circRNA was downregulated in lung cancer tissues relative to adjacent noncancerous tissues and correlated with host gene expression. In another study by Zhu et al.,68 a circRNA from ACP6 was found to be upregulated in adenocarcinoma and associated with Tumor, Node, Metastasis stage and lymphatic metastasis.
In oral squamous cell carcinomas
Chen et al.57 found many circRNAs to be differentially expressed between oral squamous cell carcinoma tissue and adjacent noncancerous tissues. Among these, an upregulated circRNA derived from the SLC30A7 gene was shown to function as a sponge for several miR-29 family members and to act as a competing endogenous RNA regulating CDK6 expression.57
In osteosarcoma
Jin et al.69 studied a single circRNA candidate from the KCNH1 oncogene. It was found to be upregulated in osteosarcoma patient samples and cell lines and to promote proliferation, invasion and metastasis in vitro and in vivo.69 In another study, Zhang et al.51 found a circRNA from UBAP2 to be the most upregulated circRNA in osteosarcoma patient samples relative to adjacent noncancerous tissues. High expression of circUBAP2 correlated with inferior overall survival of the patients and it was shown to promote osteosarcoma growth and inhibit apoptosis both in vitro and in vivo.51
In ovarian cancer
Ahmed et al.90 analyzed RNA-seq data from primary and metastatic sites from three patients with stage IIIC ovarian cancer but did not include noncancerous tissues in the study. In total, 67 580 circRNA candidates were identified in the patient samples, and it was reported that more circRNAs were differentially expressed between primary ovarian tumor and metastases than mRNAs.90 Bachmayr-Heyda et al.77 observed a global reduction in circRNA expression in immortalized normal ovarian surface epithelium cell lines compared with ovarian cancer cell lines. This may be explained by the immortalized normal ovarian surface epithelium cell lines proliferating faster than the ovarian cancer cell lines, thus allowing less time for accumulation of long-lived circRNA species.77 Finally, Bachmayr-Heyda et al.91 found reduced levels of circRNAs in miliary compared with non-miliary tumors, but this could not be explained by differences in proliferation rates.
Finally, circRNA expression has also been studied in kidney clear cell carcinoma and prostate adenocarcinoma by RNA-seq.41 In prostate adenocarcinoma, circRNAs were found to be expressed at lower levels compared with adjacent noncancerous tissues, whereas the opposite was observed in kidney clear cell carcinoma.
Hematological malignancies
Only a few studies of circRNAs using primary patient samples in hematopoietic malignancies have been published. In one of the inaugural studies of circRNAs in cancer, Salzman et al.92 not only aimed at discovering new chromosomal rearrangements in diagnostic bone marrow samples from childhood cases of hyperdiploid B-linage acute lymphoblastic leukemia (ALL) using RNA-seq but also investigated the possibility that the genome-specified exon order might be rearranged during RNA splicing. To their surprise, they found hundreds of genes that produced circRNAs in the ALL patient samples. These circRNAs were, however, also detected in Hela cells and in normal hematopoietic progenitors (CD34+), naive B-cells (CD19+) and in neutrophils. On the other hand, it has been observed that circRNAs derived from genes involved in B-cell differentiation and ALL (JAK2, PAX5, IKZF1, ETV6 and EBF1) are predominantly present in ALL compared with normal leukocytes.93 Another recent study investigated circRNA expression in acute myeloid leukemia and found several deregulated circRNAs, including a circRNA derived from WDR37.94 Finally, Guarnerio et al.95 reported that circRNAs can be derived from transcription of fusion genes generated by chromosomal translocations. The authors found circRNAs derived from multiple tumor-associated translocations, including PML–RARA in promyelocytic leukemia and MLL-AF9 in acute myeloid leukemia, and named these fusion-circRNAs. This aberrant generation of circRNA is probably a result of distant, unrelated intronic sequences of the translocated genes containing complementary repetitive sequences, which favor new back-splicing events. Importantly, Guarnerio et al.95 studied the specific functions of fusion-circRNAs and found that they contribute to tumor progression by increasing cell proliferation and clonogenicity. Moreover, a circRNA derived from the MLL-AF9 fusion protected leukemia cells from the cytotoxic effects of cytarabine, a drug used for treatment of leukemia.
