Abstract
The formation of a bipolar mitotic spindle is an essential process for the equal segregation of duplicated DNA into two daughter cells during mitosis. As a result of deregulated cellular signaling pathways, cancer cells often suffer a loss of genome integrity that might etiologically contribute to carcinogenesis. Merkel cell polyomavirus (MCV) small T (sT) oncoprotein induces centrosome overduplication, aneuploidy, chromosome breakage and the formation of micronuclei by targeting cellular ligases through a sT domain that also inhibits MCV large T oncoprotein turnover. These results provide important insight as to how centrosome number and chromosomal stability can be affected by the E3 ligase targeting capacity of viral oncoproteins such as MCV sT, which may contribute to Merkel cell carcinogenesis.
Similar content being viewed by others
Introduction
Merkel cell polyomavirus (MCV) is the causative agent of most Merkel cell carcinomas (MCCs), a highly aggressive human cancer.1 Shortly after the discovery of MCV, the large T (LT) and small T (sT) antigens encoded by the early region of the virus were identified as the two major oncoproteins responsible for tumorigenesis and tumor maintenance.2 Both LT and sT are expressed in virtually every MCV-infected MCC.3, 4 MCC tumors are characterized by the clonal integration of viral DNA within the host genome as well as by mutations in the viral early coding region that result in the expression of a C-terminally truncated form of LT; however, the coding region for sT is left intact.5
The sT antigen of MCV alone is able to transform rodent fibroblasts.6 While sT transgenic mice show dermal hyperproliferation,7 expression of sT combined with homozygous deletion of p53 in a transgenic mouse model leads to the development of fully transformed, highly anaplastic tumors in livers and spleens.8 The transformation activity of sT is independent of its ability to interact with PP2A,6, 9 but is localized to a short region of sT from amino acids 91–95 termed the LT-stabilization domain10 (LSD). The LSD allows sT to inhibit the cellular ubiquitin ligase complex SCFFbw7,10 which governs the degradation of several cellular oncoproteins, including c-myc and cyclin E.11 The LSD is also responsible for inhibiting the anaphase-promoting complex/cyclosome E3 ligase, which leads to increased mitogenesis and hyperphosphorylation of the major translation regulator 4E-BP1.12
Many cancer cells display evidence of genomic instability,13 such as the presence of centrosome amplification and multipolar mitotic spindles. Supernumerary centrosomes can lead to the assembly of multipolar mitotic spindles, which may in turn result in chromosome mis-segregation, aneuploidy and, over time, cancer development.14 MCC, similar to many other cancers, has been shown to exhibit a chromosomal instability tumor phenotype,15 which can be attributed in part to the properties of viral oncoproteins as commonly seen in other virus-induced human cancers. The Skp, Cullin, F-box containing complex (SCF) family of E3 ligases has been known to have an important role in the regulation of many of the major proteins involved in cell cycle.16, 17 In many cancers, mutational loss of SCF is associated with genomic instability resulting from the accumulation of its substrates, including cyclin E, cyclin D1, Polo-like kinase 4 (PLK4) and several others.18, 19 A number of viral oncoproteins implicated in human cancer have previously been shown to cause multipolar mitoses and aneuploidy by deregulation of host systems involved in ubiquitin-mediated degradation. Human papillomavirus 16 (HPV16) E7 induces centriole multiplication mediated by altered expression of PLK4, a target of SCFβ-TrCP,20, 21 hepatitis B virus HBx binding to DDB1 (ultraviolet-damaged DNA-binding protein1) E3 ubiquitin ligase complex subunit results in chromosome segregation defects,22 and human T-cell leukemia virus type-1 Tax oncoprotein directly binds and activates the Cdc20-associated anaphase-promoting complex23 resulting in mitotic abnormalities.24, 25, 26 In addition, primary endothelial cells infected with Kaposi’s sarcoma-associated herpesvirus display abnormal mitotic spindle assembly, supernumerary centrosomes and chromosome instability.27 These findings underscore the importance of the intricate network of host cell machinery in maintaining genome integrity, which, when deregulated by viral proteins, can lead to tumorigenesis.
MCV sT expression stabilizes SCFFbw7 targets such as c-myc and cyclin E10 and promotes microtubule destabilization.28 These effects implicate the induction of genomic instability as an important mechanism for MCV sT-induced oncogenesis. Here we show using several assays of genomic instability that MCV sT, through the targeting E3 ligases, fosters a genomically unstable phenotype. Expression of sT in vitro results in the formation of supernumerary centrosomes and increased aneuploidy and chromosomal breakage in an LSD-dependent manner. Inducible expression of sT in vivo leads to the development of micronuclei in reticulocytes. Loss of Fbw7 either by knockout or by knockdown leads to the formation of supernumerary centrosomes, an effect that can be recapitulated by sT expression when Fbw7 is replete.
Results
Overexpression of MCV sT induces centrosomal aberration and aneuploidy
To determine whether MCV sT causes centrosome amplification, NIH3T3 cells were stably transduced with lentiviruses for wild-type (WT) MCV sT, a PP2A-binding mutant sT (L142A), an LSD mutant sT (LSDm), HPV16 E7, SV40 sT or an empty vector. These stably transduced NIH3T3 cells were enriched for mitosis by arrest with the cyclin-dependent kinase 1 inhibitor RO-3306 (10 μm) at the mitotic boundary and subsequently released. Cells were then fixed and immunostained for γ-tubulin, a principle protein component of centrosomes. Abnormal centrosome numbers and multipolar mitotic assemblies were readily detected in metaphase spreads from cells expressing WT MCV sT or L142A mutant sT (Figure 1a), reaching frequencies >40% of the 70 mitotic cells counted in each experiment (Figure 1b). This 1.5- to 2-fold enrichment of supernumerary centrosomes observed in cells expressing MCV sT or a mutant unable to interact with PP2A9 was comparable to cells that expressed the HPV16 E7 oncoprotein, which has previously been reported to cause centrosome amplification.24 Mutation of the LSD of sT by alanine substitution at amino acids 91–9510 resulted in a loss of this phenotype, with the frequency of centrosome abnormalities matching cells transduced by an empty vector. Transiently transduced BJ-hTERT and Rat-1 fibroblasts also showed the formation of supernumerary centrosomes upon WT sT expression (Supplementary Figure S1).
