Abstract
The utility of tumor-derived cell lines is dependent on their ability to recapitulate underlying genomic aberrations and primary tumor biology. Here, we sequenced the exomes of 25 bladder cancer (BCa) cell lines and compared mutations, copy number alterations (CNAs), gene expression and drug response to BCa patient profiles in The Cancer Genome Atlas (TCGA). We observed a mutation pattern associated with altered CpGs and APOBEC-family cytosine deaminases similar to mutation signatures derived from somatic alterations in muscle-invasive (MI) primary tumors, highlighting a major mechanism(s) contributing to cancer-associated alterations in the BCa cell line exomes. Non-silent sequence alterations were confirmed in 76 cancer-associated genes, including mutations that likely activate oncogenes TERT and PIK3CA, and alter chromatin-associated proteins (MLL3, ARID1A, CHD6 and KDM6A) and established BCa genes (TP53, RB1, CDKN2A and TSC1). We identified alterations in signaling pathways and proteins with related functions, including the PI3K/mTOR pathway, altered in 60% of lines; BRCA DNA repair, 44%; and SYNE1–SYNE2, 60%. Homozygous deletions of chromosome 9p21 are known to target the cell cycle regulators CDKN2A and CDKN2B. This loci was commonly lost in BCa cell lines and we show the deletions extended to the polyamine enzyme methylthioadenosine (MTA) phosphorylase (MTAP) in 36% of lines, transcription factor DMRTA1 (27%) and antiviral interferon epsilon (IFNE, 19%). Overall, the BCa cell line genomic aberrations were concordant with those found in BCa patient tumors. We used gene expression and copy number data to infer pathway activities for cell lines, then used the inferred pathway activities to build a predictive model of cisplatin response. When applied to platinum-treated patients gathered from TCGA, the model predicted treatment-specific response. Together, these data and analysis represent a valuable community resource to model basic tumor biology and to study the pharmacogenomics of BCa.
Similar content being viewed by others
Introduction
Bladder cancer (BCa) is the sixth most common cancer in the United States with over 74 000 new cases and 16 000 deaths each year.1 Although there are recurrent genetic aberrations in BCa2, 3, 4, 5, 6 that relate to overall patient survival,7 tumor classification4, 8 and response to therapies,9, 10 BCa is also characterized by elevated mutational burden11 and a high degree of molecular heterogeneity.12, 13 These factors complicate the study of molecular mechanisms of disease and highlight the need to select experimental models that recapitulate the genomic variation seen in patients.
Human tumor-derived cancer cell lines have long served as model systems to study the mechanisms driving cancer and to discover potential therapeutics.14 However, and not uncommonly, drug response in cell lines does not translate to patient tumors. One of the many factors contributing to this lack of translatability is the difference in genomic aberrations between cell lines and tumors. A systematic comparison of genomic features in cell lines and tumors has only recently been possible with contributions from individual research groups5, 6 and large collaborative efforts such as The Cancer Genome Atlas (TCGA) for tumor samples,4 and the Cancer Cell Line Encyclopedia (CCLE)10 and the Genomics of Drug Sensitivity in Cancer (GDSC)15 for cancer cell lines. One such comparison looked at high-grade serous ovarian cancer and found that the two most commonly used cell line models were hyper-mutated compared with solid tumors, highlighting a potential disconnect between the genomic features of commonly used cell lines and primary tumors. Furthermore, cross-contamination of cell lines has been well documented and likely also contributes to confounding results.16 For all of these reasons, the choice of cell line models is critical if results from in vivo experiments are to have biological and clinical relevance. A number of investigators have explored similar concepts in ovarian,17 head and neck,18 colorectal,19 melanoma,20 and non-small cell lung21 cancers. Here, we provide the data and analyses to include BCa in this growing list, a necessary step to strengthen the analysis of all cancer types.
Although the CCLE and GDSC are valuable pharmacogenomics resources, they have limited mutational profiling data and are under-represented for BCa cell lines. Previous efforts characterized a panel of 40 BCa cell lines for gene expression and response to chemotherapy,22 and analyzed copy number alterations (CNAs) using single-nucleotide polymorphism (SNP) arrays.23 Here we present mutational and CNA profiling using exome sequencing on 25 BCa cell lines, along with an integrative analysis of mutations, CNA and mRNA expression to compare the cell lines with primary human bladder tumors. We use the genomic and pathway profiles of the cell lines to stratify human BCa patients according to treatment response and identify a set of pathways that are predictive of platinum chemotherapy response. To help investigators select the appropriate cell model for experimental functional studies, we make our data, analysis results and scripts publically available: https://www.synapse.org/UC25.
Results
We performed whole-exome sequencing (WES) on 25 BCa cell lines derived from 21 muscle-invasive (MI) and 4 non-MI (NMI) tumors; these lines were authenticated using DNA fingerprinting (Supplementary Table 1). Exomes were sequenced to a mean depth of ~60X coverage with 86% of exonic bases sequenced to a depth of ⩾10X (Supplementary Table 2). All WES-predicted variants passing the mutational pipeline described in the Materials and Methods section are reported in Supplementary Table 3. A previously observed high rate of somatic BCa mutations predominated by single-nucleotide variants in CpGs and associated with the apolipoprotein B mRNA editing (APOBEC) family of DNA cytosine deaminases has been described and prompted us to examine the tri-nucleotide context of the cell line variants.4, 24 This revealed a common signature in both the 21 MI tumor-derived lines (Figure 1a) and the 4 NMI tumor-derived lines (Figure 1b) that was dominated by C>T transitions in CpGs, and by C>T and C>G APOBEC-mediated changes in a T-C-W context, where the underlined cytosine is altered and W=A/T. The signature represented 16.9% and 12.3% of all alterations, respectively, in the MI tumor-derived lines. Significantly, 204 single-nucleotide variants in known or putative BCa genes were validated in the cell line exomes as described below and can be similarly analyzed (Supplementary Table 4). Exome variants associated with APOBEC enzymes and altered CpGs comprised 50/204 (24.5%) and 33/204 (16.2%) of variants, respectively, indicating a likely mechanism(s) underlying a large fraction (40.7%) of cancer-associated alterations in the exome. The proportions of the signature-associated variants in the cell lines are within the distribution reported from the analysis of somatic alterations in MI primary tumors.4, 24 Considering the cell line analysis unavoidably incorporates both germline and somatic variants, these results indicate the strength of the APOBEC- and CpG-associated signature in the exome.