circRNAs as therapeutic targets
As illustrated above, deregulation of circRNAs may influence proliferative signaling, epithelial-to-mesenchymal transition, angiogenesis, apoptosis or drug resistance and, in this manner, directly have a causative role in the development of cancer. So far, there has been no preclinical reports where circRNAs alone have been used as targets or as therapeutic vectors for cancer treatment but this scenario will likely change in the future. When designing circRNA therapeutics, there are some important issues to be considered. Targeting oncogenic circRNAs should preferably be carried out in a manner that does not interfere with linear mRNA expression. A feasible therapeutic route may be to target the unique back-splice junction of oncogenic circRNAs by exogenous delivery of siRNA fully complementary to this site. Alternatively, it may be possible to interfere with back splicing by introducing antisense oligonucleotides complementary to back-splice signals in the pre-mRNA, including flanking intronic Alu repeats or binding sites for trans-acting splicing factors involved in back splicing. To induce the expression of tumor-suppressor circRNA, exogenous expression can be accomplished by gene therapy where DNA cassettes designed for circRNA expression are delivered. circRNA expression is usually accomplished by cloning the corresponding exon(s) with accompanied splice sites into a region flanked either by an artificial long inverted repeat5 or by cognate flanking regions containing Alu repeats.96 In the future, we may also see applications for preformed circRNAs independent of nuclear splicing and export. However, care should be taken not to induce an interferon response by the introduction of foreign circRNA.97 For any of the above-mentioned strategies, a major challenge will be to target a majority of the malignant cells to avoid recurrence of the cancer.
circRNA as therapeutic vectors
The unique cellular stability and capacity of circRNA to sponge miRNA and proteins may also place circRNA as a promising vehicle for delivery of therapeutics. Harnessing the circRNA with multiple binding sites for oncogenic miRNA and/or proteins may restore controlled proliferation of the cancer cells or induce apoptosis. As circRNAs are expressed from Pol II promoters, expression can be limited to certain cell types by using cell-specific promoters or even to malignant cells by disease-activated control elements. Furthermore, the sponging repertoire can include combinations of miRNA- and protein binding sites specifically designed to target specific oncogenic profiles. In this respect, it is an important observation that the genetic content of the circRNAs generally has little influence on their capacity to form circles as long as direct complementarity to miRNAs is circumvented (unpublished observations), as this is likely to cause cleavage of the circRNA via Ago2’s slicer function as observed for miR-671-mediated cleavage of ciRS-7. Finally, it has been reported that circRNAs, provided the right translation signals, can serve as templates for protein expression.30, 31, 32 Hence, including expression cassettes for tumor-suppressor proteins may turn circRNA into a Swiss army knife for cancer therapy.
Future directions and conclusions
The study of circRNAs in cancer is still in its infancy, and a common standard for reporting and naming circRNAs is lacking. We recommend that future studies provide circBase IDs for the studied circRNAs and name them according to host gene name with the prefix ‘circ’. If the circRNA is not found in circBase, the genomic position should be provided. Unfortunately, circBase is currently static and it would be beneficial to the field if this database was regularly updated with new circRNA candidates as well as a continuous removal of false-positive circRNAs.
Currently, circRNA cancer research is also hampered by most publically available RNA-seq data sets related to cancer having been prepared using a poly(A) purification step to remove rRNA, including most samples in the TCGA. Moreover, most genome-wide studies of circRNAs have been performed using microarrays on a limited number of cancer and adjacent noncancerous tissue samples, and a thorough examination of the tumor cell content in the samples by an experienced pathologist is often lacking. Likewise, the large number of circRNAs that can be detected by RNA-seq or microarray analysis makes correcting for multiple testing a major problem in determining the significance of differentially expressed circRNAs. However, most studies have validated the differential expression of selected circRNAs, using RT-qPCR with divergent primers flanking the back-splicing junction, in larger cohorts. Likewise, many of the differentially expressed circRNAs have been shown to have oncogenic- or tumor-suppressor functions in vitro and in vivo. As most studies are retrospective, it would be interesting to see how well the best circRNA biomarker candidates perform in large prospective clinical trials.