Overexpression of sT causes centrosomal aberration in NIH3T3 cells. (a) Multipolar mitosis with centrosome amplification by viral oncoproteins. Stably transduced NIH3T3 cells were stained for centrosomes (γ-tubulin, green) and DNA (4,6-diamidino-2-phenylindole (DAPI), blue). SV40 sT antigen and HPV16 E7 were used as controls. Supernumerary centrosomes were readily visible in cells expressing WT or L142A mutant MCV sT as well as HPV16 E7 (× 1000 magnification). (b) Quantification of cells with abnormal numbers (⩾3) of centrosomes in metaphase out of 70 cells. Data show the mean±s.e. from three experiments. Expression of oncoproteins was analyzed using specific antibodies (8E6, pAb419 and HA for MCV sTs, SV40 sT and E7, respectively) by immunoblotting from the same samples used for immunofluorescence.
The presence of supernumerary centrosomes during mitosis can result in mistakes in spindle assembly and chromosome mis-segregation, which in many cases leads to cell death by mitotic catastrophe.29, 30, 31, 32, 33 In the rare cases when an aberrant mitotic event progresses, the daughter cells that result often display an aneuploid karyotype, which has been proposed to have a causal role in tumorigenesis and tumor evolution.34, 35 To ascertain the ability of MCV sT to induce an increase in aneuploidy, early-passage WI38 human diploid fibroblast cells were selected to more easily monitor changes in DNA content. These cells were first transiently transduced with lentiviruses for WT MCV sT, L142A sT, LSDm sT or an empty vector. Changes in ploidy were monitored over the course of 10 days, using flow cytometric analysis to detect changes in DNA content. A fourfold increase in the aneuploid population, evidenced by >4N staining, was present in WT sT and L142A sT-expressing cells as early as 2 days post transduction (Figure 2a and b). This increase in genome DNA content by sT expression was maintained through to 5 days post transduction, but declined at later time points with an elevated proportion of dead cells (Supplementary Figure S2a), likely triggered by acute induction of aneuploidy and genotoxic stress29 and not the by loss of sT protein levels (Supplementary Figure S2b). The degree of aneuploidy that is observed with LSDm sT is increased twofold or less compared to the empty control (Figure 2c). In addition to increased aneuploidy, an increase in the frequency of supernumerary centrosomes was present in interphase WI38 cells (Figure 2d; Supplementary Figure S3a), and the formation of multinucleated cells (Supplementary Figure S3b) was observed at 2 days post transduction by both WT and L142A sTs but not by LSDm sT expression.
sT increases aneuploidy and number of centrosomes in human diploid WI38 cells. (a) Ploidy analysis. Cells transiently transduced with WT sT, L142A sT, LSDm sT or empty expression vectors were stained for DNA content by propidium iodide and analyzed by flow cytometry. Representative cell cycle profiles are shown for cells isolated at 2 days post transduction, with 2N, 4N and >4N populations annotated. The population of cells with >4N DNA content is magnified as respective inserts, and is most elevated for WT sT- and L142A sT-expressing cells. (b) Expression of sT. Immunoblotting was simultaneously performed on a portion of the cells used for flow cytometry to validate sT expression using mouse monoclonal, anti-sT antibody (CM8E6). Results are representative of expression at 2 days post transduction from four independent experiments. (c) Fold changes in the percentage of cells with >4N content were calculated for 2, 5, 7 and 10 days after transduction in each condition. A single asterisk denotes a significant increase over empty vector control cells (P<0.05), and a double asterisk denotes a significant increase over LSDm-expressing cells (P<0.05). Results represent the mean±s.e. from three independent experiments. (d) Transiently transduced cells were fixed at 2 days post transduction and stained for γ-tubulin, a centrosome marker. The number of centrosomes per cell was analyzed by microscopy and counted in 30 interphase cells to determine the frequency of supernumerary centrosomes. Results represent the mean±s.e. from five independent experiments.
To better characterize the type of genetic damage occurring in cells from the aneuploidy analysis, karyotype (Figure 3) and metaphase chromosome spread analysis (Supplementary Figure S4) were also performed on WI38 cells. Cells were first transduced with lentiviral vectors for WT MCV sT, LSDm sT or an empty vector. At 2 days post transduction, cells were arrested in metaphase with colcemid and collected for cytogenetic analysis. Representative karyotypes were collected for each condition, and cells expressing WT MCV sT showed an elevated level of chromosomal breakage, aneuploidy and even loss of an X chromosome (Figure 3; Supplementary Figure S4). Out of the 50 metaphase spreads that were analyzed for each condition, WT sT-expressing cells showed an increase in total number of chromosome breakages and gaps, a greater number of breakages within a single cell and an overall higher average number of breakages per cell (Table 1) that did not reach statistical significance compared to empty control. The number of breakage and gaps in LSDm sT-expressing cells was not elevated above empty control levels. In looking at the specific loci of gaps and breaks, WT sT-expressing cells had a higher frequency of breaks at telomeres and known fragile sites36 (Table 1; Supplementary Tables S1–S3), but this increase was not statistically significant. Complex rearrangements, such as translocation events, were entirely absent in the empty control cells, but were observed in three instances in cells that expressed WT sT (Table 1).