Mutation signatures in MI tumor- and NMI tumor-derived bladder cell lines. (a) Mutation signatures in 21 MI tumor-derived lines. NGS variants (29 557 single-nucleotide variants (SNVs)) were grouped by the tri-nucleotide context. Arrows indicate motifs associated with altered CpGs (white) and APOBEC-family enzymes (black); arrow size indicates the relative contribution of specific motifs to variants observed in 238 MI primary tumors.24 (b) Motifs observed in 6472 SNVs in 4 NMI tumor-derived lines (MGH-U3, MGH-U4, RT4 and SW-780).
We tested a total of 309 next-generation sequencing (NGS)-predicted variants identified in known cancer genes using PCR and Sanger sequencing, and 237 (83%) NGS-predicted variants were confirmed (Supplementary Table 4). The majority of variants that were not confirmed had a low number of variant read counts. Validated variants were then computationally assessed using the Annotation, Visualization and Analysis (AVIA) pipeline, v2.0;25, 26 more than half (128/237, 54%) were predicted to be deleterious by more than one functional analysis algorithm.
Mutational landscape of BCa cell lines
Nineteen genes have NGS-predicted, Sanger sequencing-validated alterations in ⩾4 of the 25 lines, including the lysine-specific methyltransferase 2C (MLL3/KMT2C; altered in 40% of lines), FAT atypical cadherin 4 (FAT4; 24%) and retinoblastoma 1 (RB1; 20%) (Figure 2). Using Sanger sequencing, we identified telomerase reverse transcriptase (TERT) promoter variants in 84% of lines. We also detected previously unreported variants in less frequently altered, but well-known cancer-associated genes including breast cancer 1 (BRCA1) and Kirsten Rat Sarcoma homolog (KRAS). New potentially deleterious variants such as a G>T, p.E66X were detected in tumor protein p53 (TP53) in cell line UM-UC-6 and in genes encoding chromatin remodeling functions, such as a C>T, p.R4139X in MLL3 in cell line MGH-U3, an insertion and frameshift, p.D1850fs, in the chr 1p36 locus AT-rich interactive domain 1A (ARID1A) in cell line UM-UC-6, and a deletion and frameshift, p.P1139fs, in the chr X lysine-specific demethylase 6A (KDM6A) in cell line UM-UC-13. Details are in Supplementary Table 4.
Cancer-associated gene alterations in 25 BCa cell lines. Top histogram, WES-predicted variants by nucleotide change. Right histogram, percentage of cell lines with confirmed alterations by gene. Central panel, distribution of gene alterations by cell line; missense, nonsense, coding indels, UTR, promoter, and splice site alterations are displayed. Gray, no data. Mutation details are contained in Supplementary Table 4.
We evaluated the effect of reduced TERT mRNA expression in two bladder lines with predicted activating promoter mutations. We selected the TCCSUP and UM-UC-3 lines to evaluate because they have previously been reported to have high TERT mRNA expression, high TERT protein levels and high telomerase activity.3 The UM-UC-3 line contains a heterozygous c.-124C>T variant and the TCCSUP line contains a homozygous c.-124C>T variant, both suggesting increased TERT expression.3 After TERT mRNA knockdown, both lines showed a significant decrease in growth after 5 days (Supplementary Figure 1), supporting a functional impact of the promoter mutations on overall TERT expression and the BCa cell phenotype.
We mapped proteins encoded by genes with sequence variants to putative pathways to identify potentially altered signaling networks (Figure 3). Eleven genes encoding proteins involved in phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) signaling were altered in 60% of the lines, most frequently by altered PI3K regulatory subunit (PIK3CA) and the tyrosine kinase gene, ERBB2. In agreement with previous observations,2, 27 the BRCA DNA repair pathway, including BRCA1 and ATM, were altered in 44% of lines in a predominantly mutually exclusive manner. This is similar to 40/127 (32%) of tumors from patients in the TCGA,4 which also exhibited BRCA pathway gene alterations that were predominantly mutually exclusive. Cell lines exhibited variants in genes encoding related proteins and/or functions, including the spectrin repeat containing, nuclear envelope 1 and 2 (SYNE1 and SYNE2) in 60% of lines; and the lysine-specific methyltransferases MLL3, MLL2/KMT2D, MLL5/KMT2E, and MLL/KMT2A in 52% of lines. Many of the alterations we confirmed occur in genes that are frequently mutated in primary bladder tumors,2, 4 thus enabling future experimental study of these genes and pathways using these BCa cell lines.
Cell lines with altered signaling pathways and related protein functions. (a) PI3K/mTOR alterations in 15/25 (60%) cell lines. (b) BRCA pathway alterations in 11/25 (44%) cell lines. (c) Sister chromatid cohesion and segregation alterations in 6/25 (24%) cell lines. Variants are shown in an Oncoprint format (left) to identify genes that are co- or exclusively mutated; and as a pathway diagram (right) to show the frequency of altered protein components in each pathway. (d) SYNE1 and SYNE2 alterations in 15/25 (60%) cell lines. Colored boxes indicate protein domains: green, calponin homology domain; orange, spectrin repeat; blue, nuclear envelope localization domain. (e) MLL-family alterations in 12/25 (48% ) cell lines. Colored boxes indicate protein domains: green, PHD-like zinc-binding domain; orange, PHD-finger; blue, F/Y-rich N-terminus; yellow, F/Y-rich C-terminus; purple, SET domain. Variant annotation: green, missense; black, truncating (nonsense, deletion/insertion frameshift and splice junction).