For the discovery of novel circRNA species involved in cancer, more studies using an unbiased approach, such as RNA-seq, are called for. However, it is important to realize that the results are highly dependent on RNA quality (unpublished observations) and the strategy chosen for library preparation.98 In addition, a plethora of different bioinformatic algorithms for circRNA detection is now available.99 Some prediction algorithms require gene annotation lists and knowledge of exon–intron structures, while others work de novo. This requirement may reduce the number of false positives but may also preclude the detection of some circRNAs, including ciRS-7. We have recently found a dramatic difference between the output of different algorithms, emphasizing that the circRNA species detected are candidates that need further validation.100 Therefore, it may be advisable to run more than one circRNA prediction algorithm on RNA-seq data to minimize the number of false positives in the absence of a gold standard algorithm. Though RT-qPCR is a powerful technique for validating genome-wide data, there is a caveat that template switching may occur during reverse transcription,92 which may lead to false positive or biased results. In addition, exon repeats (that is, exon concatemers in linear mRNA) have been observed,101 which can only be distinguished from circRNA in an RT-PCR-based approach by RNase R treatment or polyA enrichment. Finally, appropriate normalization of RT-qPCR data is challenging102 and the use of a single, unvalidated reference gene may lead to unreliable conclusions. Thus Northern blotting is considered the gold standard for validation of novel circRNAs as no reverse transcription and amplification steps are part of the protocol.
In situ hybridization is another attractive approach toward quantification of circRNA expression, which is likely to be employed more in future circRNA cancer research. Especially because it provides spatial information about specific circRNAs and is compatible with formalin-fixed paraffin-embedded tissue samples. Thus it is possible to distinguish circRNA expression in cancer cells from background expression of non-malignant cells within the tumor. Future research should also explore the usefulness of novel technologies for circRNA detection such as nanopore RNA sequencing (Oxford Nanopore Technologies, Oxford, UK), which can potentially provide information on the entire circRNA and not just the back-splice junction, and the nCounter platform from NanoString Technologies (Seattle, WA, USA), which can quantify RNA molecules without amplification and reverse transcription.103
A causal link between many differentially expressed circRNAs and tumorigenesis is also currently lacking, and the field is somewhat biased by the hypothesis that circRNAs as a class function as miRNA sponges. A bioinformatics approach to identify putative miRNA-binding sites in circRNAs should be used only for generation of hypotheses and a starting point for further studies. Likewise, care should be taken to assess the stoichiometric relation between the miRNA-binding sites of a proposed sponge and the target sites of the miRNA, as a significant release of target repression may often require many competing target sites.104 This issue remains difficult to access when cohorts of cells with differential expression of both miRNA and circRNA are analyzed. However, future studies of single cells will help to resolve this question. Additionally, it should be noted that canonical 6-nt binding sites may compete for miRNA binding with a potency 20% of that observed for canonical 8-nt sites.105 Unfortunately, such stoichiometric relations and the nature of the miRNA-binding sites of proposed circRNA sponges have rarely been evaluated. Likewise, data from, for example, luciferase reporter assays and AGO-HITS CLIP experiments are relatively rare in circRNA cancer research. However, in light of recent studies, suggesting that the majority of circRNAs have other functions than being miRNA sponges,7, 42 other possible functions of circRNAs should be explored in future studies.
The field of circRNA cancer research has some similarities with the cancer epigenetic field.106, 107, 108 Here methylation and downregulation of many tumor-suppressor genes were discovered before much was known about what was causing these aberrant epigenetic changes. Only later, mutations in epigenetic regulators such as DNMT3A, TET2, IDH1, IDH2, EZH2 and ASXL1 were discovered and linked to aberrant epigenetic landscapes in cancer.109 At this time, we know nearly nothing about what causes the deregulation of circRNAs in cancer. Mutations in spliceosome genes, such as SF3B1, SRSF2 and U2AF1, are prevalent in cancer where they influence alternative splicing110 and miRNA expression.111 Likewise, mutations in these genes may influence circRNA biogenesis but evidence is still lacking. Altered expression or mutation of other trans-acting factors, such as QKI,16 may also cause the differential expression of circRNAs observed in most human cancers. Recently, QKI was shown to be aberrantly expressed in lung cancer and associated with poor prognosis in two independent studies,20, 21 but none of these studies investigated circRNA expression. Thus it is tempting to speculate that the aberrant expression of circRNAs observed in cancer may, in part, be explained by genetic and/or epigenetic changes of genes involved in their biogenesis. Mutations in cis regulatory elements such as inverted repeats or protein-binding motifs could, in theory, also influence circRNA expression in cancer, but this is also still an open research question that could be pursued in future.