sT expression results in increased chromosomal instability in WI38 cells. Karyotyping and G-banding analysis were performed on cells at 2 days post transduction with indicated lentivirus constructs. Representative karyotypes are shown for each condition, and chromosome breaks are indicated by red arrows. The greatest amount of breakage was seen in cells that express WT MCV sT, which is also missing an X chromosome.
sT increases aneuploidy in mice
Mis-segregation of chromosomes during mitosis can also lead to chromosomal breakage or loss, and these fractured or lagging chromosomes can become packaged in micronuclei, which appear adjacent to the main nucleus following cell division.37, 38 Therefore, the presence of micronuclei is a definitive and, more important, quantifiable marker of genomic instability. To determine whether sT expression leads to the formation of micronuclei, we made use of an inducible MCV sT transgenic mouse model for an in vivo micronucleus assay. This assay quantitatively detects the presence of micronuclei retained after enucleation occurs during erythrocyte maturation by flow cytometric analysis of peripheral blood cells.39 The primary cohort of mice used in this study allowed for ubiquitous, tamoxifen-inducible expression of MCV sT (red; Figure 4a). Because of the fact that in vivo tumor development has only been observed when sT is expressed and p53 is concurrently ablated,8 a second cohort of mice (blue) with this phenotype was also included to determine whether loss of p53 affected the frequency of micronuclei formation. Mice lacking cassettes for Cre recombinase in each of those backgrounds served as negative controls (black and gray). At 48 h post induction of sT expression, a significant increase in the presence of micronuclei in reticulocytes could be detected at nearly equal levels in mice both with and without p53 (Figure 4b). The frequency of micronuclei in reticulocytes was increased 3.3-fold upon sT expression alone (red) and 2.1-fold when combined with a p53 knockout (blue). In the same mice from which erythrocytes were collected, immunoblotting was performed on lysates extracted from a panel of tissues following necropsy to confirm that sT expression was induced ubiquitously in all experimental mice (Cre+/−) and not in tissues from control mice (Cre−/−; Figure 4c). Together, these data indicate that MCV sT is capable of inducing chromosomal damage in an in vivo setting.
sT expression in transgenic mouse model increases frequency of micronuclei. (a) Schematic of the transgenic mice used in this study. All mice were subjected to intraperitoneal injection of tamoxifen (0.2 mg/g) with blood and tissue samples collected 48 h later at necropsy. (b) In vivo micronucleus assay. Frequency of micronuclei in reticulocytes (RETs) was measured by staining DNA (MicroFlow kits, Litron Laboratories, Rochester, NY, USA) using flow cytometry. An increase in the frequency of micronucleated RETs was detected in mice after sT induction (~1%) compared to control mice (~0.3%). This corresponds to a threefold increase for sT expression alone (***P<0.0001) and twofold when p53 is concurrently knocked out (**P=0.0005). Numbers of mice within each group are indicated. (c) Near-ubiquitous expression of sT was detected in panel of mouse organs from Cre+/− mice only but not from Cre−/− controls. Representative results are shown for each mouse background. Statistical significance between two groups was determined using two-tailed Student’s t-tests in either GraphPad Prism or Excel.
sT targets E3 ligases to induce centrosome abnormality
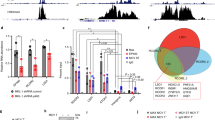
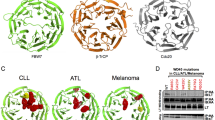
MCV sT through its LSD has been shown to inhibit multiple ubiquitin ligases, including the SCFFbw7 complex10 and the cdc20 subunit of the anaphase-promoting complex/cyclosome,12 which govern a number of substrates involved in centrosome duplication and cell division.40, 41, 42, 43 To examine the effect of E3 ligases targeted by sT in centrosome amplification, HCT116 cells deleted for the FBW7 gene18 were used. Supernumerary centrosomes form spontaneously in FBW7-null cells (−/−), but sT expression drives further centrosome amplification in this FBW7-null background, where LSDm sT cannot (Figure 5a; Supplementary Figure S5). Expression of WT but not LSDm sT in the parental FBW7 WT (+/+) cells also results in the formation of supernumerary centrosomes. Together these results underscore the importance of Fbw7 in the regulation of centrosome duplication, although it is not the sole target of MCV sT in causing a genomically unstable phenotype. Additional immunostaining against Cep170 (red) in WT sT-expressing HCT116 cells allowed us to distinguish mature ‘mother’ centrioles from newly formed daughter centrioles, which were correspondingly labeled with green fluorescent protein-tagged centrin (green). In all cases, we observed a single ‘mother’ centriole with multiple daughter centrioles, indicative of centriole overduplication as opposed to accumulation44 (Figure 5b). HPV16 E7-induced centriole overduplication requires aberrant PLK4,20 a substrate of β-TrCP E3 ubiquitin ligase and so we examined potential interaction between MCV sT and β-TrCP by immunoprecipitation. When MCV sT is coexpressed with hemagglutinin (HA)-tagged β-TrCP in 293 cells, sT was readily detectable after co-immunoprecipitation with HA antibody, and mutation of the LSD specifically impaired its binding to β-TrCP (Figure 5c). Further, we tested the effect of sT-targeting E3 ligase knockdown in centrosome amplification. Either knockdown of Fbw7 or cdc20 resulted in a twofold increase in abnormal centrosome numbers in 293 cells, with the frequency of centrosomal aberrations in these cells comparable to that achieved by expression of WT MCV sT alone (Figure 5d). Unexpectedly, knockdown of β-TrCP by short hairpin RNA (shRNA) lentiviral induction caused cell death, sustained the low efficiency of target gene silencing and resulted in a modest increase in supernumerary centrosomes (Figure 5e). The effect of sT expression on the stability of cellular Fbw7 and β-TrCP targets associated with genomic instability, including c-myc, PLK1, PLK4, YAP and Deptor, was examined in 293 cells. Consistent with our previous data10 sT inhibited the turnover of proteins targeted by E3 ligase in the presence cycloheximide in an LSD-dependent manner (Figure 5f; Supplementary Figure S6).