Several recent studies have examined genomic aberrations in the BCa cell lines.3, 23 There was complete overlap of TERT promoter mutations observed in our study and those reported in Borah et al.3 including c.-348G>C and c.-269G>A, both observed in a single line, and c.-57A>C observed in two lines. For the 25 lines studied here, Earl et al.23 reported 68 mutations in 10 genes and we confirmed 53 (78%) variants in our WES data (Supplementary Table 5). Our analysis also improved the annotation of several previously reported mutations.23 We determined that a deletion in the phosphatase and tensin homolog (PTEN) in line J82 was a homozygous loss of exons 7 and 8 (codons 212-342, NM_000314) and a deletion in RB1 in line TCCSUP was a homozygous loss of exons 18–27 (NM_000321). This comparison revealed that our knowledge of cancer gene alterations in these cell lines was far from complete, and accurate mutation detection and annotation is required to overcome inter-study variability that could have a significant impact on experiments investigating BCa cell biology and response to therapy.
Global and local copy number analysis
In addition to characterizing sequence alterations, we used WES to identify CNAs. Using the ExomeDepth method,28 we identified duplications and deletions in each of the 25 BCa cell lines. The median length of duplications was 307 kb and was 51 kb for deletions. Similarly, the CCLE analyzed a total of 26 BCa cell lines using Affymetrix SNP6.0 chips10 and Earl et al.23 assayed 40 BCa cell lines using Illumina Beadarrays. A total of 13 cell lines were assayed using all three platforms. Despite the different platforms and underlying data analyses, we found a high degree of overlap between loci with CNA using ExomeDepth (Figure 4). On average, 75% of ExomeDepth-identified duplications overlapped with amplified regions detected in the CCLE and with 53% of those detected by Earl et al. ExomeDepth deletions overlapped with 64% of the depleted regions identified in the CCLE and 75% of those in Earl et al. (Figures 4b and c). For the same 13 cell lines, we compared CCLE and Earl et al. copy number values on a gene-by-gene basis. For 12 of these lines, we found an average Pearson correlation coefficient of 0.68 ranging from 0.47 to 0.81. The one exception was the UM-UC-3 line that showed a poor correlation of 0.17, potentially because of a misidentification in either the CCLE or Earl et al. (Supplementary Figure 2).
Global copy number comparison. (a) A total of 13 cell lines were examined for CNA across three data sets: exome sequencing presented here, the CCLE10 and Earl et al.23 Copy number data are shown for each of the three data sets with copy number deletions colored blue and copy number amplifications colored red. The percentage of amplifications and deletions was determined using the program ExomeDepth28 that overlapped with data from (b) Earl et al. or (c) the CCLE.
We identified homozygous sequence alterations using WES read counts and the relative signal intensity (RSI) of mutant and WT alleles29 in Sanger sequence chromatograms produced during variant validation (Supplementary Table 4). This produced evidence of sequence and CNAs at nucleotide resolution at the sites of sequence variation (Figure 5). Seven genes, including oncogenes TERT and KRAS, and the tumor suppressors, TP53, KDM6A, SYNE1, RB1 and MLL2, exhibit allelic imbalance (RSI⩾0.7) in multiple (> 2) lines. Homozygous alterations occurred in a unique pattern in each line and likely represent irreversible commitment by each cell line to the altered function.
Loss of BCa genes on chromosome 9p21
Previously characterized in BCa, chr 9p21.3 is a genomic loci frequently altered by somatic loss that likely targets the tumor suppressors, CDKN2A and CDKN2B.4, 28, 30, 31 These genes encode proteins that form a complex with CDK4 to block cell cycle G1 progression, and an alternative transcript in the CDKN2A locus (p14/alternative reading frame (ARF)) encodes a stabilizer of TP53 through sequestration of the oncogenic ubiquitin ligase, MDM2.32, 33
Given the importance of chr 9p21.3 in human BCa, we sought to identify the cell lines with homozygous deletion of this locus for use as experimental models. To define the deletion borders, we examined WES read depth using the Integrative Genomics Viewer (IGV).34 As shown in Figure 6a, and consistent with previous human BCa data from primary tumors,4, 30 we found genes that were coordinately lost with CDKN2A/2B, which should also be considered when using these lines as experimental reagents. Homozygous deletions extending to all or part of neighboring genes were observed in 12 of the 25 lines. CDKN2A was lost in all 12 and CDKN2B in 9 of the lines. Homozygous deletions included all (six lines) or part (three lines) of MTAP, encoding a key enzyme in polyamine metabolism, indicating it is likely completely inactive in nine lines with a frequency of alteration similar to CDKN2B. The transcription factor, DMRTA1, was lost in seven lines and deletions in five lines included a microRNA, mir31, and interferon epsilon (IFNE). The remaining deletions removed varying numbers of up to 16 genes residing in an alpha interferon gene cluster, and two lines possessed deletions extending to the RNA binding protein, ELAVL2.
Deletion of chromosome 9p21.3 locus genes. (a) NGS of genes on chr 9 near CDKN2A and CDKN2B were manually examined using the IGV34 for homozygous deletions. Lines with evidence of a homozygous deletion in an exon are displayed (green). The analysis was inconclusive for ELAVL2 exon 1 (right side) but no deletions were observed that extended to the neighboring gene, IZUMO family member 3. (b-d) The expression of the indicated chr 9 gene in lines with no homozygous deletion (left) as compared with lines with ⩾1 exon exhibiting a homozygous deletion (right). (b) MTAP expression in 18 WT lines and 7 lines with a deletion. (c) CDKN2A expression in 13 WT lines and 12 lines with a deletion. (d) CDKN2B expression in 16 WT lines and 9 lines with a deletion. Wilcoxon rank sum test was used to calculate P-values.
To determine the impact of these deletions on gene expression,22 we compared the normalized expression levels for MTAP, CDKN2A and CDKN2B in lines with a homozygous deletion to lines without a deletion (Figures 6b and d). In each comparison, loss of the gene was associated with a significant reduction of expression: MTAP by 1.3-fold, CDKN2A by 8.1-fold and CDKN2B by 1.3-fold (GSE5845).22 These findings were confirmed using independent expression values generated from 15 of the same BCa cell lines in the CCLE (Supplementary Figure 3). We were unable to analyze IFNE expression because the Affymetrix gene expression array lacked IFNE probes.