Another aspect of circRNA biology that has been largely unexplored is their susceptibility to become posttranscriptionally modified. Chemical modifications of RNA such as N6-methyladenosine, N1-methyladenosine, 5-methylcytosine, 5-hydroxymethylcytosine, pseudouridine (Ψ) and inosine-to-adenine editing are central in the control of coding as well as noncoding RNA activity and stability.112 Many of the writers and erasers of these modifications are mutated or aberrantly expressed in cancer. Thus it is likely that circRNA activity is hampered or enhanced in cancer if they become aberrantly modified.
If circRNAs function as important drivers of tumorigenesis or tumor suppressors, it is logical to speculate that loci producing these become amplified and deleted, respectively, in cancer. However, to our knowledge, only one study has investigated such relationships and found that the upregulation of circPVT1 in gastric cancer can be explained, in part, by genomic DNA amplification.44
Finally, placing circRNA expression in the context of other molecular alterations observed in cancer is also something future studies should aim for. A preliminary study found circRNA expression and KRAS mutations to be linked in lung and colorectal cancer,113 and another recent study found that the oncogenic transcription factor c-Myc to regulate the expression of a large number of circRNAs.114 However, the majority of the published studies have not investigated such potential correlations.
In conclusion, circRNA research is still in its infancy and their functional role in tumorigenesis is just starting to be elucidated. The glimpse uncovered so far makes circRNA a promising candidate not only as a valuable biomarker for cancer but also as potential targets or drugs in cancer therapy.
References
Hsu MT, Coca-Prados M . Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 1979; 280: 339–340.
Cocquerelle C, Mascrez B, Hetuin D, Bailleul B . Mis-splicing yields circular RNA molecules. FASEB J 1993; 7: 155–160.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013; 19: 141–157.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013; 495: 333–338.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK et al. Natural RNA circles function as efficient microRNA sponges. Nature 2013; 495: 384–388.
Hansen TB, Kjems J, Damgaard CK . Circular RNA and miR-7 in cancer. Cancer Res 2013; 73: 5609–5612.
Guo JU, Agarwal V, Guo H, Bartel DP . Expanded identification and characterization of mammalian circular RNAs. Genome Biol 2014; 15: 409.
Veno MT, Hansen TB, Veno ST, Clausen BH, Grebing M, Finsen B et al. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol 2015; 16: 245.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 2014; 56: 55–66.
Ivanov A, Memczak S, Wyler E, Torti F, Porath HT, Orejuela MR et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep 2015; 10: 170–177.
Liang D, Wilusz JE . Short intronic repeat sequences facilitate circular RNA production. Genes Dev 2014; 28: 2233–2247.
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L . Complementary sequence-mediated exon circularization. Cell 2014; 159: 134–147.
Wang Y, Wang Z . Efficient backsplicing produces translatable circular mRNAs. RNA 2015; 21: 172–179.
Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung LH et al. Exon circularization requires canonical splice signals. Cell Rep 2015; 10: 103–111.
Barrett SP, Wang PL, Salzman J . Circular RNA biogenesis can proceed through an exon-containing lariat precursor. ELife 2015; 4: e07540.
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 2015; 160: 1125–1134.
Aktas T, Avsar Ilik I, Maticzka D, Bhardwaj V, Pessoa Rodrigues C, Mittler G et al. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 2017; 544: 115–119.
Errichelli L, Dini Modigliani S, Laneve P, Colantoni A, Legnini I, Capauto D et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat Commun 2017; 8: 14741.
Fei T, Chen Y, Xiao T, Li W, Cato L, Zhang P et al. Genome-wide CRISPR screen identifies HNRNPL as a prostate cancer dependency regulating RNA splicing. Proc Natl Acad Sci USA 2017; 114: E5207–E5215.