sT targets E3 ligases to induce centrosome abnormality. (a) Centrosome abnormality induced by sT expression or loss of Fbw7. HCT116 cells were stained for centrosomes (γ-tubulin, red) and DNA (4,6-diamidino-2-phenylindole (DAPI), blue). Supernumerary centrosomes form spontaneously in an Fbw7-null clone (FBW7−/−). Expression of WT sT, but not LSDm sT, results increased supernumerary centrosomes over empty vector control levels in both the parental (FBW7+/+) and knockout (FBW7−/−) cell lines. Images were captured at × 1000 magnification with oil. (b) sT induces centrosome overduplication. HCT116 were transfected with green fluorescent protein (GFP)-centrin to visualize centrioles (green) along with MCV sT. CEP170 protein (red) was stained to specifically visualize ‘mother’ centrioles. The single mother with multiple daughter centrioles is induced by sT overexpression. Images were captured at × 1000 magnification. (c) MCV sT interacts with β-TrCP in an LSD-dependent manner. HEK293 cells were transfected with increasing amounts of WT sT (1, 3 and 5 μg) or LSDm (2, 6 and 10 μg) along with HA-tagged β-TrCP (5 μg with sT WT and 10 μg with LSDm) to obtain comparable levels of protein expression. (d) E3 ligase knockdown is sufficient for centrosome amplification. Depletion of Fbw7, or cdc20 by shRNA induces a significant increase in centrosome amplification, as does sT overexpression alone (P<0.05). Data show the mean±s.e. from three experiments, where the number of centrosomes in 30 metaphase cells was counted. (e) Efficiency of knockdown was determined by quantitative PCR instead of immunoblotting due to the low expression of cellular E3 ligases and the limited availability of Fbw7 antibody.10 (f) sT stabilizes Fbw7 and β-TrCP substrate targets relevant for genome stability. The half-lives of the target proteins of these E3 ligases are extended by expression of WT sT but not empty vector or LSDm sT. Specific targets of Fbw7, β-TrCP or both are indicated by side brackets (left). 293 cells were treated with cycloheximide (CHX; 100 μg/ml) and collected at the indicated time points. A representative α-tubulin loading control is shown. Representative results are shown from three independent experiments.
Discussion
As a recognized hallmark of cancer, the genomic instability phenotype induced by MCV sT may be a contributing factor to its transformation activity in the context of MCC development. Genomic instability is frequently characterized by centrosomal aberrations, namely the presence of supernumerary centrosomes. While a diverse set of proteins contribute to the tight regulation of centrosome duplication, it is known that members of the SCF complex are integral to this process.19 Specifically, inactivation of SCF/Fbw7 has previously been shown to be associated with increased centrosome amplification due to increased levels of c-myc, cyclin E,18 PLK243 and possibly PLK4.16 Our results demonstrate that sT expression results in the presence of supernumerary centrosomes in multiple cell lines in an LSD-dependent manner. It has been shown that the LSD is required for the binding of sT to Fbw7,10 and our results further show that turnover of Fbw7 target proteins, including c-myc, PLK1 and YAP is attenuated in the presence of sT. The accumulation of these substrates as a result of Fbw7 inhibition could in part be responsible for centrosome overduplication in sT-expressing cells.
The overlap between multiple E3 ligase pathways that regulate mitotic controls makes it particularly difficult to implicate the targeting of any single E3 ligase by MCV sT as a cause for genome instability. For example, because both Fbw7 and β-TrCP are F-box proteins that function as part of the SCF complex,45 there is overlap in the proteins they target for degradation. Similarly, although β-TrCP has been identified as the E3 ligase responsible for PLK4 degradation, which allows for tight control of centriole duplication,46 it also has been proposed that Fbw7 may participate in PLK4 regulation.16 A combination of promiscuous targeting of such E3 ligases by sT along with overlap within those E3 ligase networks may contribute to the ability of sT to produce an unstable genomic phenotype. Although sT targeting of any single E3 ligase cannot be shown to be exclusively responsible for its disturbance of genome stability, what is clear from our results, however, is that the observed phenotype requires the presence of the LSD of sT in multiple assays.
Not only is the presence of supernumerary centrosomes a marker for genomic instability in its own right, it has also been proposed as a precursor to the development of aneuploidy.47 Our results indicate that sT is capable of acutely inducing an aneuploid cell population. A subset of these populations is lost over time, which is consistent with cell death following acute genotoxic stress, in which cells with the most severe genetic abnormalities die.29 We also noted that WT or L142A sT expression both led to an increased proportion of cells in the G2/M population, which is in line with the ability of sT to promote mitogenesis as a result of targeting the anaphase-promoting complex/cyclosome.12 Our karyotyping analysis showed that WT sT-expressing cells contained more breakages at fragile sites that are uniquely induced by oncogene expression (such as cyclin E or Ras) as well as general replicative stress36 (Supplementary Tables S1–S3). In addition, WT sT-expressing cells exhibited a higher number of breaks in telomeric regions, which is of interest because telomere dysfunction has been associated with tumorigenesis.48 Although the differences noted in our karyotype analysis are not statistically significant, they show a trend toward the types of genetic damage that likely result from sT expression over time and are consistent with the small population of cells with aberrant DNA content detected by flow cytometry (Figure 2a). These single-oncoprotein overexpression experiments may not completely recapitulate the precise biology of the virus in the context of MCC, in which the LT antigen is also expressed, but these data do provide evidence that sT expression alone is capable of inducing a genomically unstable phenotype.