Finally, we examined the expression of proteins encoded by genes deleted on chr 9p21 using publically available data (http://www.proteinatlas.org). DMRTA1 was expressed at medium and high levels in normal urothelial tissue and in 52% of bladder tumors. IFNE was expressed in fewer healthy tissues and in 23% of cancers, including moderate expression in BCa, but not in healthy urothelium. IFNA1 showed limited expression in healthy tissues and in 7% of cancers, whereas IFNB1, located outside the deletions detected in the cell lines, was expressed in almost all healthy tissues and 100% of cancers, including BCa. No protein expression data were available for MTAP, however, we were able to confirm MTAP loss in tumors from 34/127 (27%) patients analyzed by the TCGA,4 confirming the relevance of MTAP deficiency in both cell lines and primary disease.
BCa cell lines recapitulate genomic aberrations in tumors
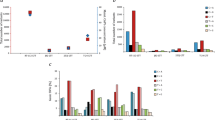
Small insertions and deletions (indels), and larger CNAs have been comprehensively characterized in both cell lines and BCa patient tumors. We used patient data from TCGA,4 cell line CNAs from Earl et al.,23 and the WES data presented here to calculate the ‘fraction of genome altered’ and ‘mutations per million bases.’ As shown in Figure 7a, the cell lines have a higher number of mutations, but lower overall number of CNAs (both significant P<0.01, Wilcoxon rank sum test). The numbers of alterations in the cell lines were within the range of genomic aberrations observed in the tumors. This is in contrast to the results reported by Domcke et al.17 for high-grade serous ovarian cancer where they found a set of commonly used cell lines were significantly hyper-mutated compared with patient tumors.
Genomic comparison of BCa cell lines to tumors. (a) Mutation frequency (x axis) compared with the fraction of the genome altered (y axis) for bladder tumors (blue) and cell lines (red). (b) Frequently altered genes identified from the TCGA analysis of BCa tumors4 are ordered by the percentage of samples with mutations in TCGA tumors as compared with the cell lines. Amplified, red; deleted, blue.
We also observed that many of the well-known BCa driver genes defined by TCGA4 showed similar frequencies of alterations in the cell lines when compared with patient tumors, suggesting that the cell lines are appropriate as experimental reagents to model bladder tumors (Figure 7b). The difference in frequency of alterations between the tumors and cell lines that are observed for some genes may be specific to the samples that were examined, but may also be biologically relevant and reflect the effects of selection during cell culture. For example, forkhead box Q1 encodes a transcription factor that regulates epithelial–mesenchymal transition35 and the reduced number of alterations in the cell lines as compared with patient tumors may reflect an essential role in cell line formation. Figure 7b also demonstrates that sequence alterations and CNAs occur as distinct mechanisms acting on individual cancer genes, with most genes primarily altered by a single mechanism, such as CDKN2A by CNA.
Cell line-derived signatures predict patient treatment response
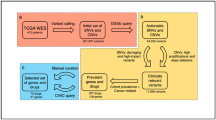
Cell lines are commonly used as pre-clinical drug screening platforms and are associated with large amounts of data generated across many cell lines and cancer types.10, 14, 15 It has been recognized that alignment of genomic profiles between cell lines and patients can be used to more effectively translate cell line drug response to patient cohorts.14 We took an integrative approach to characterize pathway activity including gene expression22, 36 and CNAs23 as inputs to PARADIGM to calculate inferred pathway activity (IPA)37 (Supplementary Table 6). We used elastic net regression38 and IPAs to build a predictive model of cisplatin response (Figure 8a). We used three-fold cross-validation to tune model parameters and ran 10 000 iterations to identify the most consistently selected IPAs. We built a final model using IPAs selected in 80% of the iterations, then used the model to predict platinum-based drug sensitivity in TCGA patient cohorts that were either treated with a platinum drug (n=44) or were not treated with chemotherapy as defined in the TCGA clinical information files (n=104). TCGA patient IPAs can be found in Supplementary Table 7. Kaplan–Meier survival analysis showed significant separation among patients treated with platinum-based drugs between those predicted to be sensitive and those predicted to be resistant (P=0.05, log-rank test). The patient cohort not treated with chemotherapy showed no separation between patients predicted to be either sensitive or resistant, consistent with this patient cohort being a negative control (Figure 8b).
Pathway signature from cisplatin-treated cell lines predicts patient response. (a) Cell lines are ranked ordered according to measured response to cisplatin. Heatmap of PARADIGM scores for the top concepts from the elastic net analysis. The PARADIGM class is in parentheses and the fraction of times the concept was selected over 10 000 iterations of model training is shown in brackets. Concepts selected in over 80% of the iterations were used in the final model. (c) Patients from the TCGA were separated into two groups according to whether they received a platinum-based chemotherapy or not treated with chemotherapy. The cisplatin signature derived from the cell lines was used to predict sensitive and resistance in the two patient populations. Predictions were evaluated using a Kaplan–Meier survival plot (log-rank test).
Discussion
There has been little advance in chemotherapeutic or targeted therapies for BCa in the past 20 years and robust pharmacogenomic models are needed to study molecular mechanisms of response and resistance. Here, we show that BCa cell lines model the genomic features of patient tumors and these features can be used to model response to platinum drugs. The generation of genomic data and clinical trials that leverage these data are increasing (for example, MATCH (NCT02465060) and SWOG1314 (NCT02177695)), highlighting the importance of model systems such as cell lines to study the mechanisms of drug response and resistance.
The GDSC and CCLE remain the most comprehensive pharmacogenomic cell line collections, yet both data sets lack comprehensive mutational profiling. According to a recent survey of major BCa genomics data sets, there are 34 genes frequently and significantly mutated by somatic alterations in BCa tumors.24 Many of these newly identified cancer genes have not been characterized in commonly utilized BCa cell lines, including important chromatin-associated genes such as STAG2 and MLL2.10, 39 Over the 25 bladder cell lines analyzed using WES, we report 9042 genes that have at least one sequence variant or indel (Supplementary Table 3), many of which remain to be further studied, and representing a major improvement to the current state of mutational profiling for these lines.
Importantly, we found that the bladder lines replicate many aberrations associated with BCa tumor development. Analysis of the alterations in the cell lines revealed remarkably similar overall proportions of altered sequence motifs when compared with MI primary tumors;24 this demonstrates a larger fraction of cancer-associated variants (40.7%) are associated with APOBEC enzymes and altered CpG sites. This analysis likely underestimates the significance of these alterations as we did not count variants that create new CpGs with unknown regulatory impact, and there is an unknown proportion of germline variants in the cell lines that will dilute the somatic mutation signal.