Zong FY, Fu X, Wei WJ, Luo YG, Heiner M, Cao LJ et al. The RNA-binding protein QKI suppresses cancer-associated aberrant splicing. PLoS Genet 2014; 10: e1004289.
de Miguel FJ, Pajares MJ, Martinez-Terroba E, Ajona D, Morales X, Sharma RD et al. A large-scale analysis of alternative splicing reveals a key role of QKI in lung cancer. Mol Oncol 2016; 10: 1437–1449.
Lee T, Pelletier J . The biology of DHX9 and its potential as a therapeutic target. Oncotarget 2016; 7: 42716–42739.
Hansen TB, Wiklund ED, Bramsen JB, Villadsen SB, Statham AL, Clark SJ et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J 2011; 30: 4414–4422.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Wang F, Nazarali AJ, Ji S . Circular RNAs as potential biomarkers for cancer diagnosis and therapy. Am J Cancer Res 2016; 6: 1167–1176.
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 2015; 25: 981–984.
Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem 2015; 61: 221–230.
Memczak S, Papavasileiou P, Peters O, Rajewsky N . Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS One 2015; 10: e0141214.
Kim KM, Abdelmohsen K, Mustapic M, Kapogiannis D, Gorospe M . RNA in extracellular vesicles. Wiley Interdiscip Rev RNA 2017; 8: e1413.
Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O et al. Circ-ZNF609 Is a circular RNA that can be translated and functions in myogenesis. Mol Cell 2017; 66: 22–37.e29.
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L et al. Translation of CircRNAs. Mol Cell 2017; 66: 9–21.e7.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res 2017; 27: 626–641.
Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M . CircInteractome: a web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 2016; 13: 34–42.
Rupaimoole R, Calin GA, Lopez-Berestein G, Sood AK . miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov 2016; 6: 235–246.
Kohlhapp FJ, Mitra AK, Lengyel E, Peter ME . MicroRNAs as mediators and communicators between cancer cells and the tumor microenvironment. Oncogene 2015; 34: 5857–5868.
Thomas LF, Saetrom P . Circular RNAs are depleted of polymorphisms at microRNA binding sites. Bioinformatics 2014; 30: 2243–2246.
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP . A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010; 465: 1033–1038.
Karreth FA, Pandolfi PP . ceRNA cross-talk in cancer: when ce-bling rivalries go awry. Cancer Discov 2013; 3: 1113–1121.
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS et al. Non-coding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res 2017; 77: 2339–2350.
Yang W, Du WW, Li X, Yee AJ, Yang BB . Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene 2016; 35: 3919–3931.
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun 2016; 7: 11215.
Boeckel JN, Jae N, Heumuller AW, Chen W, Boon RA, Stellos K et al. Identification and characterization of hypoxia-regulated endothelial circular RNA. Circ Res 2015; 117: 884–890.
Panda AC, Grammatikakis I, Kim KM, De S, Martindale JL, Munk R et al. Identification of senescence-associated circular RNAs (SAC-RNAs) reveals senescence suppressor CircPVT1. Nucleic Acids Res 2016; 45: 4021–4035.
Chen J, Li Y, Zheng Q, Bao C, He J, Chen B et al. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett 2017; 388: 208–219.
Abdelmohsen K, Panda AC, Munk R, Grammatikakis I, Dudekula DB, De S et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol 2017; 14: 361–369.
Lebedeva S, Jens M, Theil K, Schwanhausser B, Selbach M, Landthaler M et al. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol Cell 2011; 43: 340–352.
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ 2017; 24: 357–370.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 2015; 22: 256–264.
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB . Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res 2016; 44: 2846–2858.
Zheng J, Liu X, Xue Y, Gong W, Ma J, Xi Z et al. TTBK2 circular RNA promotes glioma malignancy by regulating miR-217/HNF1beta/Derlin-1 pathway. J Hematol Oncol 2017; 10: 52.
Zhang H, Wang G, Ding C, Liu P, Wang R, Ding W et al. Increased circular RNA UBAP2 acts as a sponge of miR-143 to promote osteosarcoma progression. Oncotarget 2017; 8: 61687–61697.