Materials and methods
Cell culture and transfection
HEK293 and WI38 (American Type Culture Collection CCL-75, Manassas, VA, USA) cell lines were maintained in Dulbecco’s modified Eagle’s medium (Cellgro, #10-013, Manassas, VA, USA), supplemented with 10% fetal bovine serum (VWR Seradigm, Radnor, PA, USA). hCDC4+/+ and hCDC4−/− HCT116 cells (kindly provided by Dr Bert Vogelstein and obtained from Johns Hopkins University Genetic Resources Core Facility) were grown in McCoy’s 5A medium with 10% fetal bovine serum. NIH3T3 cells were maintained in Dulbecco’s modified Eagle’s medium with 5% calf serum (Gibco, Gaithersburg, MD, USA). Transfections were performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. The short tandem repeat profiles of the HEK293 and WI38 cells were authenticated by the University of Arizona Genetics Core and NIH3T3 cells by IDEXX (Westbrook, ME, USA). All cells used in experiments were confirmed to be negative for mycoplasma at the time of each experiment.
Plasmids and lentivirus production
Codon-optimized cDNAs for MCV sT WT (GenBank accession number AEM01096.1, Addgene, Cambridge, MA, USA, #40201), sTL142A,9 sTLSDm10 and for SV40 sT (GenBank accession number KM359729),9 and HPV E7 (NCBI reference sequence NP_041326.1)49 plasmids for lentiviral transduction and transfection were described previously as referenced. shRNA-targeting sequences of CDC20 (TRCN0000003790, UniProtKB ID: Q12834) and BTRC (TRCN0000006541, UniProtKB ID: Q9Y297) cloned in pLKO.1 vector6 were used for the knockdown study. shRNAs for human Fbw7 (UniProtKB ID: Q969H0) were described previously,10 and a control shRNA construct was obtained from Addgene (#1864). Lentiviral and retroviral particles were generated in 293FT (Invitrogen) cells maintained in Dulbecco’s modified Eagle’s medium with 10% fetal bovine serum by co-transfection with the packaging vector plasmids psPAX2 and pMD2 or pCL-Ampho (Addgene) using Lipofectamine 2000 (Invitrogen). All constructs used in this study are listed in Supplementary Table S4.
Immunoblotting and antibodies
Cells were lysed in buffer (50 mm Tris-HCl (pH 8.0), 150 mm NaCl, 0.6% SDS and 5 mm NaF) containing protease inhibitors (Roche, Basel, Switzerland). Lysates were resolved by SDS-polyacrylamide gel electrophoresis and transferred to nitrocellulose membranes (GE Healthcare, Chicago, IL, USA), then incubated in primary antibodies for at least 2 h at room temperature. For quantitative infrared western blot detection, IRDye 800CW goat anti-mouse (926-32210, LI-COR) or IRDye 800CW goat anti-rabbit antibody (929-32211, LI-COR) were used as secondary antibodies. Signal intensities were analyzed at 800 nm, using the Odyssey IR Imaging System (LI-COR, Lincoln, NE, USA). The following primary antibodies were used: anti-MCV sT CM8E6;50 anti-α tubulin (B-5-1-2, T5168, Sigma-Aldrich, St Louis, MO, USA); pAb419 (sc-58665, Santa Cruz, Dallas, TX, USA); Myc (9E10, sc-40, Santa Cruz); FLAG (M2, F3165, Sigma, St Louis, MO, USA); anti-HA (16B12, MMS-101R, Biolegend, San Diego, CA, USA); and Hsc70 (sc-7298, Santa Cruz).
Flow cytometry
Cells were trypsinized and stained with a fixable viability dye (eBioscience, Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. Fixation was performed in cold 70% (vol/vol) ethanol. Fixed cells were washed with phosphate-buffered saline (PBS) containing 1% bovine serum albumin and permeabilized by addition of 0.25% Triton X-100 for 30 min on ice. To label DNA, cells were resuspended in propidium iodide/RNaseA staining solution (0.05 mg/ml propidium iodide, 0.1 mg/ml RNaseA in 1 × PBS) and incubated for 30 min at room temperature and analyzed using a Becton Dickinson (Franklin Lakes, NJ, USA) Accuri C6 flow cytometer.
Immunofluorescence microscopy
Cells were seeded on glass slides and transfected or transduced with designated plasmids. Cells were fixed with 4% buffered formalin for 10 min and permeabilized with PBS with 0.1% Triton X-100 for 10 min. After blocking with 10% goat serum in PBS-T (PBS with 0.5% Tween 20), cells were incubated overnight at 4 °C with primary antibody γ-tubulin (ab84355, Abcam, Cambridge, MA, USA) or CEP170 (72-413-1, Thermo Fisher Scientific, Waltham, MA, USA) for staining, followed by secondary antibody incubation (Alexa Fluor 488- or 568-conjugated goat anti-rabbit (A11034 or A11036) or anti-mouse (A11029 or A11004, 1:1000, Invitrogen) for 1 h at room temperature. Stained cells were mounted in aqueous medium containing 4,6-diamidino-2-phenylindole (Vector Laboratories, Burlingame, CA, USA). Cells were analyzed by fluorescence imaging using an Olympus AX70 epifluorescence microscope equipped with a U-CMAD3 camera (Olympus, Tokyo, Japan). Centrosome counting was performed in a blinded manner by two observers.