The chr 9p21.3 deletion typically associated with CDKN2A, p14ARF and CDKN2B loss31 was frequently observed in the cell lines, similar to primary tumors; however, to date the presence, extent and effect of the deletions on the surrounding genes has been unclear in BCa and in these lines, representing key experimental gaps in these important reagents. Thus, these data indicate these genes—MTAP and possibly others on chr 9p21- are BCa-associated genes that are distinct from CDKN2A, p14ARF and CDKN2B. A review of published studies confirms they are targeted for frequent alteration in a wide range of cancers primarily by CNA loss, including BCa in agreement with our data40 (Supplementary Figure 4). Additionally, recent work by Mavrakis et al.41 supports the importance of MTAP independent of CDKN2A.
These findings have implications for BCa etiology and treatment, and require consideration for future experiments in the ~50% of lines with homozygous deletions extending beyond CDKN2A and CDKN2B. For example, MTAP encodes a key enzyme in the salvage of adenine and methionine in polyamine metabolism and loss may contribute to metabolic changes that are commonly observed in cancer.42 Loss of MTAP leads to increased concentrations of its substrate, methylthioadenosine (MTA), which drives the reverse reactions in polyamine synthesis producing damaging hydrogen peroxide. These data indicate MTAP deficiency in tumor cells occurs at an appreciable frequency and polyamine synthesis may be an attractive therapeutic target that is particularly effective when applied to MTAP-deficient BCa.43, 44 Although a previous attempt to target polyamine biosynthesis through inhibition of ornithine decarboxylase (ODC1) did not prevent recurrence in low-risk superficial BCas,45 MTAP-deficient tumors were not analyzed and this distinction may have clarified a stratified patient response.
The alpha interferon gene cluster residing next to CDKN2A and MTAP, including the recently described IFNE, encodes cytokines that maintain interferon-regulated gene expression, inhibit cell proliferation, and protect against viral and bacterial infection.46, 47 The constitutive expression of IFNE in epithelial cells of the female and male reproductive tract appears to indicate an essential role to protect against viral and microbial infection, and common sexually transmitted disease.45, 48 Thus, as IFNE is the most frequently lost type I interferon gene at the chr 9p21 locus, the loss may be associated with increased viral and microbial infections of the urinary tract and bladder. We hypothesize that anti-microbial agents, such as the Bacillus Calmette-Guérin (BCG) vaccine already used in BCa, or antiviral therapy may be particularly effective in IFNE-deficient BCa.
These data indicate sequence changes and CNA affects both copies of an unexpected number of cancer genes across BCa cell lines. The combinations of homozygous losses and gains shown in Figures 4 and 6 represent, in theory, complete commitment of a cell line to the altered functions and therefore represent a unique combination of therapeutic targets (a therapeutic fingerprint) that is specific to each individual cancer from which the cell lines are derived. Thus, this high-resolution WES analysis vastly improves the utility of these cell lines as experimental reagents for combinatorial testing of new and existing therapies against altered BCa genes. Comparisons of tumor-derived cell lines with patient samples have been performed in ovarian,17 head and neck,18 colorectal,19 melanoma20 and non-small cell lung21 cancers. Here we report a genomics resource for the BCa community (https://www.synapse.org/UC25) and demonstrate its use for a pharmacogenomics analysis that can be applied to other cancer types based on cell line profiling.
Materials and methods
Exome capture, sequencing, and variant analysis
All cell lines were fingerprinted using 15 short tandem repeat loci at the University of Colorado core facility. DNA from cell lines (3 μg) was quantitated by fluorimeter and agarose gel electrophoresis and fragmented. Exome capture was performed using the Agilent SureSelect Human All Exon 38 Mb kit (Santa Clara, CA, USA). The exome of the T24 cell line was collected using the Agilent SureSelect Human All Exon 50 Mb kit and was sequenced to a greater depth than the other cell lines for future comparative studies to related derivative lines. NGS was used to generate 100 bp, paired-end reads using a HiSeq 2000 platform (Illumina Inc., San Diego, CA, USA) and was analyzed using the Genome Analyzer Pipeline, v. 1.3 (Illumina Inc.) with standard parameters. Base calls produced by Illumina base calling software 1.7 were aligned to the human genome (hg19) reference sequence using BWA,49 and duplicate reads were removed by SAMtools.50 SOAPsnp51 and SAMtools were used to identify SNPs and short indels, respectively, and variants were annotated using ANNOVAR.52 We removed SNPs with a base quality <20 and/or covered by fewer than four high-quality sequence reads. We intentionally used liberal filtering as we intended to validate all variants of interest by a second method (PCR and Sanger sequencing). Identical variants called in >5 cell lines or in known segmental duplications (segdups) were also excluded from further analysis (Supplementary Table 3). The tri-nucleotide context of variants that were not observed in the 1000 Genomes Project samples53 was examined using twoBitToFa (http://hgdownload.soe.ucsc.edu/admin/exe/linux.x86_64/twoBitToFa). NGS analysis utilized the high-performance computational capabilities of the Biowulf Linux cluster at the National Institutes of Health (Bethesda, MD, USA) (http://biowulf.nih.gov). A subset of NGS-predicted variants that altered proteins (n=309) were examined using PCR and Sanger sequencing as previously described.2
Cell growth assay and quantitative reverse transcriptase–PCR
UM-UC-3 and TCCSUP cells were split at 1x105 cells per well onto a 24-well plate. After 24 h, cells were transfected with Smartpool of 4 different siRNAs (Dharmacon, Lafayette, CO, USA) for non-target control and hTERT at 25 nM conc. using Lipofectamine RNAiMax transfection reagent from Thermo Fisher Scientific (Waltham, MA, USA). After 5 days of transfection, cells were trypsinized, resuspended in 1 ml culture media and live cells were counted by Trypan blue exclusion method. To confirm the knockdown of hTERT, cells were harvested 24 h after transfection and quantitative reverse transcriptase–PCR was performed. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as an internal control gene. Expression was normalized to control-treated cells to determine gene expression in knockdown cells. Primer sets used for quantitative reverse transcriptase–PCR are, hTERT forward 5′-CATTTCATCAGCAAGTTTGGAAG-3′ and reverse 5′-TTTCAGGATGGAGTAGCAGAGG-3′ and GAPDH forward 5′-TCTTTTGCGTCGCCAGCCGA-3′ and reverse 5′-ACCAGGCGCCCAATACGACC-3′. Triplicates were run for each cell line and values are indicated as mean±s.e.