Yang P, Qiu Z, Jiang Y, Dong L, Yang W, Gu C et al. Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/beta-catenin signaling pathway. Oncotarget 2016; 7: 63449–63455.
Zhong Z, Huang M, Lv M, He Y, Duan C, Zhang L et al. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett 2017; 403: 305–317.
Li F, Zhang L, Li W, Deng J, Zheng J, An M et al. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/beta-catenin pathway. Oncotarget 2015; 6: 6001–6013.
Wan L, Zhang L, Fan K, Cheng ZX, Sun QC, Wang JJ . Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the Wnt/beta-catenin pathway. Biomed Res Int 2016; 2016: 1579490.
Weng W, Wei Q, Toden S, Yoshida K, Nagasaka T, Fujiwara T et al. Circular RNA ciRS-7 - a promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin Cancer Res 2017; 23: 3918–3928.
Chen L, Zhang S, Wu J, Cui J, Zhong L, Zeng L et al. circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Oncogene 2017; 36: 4551–4561.
Han D, Li J, Wang H, Su X, Hou J, Gu Y et al. Circular RNA MTO1 acts as the sponge of miR-9 to suppress hepatocellular carcinoma progression. Hepatology 2017; 66: 1151–1164.
Chen S, Li T, Zhao Q, Xiao B, Guo J . Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta 2017; 466: 167–171.
Guo JN, Li J, Zhu CL, Feng WT, Shao JX, Wan L et al. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation, migration, and invasion in colorectal cancer. Oncotargets Ther 2016; 9: 7451–7458.
Li P, Chen H, Chen S, Mo X, Li T, Xiao B et al. Circular RNA 0000096 affects cell growth and migration in gastric cancer. Br J Cancer 2017; 116: 626–633.
Li P, Chen S, Chen H, Mo X, Li T, Shao Y et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta 2015; 444: 132–136.
Xia W, Qiu M, Chen R, Wang S, Leng X, Wang J et al. Circular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferation. Sci Rep 2016; 6: 35576.
Xuan L, Qu L, Zhou H, Wang P, Yu H, Wu T et al. Circular RNA: a novel biomarker for progressive laryngeal cancer. Am J Transl Res 2016; 8: 932–939.
Yao Z, Luo J, Hu K, Lin J, Huang H, Wang Q et al. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration, and invasion but through different signaling pathways. Mol Oncol 2017; 11: 422–437.
Zhang P, Zuo Z, Shang W, Wu A, Bi R, Wu J et al. Identification of differentially expressed circular RNAs in human colorectal cancer. Tumour Biol 2017; 39: 1010428317694546.
Fu L, Yao T, Chen Q, Mo X, Hu Y, Guo J . Screening differential circular RNA expression profiles reveals hsa_circ_0004018 is associated with hepatocellular carcinoma. Oncotarget 2017; 8: 58405–58416.
Zhu X, Wang X, Wei S, Chen Y, Chen Y, Fan X et al. hsa_circ_0013958: a circular RNA and potential novel biomarker for lung adenocarcinoma. FEBS J 2017; 284: 2170–2182.
Jin H, Jin X, Zhang H, Wang W . Circular RNA hsa-circ-0016347 promotes proliferation, invasion and metastasis of osteosarcoma cells. Oncotarget 2017; 8: 25571–25581.
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO . Cell-type specific features of circular RNA expression. PLoS Genet 2013; 9: e1003777.
Sand M, Bechara FG, Sand D, Gambichler T, Hahn SA, Bromba M et al. Circular RNA expression in basal cell carcinoma. Epigenomics 2016; 8: 619–632.
Sand M, Bechara FG, Sand D, Gambichler T, Hahn SA, Bromba M et al. Long-noncoding RNAs in basal cell carcinoma. Tumour Biol 2016; 37: 10595–10608.
Zhong Z, Lv M, Chen J . Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep 2016; 6: 30919.
Nair AA, Niu N, Tang X, Thompson KJ, Wang L, Kocher JP et al. Circular RNAs and their associations with breast cancer subtypes. Oncotarget 2016; 7: 80967–80979.
Lu WY . Roles of the circular RNA circ-Foxo3 in breast cancer progression. Cell Cycle 2017; 16: 589–590 1-2.