Karyotype analysis
Cells were seeded in flasks and transduced with specified lentiviruses. The following day cells were shipped to the Cytogenomics Laboratory at the University of Minnesota Masonic Cancer Center, and culture medium was replaced upon arrival. At 48 h post transduction, cells were subjected to overnight (15 h) treatment with colcemid to arrest cells in metaphase. Cells were then collected, fixed and dropped on slides according to standard cytogenetic protocol. Fifty metaphases per condition were analyzed for chromosomal gaps, breakages and losses by G-banding at 400 band resolution.
Mouse studies
All mice were maintained in accordance with protocols approved by the Institutional Animal Care and Use Committee at the University of Pittsburgh. In brief, a previously characterized tamoxifen-inducible, cre-loxP recombination system was used to achieve ubiquitous expression of MCV sT from the ROSA26 locus in C57BL/6 mice.8 Two main cohorts of mice (8–10 weeks of age) were used: UbcCreERT2−/+ROSAsT/+ for sT expression alone (five female and four male) and UbcCreERT2−/+p53flox/floxROSAsT/+ (six female and four male) for additional p53 knockout. UbcCreERT2−/− mice (four female and sevem male) from either genetic background served as negative controls, as they lack expression of Cre recombinase. All mice were subjected to intraperitoneal injection of tamoxifen (0.2 mg/g), followed by killing after 48 h. Mouse tissues were isolated and processed for immunoblotting as described previously.8 Mouse blood samples were isolated via cardiac punture, fixed in ultracold methanol and analyzed by Litron Laboratories (Rochester, NY, USA). All mice used for experiments were randomly assigned to non-blinded treatment groups. Sample size determined by Mead’s resource equation.
Immunoprecipitation
HEK293 were transfected with plasmids for expression of MCV sT WT or sT LSDm along with HA-tagged β-TrCP. Cells were lysed in immunoprecipitation buffer (50 mm Tris-HCl (pH 7.4), 150 mm NaCl and 1% Triton X-100) freshly supplemented with protease inhibitor cocktail (Roche), 1 mm phenylmethylsulfonyl fluoride and 1 mm benzamidine. Lysates were incubated overnight at 4 °C with HA-specific antibody, followed by incubation with a slurry of Protein A/G Plus Agarose beads (Santa Cruz) for 3 additional hours. Following a series of wash steps, bound proteins were eluted in 2 × SDS loading buffer, resolved by SDS-polyacrylamide gel electrophoresis, and immunoblotted for MCV sT as described above.
Quantitative real-time PCR analysis
shRNA-transfected cells were selected with puromycin (2 μg/ml) for 4 days after infection. Total RNA was isolated 2 days after transduction using TRIzol LS reagent (Invitrogen) and treated with DNase I (NEB, Ipswich, MA, USA). The cDNA was generated using random hexamer primers and Superscript III First-Strand Synthesis reverse transcriptase (Invitrogen). Quantitative PCR with cDNA (0.1 μg) was carried out using SsoFast EvaGreen Supermix (Bio-Rad, Hercules, CA, USA) and a CFX96 real-time PCR detection system (Bio-Rad) according to the manufacturer’s protocol with primer pairs for Fbw7 (5′-AAAGAGTTGTTAGCGGTTCTCG-3′ and 5′-CCACATGGATACCATCAAACTG-3′), β-TrCP (5′-TGCCGAAGTGAAACAAGC-3′ and 5′-CCTGTGAGAATTCGCTTG-3′), cdc20 (5′- CGCCAACCGATCCCACAG-3′ and 5′-CAGGTTCAAAGCCCAGGC-3′) and GAPDH (5′-ACTTCAACAGCGACACCC-3′ and 5′-TCTTCCTCTTGTGCTCTTGC-3′) cDNA detection.
Statistical analysis
Statistical significance between two groups was determined using one- or two-tailed Student’s t-tests in either GraphPad Prism (GraphPad Software, Inc., La Jolla, CA, USA) or Excel. Specifically for the karyotype results, statistical significance was determined using χ2 or Fisher’s exact tests where appropriate.
Accession codes
References
Feng H, Shuda M, Chang Y, Moore PS . Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 2008; 319: 1096–1100.
Houben R, Shuda M, Weinkam R, Schrama D, Feng H, Chang Y et al. Merkel cell polyomavirus-infected Merkel cell carcinoma cells require expression of viral T antigens. J Virol 2010; 84: 7064–7072.
Busam KJ, Jungbluth AA, Rekthman N, Coit D, Pulitzer M, Bini J et al. Merkel cell polyomavirus expression in merkel cell carcinomas and its absence in combined tumors and pulmonary neuroendocrine carcinomas. Am J Surg Pathol 2009; 33: 1378–1385.
Rodig SJ, Cheng J, Wardzala J, DoRosario A, Scanlon JJ, Laga AC et al. Improved detection suggests all Merkel cell carcinomas harbor Merkel polyomavirus. J Clin Invest 2012; 122: 4645–4653.
Shuda M, Feng H, Kwun HJ, Rosen ST, Gjoerup O, Moore PS et al. T antigen mutations are a human tumor-specific signature for Merkel cell polyomavirus. Proc Natl Acad Sci USA 2008; 105: 16272–16277.