Copy number and gene expression analysis
To identify deletions on chr 9, we examined NGS sequence depth for each exon in genes in a 3-Mb region of chr 9p21.3 encompassing CDKN2A and CDKN2B and visualized using the IGV.32 We observed >95% fewer NGS reads in a subset of lines that likely indicated homozygous deletions. On average, <3 NGS reads per exon were observed in the deleted region, which was used as the cutoff to define a homozygous deleted exon. Putative heterozygous losses of one allele were noted in additional cell lines, however, determining the borders of the deletions (our goal) was lower confidence.28 The gene expression (GSE5845)22 for CDKN2A, CDKN2B, MTAP and IFNA8 were available, but other genes and miRNAs were not. IFNA8 was not analyzed because of few lines with a deletion (n=4) and a low IFNA8 expression across all lines. Gene expression differences between groups of cell lines were tested using the Wilcoxon rank sum test.
For copy number comparisons, the gene coordinates were mapped to the copy number segments defined in the CCLE and Earl et al.10, 23 The log-fold change measures were transferred to the genes in the copy number segments and genes overlapping segments were removed. These values were used to compare CNA measurements in the CCLE with those in Earl et al.
Focal CNAs from exome sequencing data
WES data for all cell lines were pooled and simultaneously analyzed using ExomeDepth28 according to the script provided in the Supplementary Text. As described in Plagnol et al.,28 CNAs were called according to the expected vs observed reads based on a beta-binomial model. The reference set of cell lines was constructed by ranking all cells against the test cell line using correlation, then sequentially adding cell lines, fitting the model and testing the posterior probability in favor of a single-exon heterozygous deletion model. CNAs predicted by ExomeDepth were compared with segmented copy number profiles from the CCLE10 and Earl et al.23 Data processing, normalization and segmentation from CCLE and Earl et al. were done according to the associated publications. A total of 13 cell lines were shared across all 3 data sets (WES, CCLE and Earl et al.). An ExomeDepth CNA was called a positive overlap if the segment mapped to CCLE or Earl et al. showed a positive sign for duplications and negative sign for deletions.
TCGA patient data
RNAseq V2 gene expression, SNP6 CNAs, clinical and survival data of BCa (BLCA) patients was collected from TCGA.4 For further analyses, the data were filtered to only include patients with MI disease (T2–T4) according to the TCGA clinical file. Drug treatment information for all patients in the TCGA cohort was taken from a clinical data file (https://tcga-data.nci.nih.gov/tcga/dataAccessMatrix.htm).
Calculating fraction of genome altered and mutations per million bases
The calculations of the fraction of genome altered and mutations per million bases was described by Domcke et al.17 Cell line CNAs were smoothed and segmented using the DNAcopy R package54 for the processed Illumina Human1M-Duov3 DNA Analysis BeadChip (GSE64572),23 along with the probe locations (GPL6984). The TCGA patient segmentation files were downloaded from the GDAC Firehose website (doi:10.7908/C16T0KXB). A threshold of 0.3 was used for the cell lines and 0.2 for the patient tumors.
The TCGA patient mutation rates were taken from previous calculations.11 For each cell line, the number of bases with at least eight reads that mapped to the SureSelect Human All Exon 38 Mb Kit BED file defined the number of bases covered. Mutations were counted if they met the following criteria: quality score ⩾10, allele frequency ⩾0.1, depth ⩾8, missense, stopgain, or frameshift mutation, and not found in dbSNP (v138). The Wilcoxon rank sum test was used to compare the distribution of mutations and copy numbers between patient and cell lines.
UC40 gene expression and dose response
We used the gene expression (GSE5845) and dose response for cisplatin for the UC40 BCa cell lines as reported in Lee et al.22
PARADIGM analyses
PARADIGM (http://sbenz.github.io/Paradigm/), an integrative -omics tool for inferring pathway activities, was used to calculate IPA by integrating CNA and gene expression data for each patient and cell line.37 Patient input data consisted of the TCGA gene expression (RSEM55 normalized RNAseq) and segmented, log-transformed copy number values from GISTIC2.4 Cell line input data consisted of RMA-normalized gene expression22 and segmented, log-transformed copy number values from Earl et al.23 using GISTIC2. A total of 20 (of the 25) cell lines had both gene expression and CNA data. Gene expression and CNA data were normalized according to the PARADIGM documentation.37 The UCSC ‘Super Pathway’ input file was used: https://www.synapse.org/#!Synapse:syn1528312.
Elastic net predictions and survival analysis
The elastic net model (glmnet R package)56 was trained on cell line PARADIGM scores (IPAs) with the response variable being the corresponding cisplatin GI50 values.9 Only IPAs that showed a Spearman correlation P-value ⩽0.05 when compared with the GI50 values were included. Optimal α and λ values were selected using glmnet’s cross-validation function by looping through α between 0 and 1 at a 0.05 interval. Mean squared error was used to evaluate performance and λ values were the largest value within 1 s.e. of the minimum error. The α and λ values with the smallest mean three-fold cross-validation error were chosen to train the final elastic net model. We performed 10 000 iterations of model training and built a final elastic net model using IPAs that were selected in over 80% of the iterations. The final model was used to predict drug response for each patient in the TCGA cohort.
Based on predicted drug response, patients were rank ordered and median-split into two cohorts, predicted sensitive and resistant to treatment. Using the TCGA clinical information, patients were grouped into platinum-treated (n=44) and no evidence of chemotherapy treatment (n=104) cohorts. Kaplan–Meier survival curves were calculated separately for the platinum-treated and no evidence of drug treatment cohorts (log-rank statistical test).