Liang HF, Zhang XZ, Liu BG, Jia GT, Li WL . Circular RNA circ-ABCB10 promotes breast cancer proliferation and progression through sponging miR-1271. Am J Cancer Res 2017; 7: 1566–1576.
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T et al. Correlation of circular RNA abundance with proliferation–exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep 2015; 5: 8057.
Huang G, Zhu H, Shi Y, Wu W, Cai H, Chen X . cir-ITCH plays an inhibitory role in colorectal cancer by regulating the Wnt/beta-catenin pathway. PLoS One 2015; 10: e0131225.
Zhu M, Xu Y, Chen Y, Yan F . Circular BANP, an upregulated circular RNA that modulates cell proliferation in colorectal cancer. Biomed Pharmacother 2017; 88: 138–144.
Sand M, Bechara FG, Gambichler T, Sand D, Bromba M, Hahn SA et al. Circular RNA expression in cutaneous squamous cell carcinoma. J Dermatol Sci 2016; 83: 210–218.
Su H, Lin F, Deng X, Shen L, Fang Y, Fei Z et al. Profiling and bioinformatics analyses reveal differential circular RNA expression in radioresistant esophageal cancer cells. J Transl Med 2016; 14: 225.
Zhang Y, Li J, Yu J, Liu H, Shen Z, Ye G et al. Circular RNAs signature predicts the early recurrence of stage III gastric cancer after radical surgery. Oncotarget 2017; 8: 22936–22943.
Song X, Zhang N, Han P, Moon BS, Lai RK, Wang K et al. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res 2016; 44: e87.
Barbagallo D, Condorelli A, Ragusa M, Salito L, Sammito M, Banelli B et al. Dysregulated miR-671-5p / CDR1-AS / CDR1 / VSNL1 axis is involved in glioblastoma multiforme. Oncotarget 2016; 7: 4746–4759.
Xu L, Zhang M, Zheng X, Yi P, Lan C, Xu M . The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol 2016; 143: 17–27.
Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z et al. Hsa_circ_0001649: a circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomarkers 2016; 16: 161–169.
Shang X, Li G, Liu H, Li T, Liu J, Zhao Q et al. Comprehensive circular RNA profiling reveals that hsa_circ_0005075, a new circular RNA biomarker, is involved in hepatocellular crcinoma development. Medicine 2016; 95: e3811.
Fu L, Wu S, Yao T, Chen Q, Xie Y, Ying S et al. Decreased expression of hsa_circ_0003570 in hepatocellular carcinoma and its clinical significance. J Clin Lab Anal 2017; e-pub ahead of print 11 May 2017; doi:10.1002/jcla.22239.
Guo W, Zhang J, Zhang D, Cao S, Li G, Zhang S et al. Polymorphisms and expression pattern of circular RNA circ-ITCH contributes to the carcinogenesis of hepatocellular carcinoma. Oncotarget 2017; 8: 48169–48177.
Ahmed I, Karedath T, Andrews SS, Al-Azwani IK, Mohamoud YA, Querleu D et al. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget 2016; 7: 36366–36381.
Bachmayr-Heyda A, Auer K, Sukhbaatar N, Aust S, Deycmar S, Reiner AT et al. Small RNAs and the competing endogenous RNA network in high grade serous ovarian cancer tumor spread. Oncotarget 2016; 7: 39640–39653.
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO . Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 2012; 7: e30733.
Bonizzato A, Gaffo E, Te Kronnie G, Bortoluzzi S . CircRNAs in hematopoiesis and hematological malignancies. Blood Cancer J 2016; 6: e483.
Li W, Zhong C, Jiao J, Li P, Cui B, Ji C et al. Characterization of hsa_circ_0004277 as a new biomarker for acute myeloid leukemia via circular RNA profile and bioinformatics analysis. Int J Mol Sci 2017; 18: 597.
Guarnerio J, Bezzi M, Jeong JC, Paffenholz SV, Berry K, Naldini MM et al. Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell 2016; 165: 289–302.
Kramer MC, Liang D, Tatomer DC, Gold B, March ZM, Cherry S et al. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev 2015; 29: 2168–2182.