Shuda M, Kwun HJ, Feng H, Chang Y, Moore PS . Human Merkel cell polyomavirus small T antigen is an oncoprotein targeting the 4E-BP1 translation regulator. J Clin Invest 2011; 121: 3623–3634.
Verhaegen ME, Mangelberger D, Harms PW, Vozheiko TD, Weick JW, Wilbert DM et al. Merkel cell polyomavirus small T antigen is oncogenic in transgenic mice. J Invest Dermatol 2015; 135: 1415–1424.
Shuda M, Guastafierro A, Geng X, Shuda Y, Ostrowski SM, Lukianov S et al. Merkel cell polyomavirus small T antigen induces cancer and embryonic Merkel cell proliferation in a transgenic mouse model. PLoS One 2015; 10: e0142329.
Kwun HJ, Shuda M, Camacho CJ, Gamper AM, Thant M, Chang Y et al. Restricted protein phosphatase 2A targeting by Merkel cell polyomavirus small T antigen. J Virol 2015; 89: 4191–4200.
Kwun HJ, Shuda M, Feng H, Camacho CJ, Moore PS, Chang Y . Merkel cell polyomavirus small T antigen controls viral replication and oncoprotein expression by targeting the cellular ubiquitin ligase SCFFbw7. Cell Host Microbe 2013; 14: 125–135.
Welcker M, Clurman BE . FBW7 ubiquitin ligase: a tumour suppressor at the crossroads of cell division, growth and differentiation. Nat Rev Cancer 2008; 8: 83–93.
Shuda M, Velasquez C, Cheng E, Cordek DG, Kwun HJ, Chang Y et al. CDK1 substitutes for mTOR kinase to activate mitotic cap-dependent protein translation. Proc Natl Acad Sci USA 2015; 112: 5875–5882.
Negrini S, Gorgoulis VG, Halazonetis TD . Genomic instability—an evolving hallmark of cancer. Nat Rev Mol Cell Biol 2010; 11: 220–228.
Nigg EA . Centrosome aberrations: cause or consequence of cancer progression? Nat Rev Cancer 2002; 2: 815–825.
Sahi H, Savola S, Sihto H, Koljonen V, Bohling T, Knuutila S . RB1 gene in Merkel cell carcinoma: hypermethylation in all tumors and concurrent heterozygous deletions in the polyomavirus-negative subgroup. APMIS 2014; 122: 1157–1166.
Peel N . Everything in moderation: proteolytic regulation of centrosome duplication. Worm 2013; 2: e22497.
Ang XL, Wade Harper J . SCF-mediated protein degradation and cell cycle control. Oncogene 2005; 24: 2860–2870.
Rajagopalan H, Jallepalli PV, Rago C, Velculescu VE, Kinzler KW, Vogelstein B et al. Inactivation of hCDC4 can cause chromosomal instability. Nature 2004; 428: 77–81.
Silverman JS, Skaar JR, Pagano M . SCF ubiquitin ligases in the maintenance of genome stability. Trends Biochem Sci 2012; 37: 66–73.
Korzeniewski N, Treat B, Duensing S . The HPV-16 E7 oncoprotein induces centriole multiplication through deregulation of Polo-like kinase 4 expression. Mol Cancer 2011; 10: 61.
Holland AJ, Lan W, Niessen S, Hoover H, Cleveland DW . Polo-like kinase 4 kinase activity limits centrosome overduplication by autoregulating its own stability. J Cell Biol 2010; 188: 191–198.
Martin-Lluesma S, Schaeffer C, Robert EI, van Breugel PC, Leupin O, Hantz O et al. Hepatitis B virus X protein affects S phase progression leading to chromosome segregation defects by binding to damaged DNA binding protein 1. Hepatology 2008; 48: 1467–1476.
Liu B, Hong S, Tang Z, Yu H, Giam CZ . HTLV-I Tax directly binds the Cdc20-associated anaphase-promoting complex and activates it ahead of schedule. Proc Natl Acad Sci USA 2005; 102: 63–68.
Duensing S, Lee LY, Duensing A, Basile J, Piboonniyom S, Gonzalez S et al. The human papillomavirus type 16 E6 and E7 oncoproteins cooperate to induce mitotic defects and genomic instability by uncoupling centrosome duplication from the cell division cycle. Proc Natl Acad Sci USA 2000; 97: 10002–10007.
Yun C, Cho H, Kim SJ, Lee JH, Park SY, Chan GK et al. Mitotic aberration coupled with centrosome amplification is induced by hepatitis B virus X oncoprotein via the Ras-mitogen-activated protein/extracellular signal-regulated kinase-mitogen-activated protein pathway. Mol Cancer Res 2004; 2: 159–169.
Peloponese Jr JM, Haller K, Miyazato A, Jeang KT . Abnormal centrosome amplification in cells through the targeting of Ran-binding protein-1 by the human T cell leukemia virus type-1 Tax oncoprotein. Proc Natl Acad Sci USA 2005; 102: 18974–18979.
Pan H, Zhou F, Gao SJ . Kaposi's sarcoma-associated herpesvirus induction of chromosome instability in primary human endothelial cells. Cancer Res 2004; 64: 4064–4068.
Knight LM, Stakaityte G, Wood JJ, Abdul-Sada H, Griffiths DA, Howell GJ et al. Merkel cell polyomavirus small T antigen mediates microtubule destabilisation to promote cell motility and migration. J Virol 2015; 89: 35–47.
Chi YH, Jeang KT . Aneuploidy and cancer. J Cell Biochem 2007; 102: 531–538.
Kops GJ, Weaver BA, Cleveland DW . On the road to cancer: aneuploidy and the mitotic checkpoint. Nat Rev Cancer 2005; 5: 773–785.