Code availability
The code used to generate the copy number calls is available as Supplementary Text and at: https://www.synapse.org/UC25. The code used to generate the predictions of cisplatin sensitivity is available at: https://www.synapse.org/UC25.
Abbreviations
- BCa:
-
bladder cancer
- CCLE:
-
the Cancer Cell Line Encyclopedia
- CNA:
-
copy number alteration
- GDSC:
-
the Genomics of Drug Sensitivity in Cancer
- MTA:
-
methylthioadenosine
- NGS:
-
next-generation sequencing
- SNP:
-
single-nucleotide polymorphism
- TCGA:
-
The Cancer Genome Atlas
- WES:
-
whole-exome sequencing.
References
Siegel RL, Miller KD, Jemal A . Cancer statistics, 2015. CA Cancer J Clin 2015; 65: 5–29.
Nickerson ML, Dancik GM, Im KM, Edwards MG, Turan S, Brown J et al. Concurrent alterations in TERT, KDM6A, and the BRCA pathway in bladder cancer. Clin Cancer Res 2014; 20: 4935–4948.
Borah S, Xi L, Zaug AJ, Powell NM, Dancik GM, Cohen SB et al. TERT promoter mutations and telomerase reactivation in urothelial cancer. Science (80- ) 2015; 347: 1006–1010.
The Cancer Genome Atlas Research Network, Network C genome AR. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 2014; 507: 315–322.
Guo G, Sun X, Chen C, Wu S, Huang P, Li Z et al. Whole-genome and whole-exome sequencing of bladder cancer identifies frequent alterations in genes involved in sister chromatid cohesion and segregation. Nat Genet 2013; 45: 1459–1463.
Iyer G, Al-Ahmadie H, Schultz N, Hanrahan AJ, Ostrovnaya I, Balar A V et al. Prevalence and co-occurrence of actionable genomic alterations in high-grade bladder cancer. J Clin Oncol 2013; 31: 3133–3140.
Riester M, Taylor JM, Feifer A, Koppie T, Rosenberg JE, Downey RJ et al. Combination of a novel gene expression signature with a clinical nomogram improves the prediction of survival in high-risk bladder cancer. Clin Cancer Res 2012; 18: 1323–1333.
Smith SC, Baras AS, Dancik G, Ru Y, Ding K-F, Moskaluk CA et al. A 20-gene model for molecular nodal staging of bladder cancer: development and prospective assessment. Lancet Oncol 2011; 12: 137–143.
Choi W, Porten S, Kim S, Willis D, Plimack ER, Hoffman-Censits J et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 2014; 25: 152–165.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C et al. Mutational landscape and significance across 12 major cancer types. Nature 2013; 502: 333–339.
Morrison CD, Liu P, Woloszynska-Read A, Zhang J, Luo W, Qin M et al. Whole-genome sequencing identifies genomic heterogeneity at a nucleotide and chromosomal level in bladder cancer. Proc Natl Acad Sci USA 2014; 111: E672–E681.
Cao W, Chi W-H, Wang J, Tang J-J, Lu Y-J . TNF-alpha promotes doxorubicin-induced cell apoptosis and anti-cancer effect through downregulation of p21 in p53-deficient tumor cells. Biochem Biophys Res Commun 2005; 330: 1034–1040.
Goodspeed A, Heiser LM, Gray JW, Costello JC . Tumor-derived cell lines as molecular models of cancer pharmacogenomics. Mol Cancer Res 2015; 14: 3–13.
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012; 483: 570–575.
Yu M, Selvaraj SK, Liang-Chu MM, Aghajani S, Busse M, Yuan J et al. A resource for cell line authentication, annotation and quality control. Nature 2015; 520: 307–311.
Domcke S, Sinha R, Levine DA, Sander C, Schultz N . Evaluating cell lines as tumour models by comparison of genomic profiles. Nat Commun 2013; 4: 2126.
Li H, Wawrose JS, Gooding WE, Garraway LA, Lui VWY, Peyser ND et al. Genomic analysis of head and neck squamous cell carcinoma cell lines and human tumors: a rational approach to preclinical model selection. Mol Cancer Res 2014; 12: 571–582.
Mouradov D, Sloggett C, Jorissen RN, Love CG, Li S, Burgess AW et al. Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. Cancer Res 2014; 74: 3238–3247.
Lin WM, Baker AC, Beroukhim R, Winckler W, Feng W, Marmion JM et al. Modeling genomic diversity and tumor dependency in malignant melanoma. Cancer Res 2008; 68: 664–673.
Sos ML, Michel K, Zander T, Weiss J, Frommolt P, Peifer M et al. Predicting drug susceptibility of non-small cell lung cancers based on genetic lesions. J Clin Invest 2009; 119: 1727–1740.
Lee JK, Havaleshko DM, Cho H, Weinstein JN, Kaldjian EP, Karpovich J et al. A strategy for predicting the chemosensitivity of human cancers and its application to drug discovery. Proc Natl Acad Sci USA 2007; 104: 13086–13091.
Earl J, Rico D, Carrillo-de-Santa-Pau E, Rodríguez-Santiago B, Méndez-Pertuz M, Auer H et al. The UBC-40 Urothelial Bladder Cancer cell line index: a genomic resource for functional studies. BMC Genomics 2015; 16: 403.
Kim J, Akbani R, Creighton CJ, Lerner SP, Weinstein JN, Getz G et al. Invasive bladder cancer: genomic insights and therapeutic promise. Clin Cancer Res 2015; 21: 4514–4524.
Vuong H, Che A, Ravichandran S, Luke BT, Collins JR, Mudunuri US . AVIA v2.0: annotation, visualization and impact analysis of genomic variants and genes. Bioinformatics 2015; 31: 2748–2750.
Vuong H, Stephens RM, Volfovsky N . AVIA: an interactive web-server for annotation, visualization and impact analysis of genomic variations. Bioinformatics 2014; 30: 1013–1014.
Yap KL, Kiyotani K, Tamura K, Antic T, Jang M, Montoya M et al. Whole-exome sequencing of muscle-invasive bladder cancer identifies recurrent mutations of UNC5C and prognostic importance of DNA repair gene mutations on survival. Clin Cancer Res 2014; 20: 6605–6617.