Chen YG, Kim MV, Chen X, Batista PJ, Aoyama S, Wilusz JE et al. Sensing self and foreign circular RNAs by intron identity. Mol Cell 2017; 67: 228–238.e5.
Panda AC, De S, Grammatikakis I, Munk R, Yang X, Piao Y et al. High-purity circular RNA isolation method (RPAD) reveals vast collection of intronic circRNAs. Nucleic Acids Res 2017.
Szabo L, Salzman J . Detecting circular RNAs: bioinformatic and experimental challenges. Nat Rev Genet 2016; 17: 679–692.
Hansen TB, Veno MT, Damgaard CK, Kjems J . Comparison of circular RNA prediction tools. Nucleic Acids Res 2016; 44: e58.
Rigatti R, Jia JH, Samani NJ, Eperon IC . Exon repetition: a major pathway for processing mRNA of some genes is allele-specific. Nucleic Acids Res 2004; 32: 441–446.
Bustin SA, Benes V, Garson J, Hellemans J, Huggett J, Kubista M et al. The need for transparency and good practices in the qPCR literature. Nat Methods 2013; 10: 1063–1067.
Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol 2008; 26: 317–325.
Denzler R, Agarwal V, Stefano J, Bartel DP, Stoffel M . Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol Cell 2014; 54: 766–776.
Denzler R, McGeary SE, Title AC, Agarwal V, Bartel DP, Stoffel M . Impact of microRNA levels, target-site complementarity, and cooperativity on competing endogenous RNA-regulated gene expression. Mol Cell 2016; 64: 565–579.
Kristensen LS, Nielsen HM, Hansen LL . Epigenetics and cancer treatment. Eur J Pharmacol 2009; 625: 131–142.
Jones PA . DNA methylation and cancer. Oncogene 2002; 21: 5358–5360.
Jones PA, Issa JP, Baylin S . Targeting the cancer epigenome for therapy. Nat Rev Genet 2016; 17: 630–641.
You JS, Jones PA . Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell 2012; 22: 9–20.
Chabot B, Shkreta L . Defective control of pre-messenger RNA splicing in human disease. J Cell Biol 2016; 212: 13–27.
Aslan D, Garde C, Nygaard MK, Helbo AS, Dimopoulos K, Hansen JW et al. Tumor suppressor microRNAs are downregulated in myelodysplastic syndrome with spliceosome mutations. Oncotarget 2016; 7: 9951–9963.
Esteller M, Pandolfi PP . The epitranscriptome of noncoding RNAs in cancer. Cancer Discov 2017; 7: 359–368.
Dou Y, Cha DJ, Franklin JL, Higginbotham JN, Jeppesen DK, Weaver AM et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep 2016; 6: 37982.
Gou Q, Wu K, Zhou JK, Xie Y, Liu L, Peng Y . Profiling and bioinformatic analysis of circular RNA expression regulated by c-Myc. Oncotarget 2017; e-pub ahead of print 14 May 2017; doi.org/10.18632/oncotarget.17788.
Acknowledgements
We thank Maria Schertz Andersen and Mette Dahl for a careful and critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
TBH and JK are listed as inventors on a patent filed by Aarhus University concerning the use of artificial circRNA as microRNA sponges. The other authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kristensen, L., Hansen, T., Venø, M. et al. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37, 555–565 (2018). https://doi.org/10.1038/onc.2017.361
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.361
This article is cited by
-
N6-methyladenosine-modified circ_104797 sustains cisplatin resistance in bladder cancer through acting as RNA sponges
Cellular & Molecular Biology Letters (2024)
-
Circular RNAs in tumor immunity and immunotherapy
Molecular Cancer (2024)
-
RUNX1, FUS, and ELAVL1-induced circPTPN22 promote gastric cancer cell proliferation, migration, and invasion through miR-6788-5p/PAK1 axis-mediated autophagy
Cellular & Molecular Biology Letters (2024)
-
CircRNA and lncRNA-associated competing endogenous RNA networks in medulloblastoma: a scoping review
Cancer Cell International (2024)
-
Comprehensive analysis of circRNA-miRNA-mRNA network related to angiogenesis in recurrent implantation failure
BMC Medical Genomics (2024)