Fujiwara T, Bandi M, Nitta M, Ivanova EV, Bronson RT, Pellman D . Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 2005; 437: 1043–1047.
Vitale I, Galluzzi L, Castedo M, Kroemer G . Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat Rev Mol Cell Biol 2011; 12: 385–392.
Storchova Z, Pellman D . From polyploidy to aneuploidy, genome instability and cancer. Nat Rev Mol Cell Biol 2004; 5: 45–54.
Fang X, Zhang P . Aneuploidy and tumorigenesis. Semin Cell Dev Biol 2011; 22: 595–601.
Ganem NJ, Godinho SA, Pellman D . A mechanism linking extra centrosomes to chromosomal instability. Nature 2009; 460: 278–282.
Miron K, Golan-Lev T, Dvir R, Ben-David E, Kerem B . Oncogenes create a unique landscape of fragile sites. Nat Commun 2015; 6: 7094.
Iarmarcovai G, Bonassi S, Botta A, Baan RA, Orsiere T . Genetic polymorphisms and micronucleus formation: a review of the literature. Mutat Res 2008; 658: 215–233.
Bonassi S, Znaor A, Ceppi M, Lando C, Chang WP, Holland N et al. An increased micronucleus frequency in peripheral blood lymphocytes predicts the risk of cancer in humans. Carcinogenesis 2007; 28: 625–631.
Witt KL, Livanos E, Kissling GE, Torous DK, Caspary W, Tice RR et al. Comparison of flow cytometry- and microscopy-based methods for measuring micronucleated reticulocyte frequencies in rodents treated with nongenotoxic and genotoxic chemicals. Mutat Res 2008; 649: 101–113.
Nakayama KI, Nakayama K . Ubiquitin ligases: cell-cycle control and cancer. Nat Rev Cancer 2006; 6: 369–381.
Crusio KM, King B, Reavie LB, Aifantis I . The ubiquitous nature of cancer: the role of the SCF(Fbw7) complex in development and transformation. Oncogene 2010; 29: 4865–4873.
Li M, Fang X, Wei Z, York JP, Zhang P . Loss of spindle assembly checkpoint-mediated inhibition of Cdc20 promotes tumorigenesis in mice. J Cell Biol 2009; 185: 983–994.
Cizmecioglu O, Krause A, Bahtz R, Ehret L, Malek N, Hoffmann I . Plk2 regulates centriole duplication through phosphorylation-mediated degradation of Fbxw7 (human Cdc4). J Cell Sci 2012; 125 (Pt 4): 981–992.
Duensing A, Chin A, Wang L, Kuan SF, Duensing S . Analysis of centrosome overduplication in correlation to cell division errors in high-risk human papillomavirus (HPV)-associated anal neoplasms. Virology 2008; 372: 157–164.
Lau AW, Fukushima H, Wei W . The Fbw7 and betaTRCP E3 ubiquitin ligases and their roles in tumorigenesis. Front Biosci 2012; 17: 2197–2212.
Guderian G, Westendorf J, Uldschmid A, Nigg EA . Plk4 trans-autophosphorylation regulates centriole number by controlling betaTrCP-mediated degradation. J Cell Sci 2010; 123 (Pt 13): 2163–2169.
Duensing S, Munger K . Human papillomaviruses and centrosome duplication errors: modeling the origins of genomic instability. Oncogene 2002; 21: 6241–6248.
Desmaze C, Soria JC, Feulet-Marrière MA, Mathieu N, Sabatier L . Telomere-driven genomic instability in cancer cells. Cancer Lett 2003; 194: 173–182.
White EA, Sowa ME, Tan MJ, Jeudy S, Hayes SD, Santha S et al. Systematic identification of interactions between host cell proteins and E7 oncoproteins from diverse human papillomaviruses. Proc Natl Acad Sci USA 2012; 109: E260–E267.
Kwun HJ, Guastafierro A, Shuda M, Meinke G, Bohm A, Moore PS et al. The minimum replication origin of merkel cell polyomavirus has a unique large T-antigen loading architecture and requires small T-antigen expression for optimal replication. J Virol 2009; 83: 12118–12128.
Acknowledgements
We thank Gutian Xiao for providing the HA-βTrCP plasmid, Bert Vogelstein for providing the HCT116 cells, and Megan Lambert and Robin Frederick for assistance in mouse studies, and Meredith Monk for help with the manuscript. The cytogenetic analyses were performed in the Cytogenomics Shared Resource at the University of Minnesota with support from the comprehensive Masonic Cancer Center grant P30CA077598. This study was supported by National Institutes of Health (NIH) grants CA136363 and CA120726 to PSM and YC, who are also supported as American Cancer Society Research Professors. Additional support was provided by the NIH grant T32AI049820 to JAW as a predoctoral trainee. This project used the UPCI Animal and Cytometry Facilities that are supported in part by award P30CA047904.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Kwun, H., Wendzicki, J., Shuda, Y. et al. Merkel cell polyomavirus small T antigen induces genome instability by E3 ubiquitin ligase targeting. Oncogene 36, 6784–6792 (2017). https://doi.org/10.1038/onc.2017.277
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.277
This article is cited by
-
Surface charge of Merkel cell polyomavirus small T antigen determines cell transformation through allosteric FBW7 WD40 domain targeting
Oncogenesis (2020)
-
A pilot study of alternative TrkAIII splicing in Merkel cell carcinoma: a potential oncogenic mechanism and novel therapeutic target
Journal of Experimental & Clinical Cancer Research (2019)
-
Morphologic and immunophenotypical features distinguishing Merkel cell polyomavirus-positive and negative Merkel cell carcinoma
Modern Pathology (2019)