Plagnol V, Curtis J, Epstein M, Mok KY, Stebbings E, Grigoriadou S et al. A robust model for read count data in exome sequencing experiments and implications for copy number variant calling. Bioinformatics 2012; 28: 2747–2754.
Nickerson ML, Jaeger E, Shi Y, Durocher JA, Mahurkar S, Zaridze D et al. Improved identification of von Hippel-Lindau gene alterations in clear cell renal tumors. Clin Cancer Res 2008; 14: 4726–4734.
Heidenblad M, Lindgren D, Jonson T, Liedberg F, Veerla S, Chebil G et al. Tiling resolution array CGH and high density expression profiling of urothelial carcinomas delineate genomic amplicons and candidate target genes specific for advanced tumors. BMC Med Genomics 2008; 1: 3.
Stadler WM, Sherman J, Bohlander SK, Roulston D, Dreyling M, Rukstalis D et al. Homozygous deletions within chromosomal bands 9p21-22 in bladder cancer. Cancer Res 1994; 54: 2060–2063.
Pomerantz J, Schreiber-Agus N, Liégeois NJ, Silverman A, Alland L, Chin L et al. The Ink4a tumor suppressor gene product, p19Arf, interacts with MDM2 and neutralizes MDM2’s inhibition of p53. Cell 1998; 92: 713–723.
Zhang Y, Xiong Y, Yarbrough WG . ARF promotes MDM2 degradation and stabilizes p53: ARF-INK4a locus deletion impairs both the Rb and p53 tumor suppression pathways. Cell 1998; 92: 725–734.
Thorvaldsdóttir H, Robinson JT, Mesirov JP . Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 2013; 14: 178–192.
Qiao Y, Jiang X, Lee ST, Karuturi RKM, Hooi SC, Yu Q . FOXQ1 regulates epithelial-mesenchymal transition in human cancers. Cancer Res 2011; 71: 3076–3086.
Havaleshko DM, Cho H, Conaway M, Owens CR, Hampton G, Lee JK et al. Prediction of drug combination chemosensitivity in human bladder cancer. Mol Cancer Ther 2007; 6: 578–586.
Vaske CJ, Benz SC, Sanborn JZ, Earl D, Szeto C, Zhu J et al. Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics 2010; 26: i237–i245.
Zou H, Hastie T . Regularization and variable selection via the elastic net. J R Stat Soc Ser B Stat Methodol 2005; 67: 301–320.
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 2013; 41: D955–D961.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2012; 2: 401–404.
Mavrakis KJ, McDonald ER 3rd, Schlabach MR, Gross BE, Billy E, Hoffman GR, deWeck A et al. Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5. Science 2016; 351: 1208–1213.
Scuoppo C, Miething C, Lindqvist L, Reyes J, Ruse C, Appelmann I et al. A tumour suppressor network relying on the polyamine–hypusine axis. Nature 2012; 487: 244–248.
Bertino JR, Waud WR, Parker WB, Lubin M . Targeting tumors that lack methylthioadenosine phosphorylase (MTAP) activity: current strategies. Cancer Biol Ther 2011; 11: 627–632.
Casero RA, Marton LJ . Targeting polyamine metabolism and function in cancer and other hyperproliferative diseases. Nat Rev Drug Discov 2007; 6: 373–390.
Messing E, Kim KM, Sharkey F, Schultz M, Parnes H, Kim D et al. Randomized prospective phase III trial of difluoromethylornithine vs placebo in preventing recurrence of completely resected low risk superficial bladder cancer. J Urol 2006; 176: 500–504.
Fung KY, Mangan NE, Cumming H, Horvat JC, Mayall JR, Stifter SA et al. Interferon-ɛ protects the female reproductive tract from viral and bacterial infection. Science 2013; 339: 1088–1092.
Wijesundara DDKD Xi Y, Ranasinghe C . Unraveling the convoluted biological roles of type I interferons in infection and immunity: a way forward for therapeutics and vaccine design. Front Immunol 2014; 5: 412.
Hermant P, Francius C, Clotman F, Michiels T . IFN-ɛ is constitutively expressed by cells of the reproductive tract and is inefficiently secreted by fibroblasts and cell lines. PLoS One 2013; 8: e71320.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
Li R, Li Y, Fang X, Yang H, Wang J, Kristiansen K et al. SNP detection for massively parallel whole-genome resequencing. Genome Res 2009; 19: 1124–1132.
Wang K, Li M, Hakonarson H . ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 2010; 38: e164.
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE et al. An integrated map of genetic variation from 1,092 human genomes. Nature 2012; 491: 56–65.
Seshan VV, Olshen A . DNAcopy. DNA Copy Number Analysis R package version 1.44.0. Bioconductor, software for bioinformatics Available from http://www.bioconductor.org.
Li B, Dewey CNN . RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 2011; 12: 323.
Friedman J, Hastie T, Tibshirani R . Regularization paths for generalized linear models via coordinate descent. J Stat Softw 2010; 33: 1–22.
Acknowledgements
We would like to thank Dr Neeraj Agarwal for providing the TERT experimental data. This work was in part supported by NIH grants CA143971 and CA075115 for DT; Intramural Research Program of the NIH, the National Cancer Institute, Center for Cancer Research for MLN and MD; National Library of Medicine T15 training grant LM009451 for NW; Shenzhen municipal science and technology project CXZZ20140826163906370 for SW.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Nickerson, M., Witte, N., Im, K. et al. Molecular analysis of urothelial cancer cell lines for modeling tumor biology and drug response. Oncogene 36, 35–46 (2017). https://doi.org/10.1038/onc.2016.172
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.172
This article is cited by
-
Erstlinientherapie beim metastasierten Nierenzellkarzinom
Die Onkologie (2024)
-
Human amniotic membrane inhibits migration and invasion of muscle-invasive bladder cancer urothelial cells by downregulating the FAK/PI3K/Akt/mTOR signalling pathway
Scientific Reports (2023)
-
Bladder cancer
Nature Reviews Disease Primers (2023)
-
Erstlinientherapie beim metastasierten Nierenzellkarzinom
best practice onkologie (2023)
-
Erstlinientherapie beim metastasierten Nierenzellkarzinom
Die Urologie (2023)