Abstract
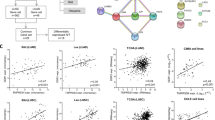
Non-small-cell lung cancer (NSCLC) is a common malignancy with a poor prognosis. Despite progress targeting oncogenic drivers, there are no therapies targeting tumor-suppressor loss. Smad4 is an established tumor suppressor in pancreatic and colon cancer; however, the consequences of Smad4 loss in lung cancer are largely unknown. We evaluated Smad4 expression in human NSCLC samples and examined Smad4 alterations in large NSCLC data sets and found that reduced Smad4 expression is common in human NSCLC and occurs through a variety of mechanisms, including mutation, homozygous deletion and heterozygous loss. We modeled Smad4 loss in lung cancer by deleting Smad4 in airway epithelial cells and found that Smad4 deletion both initiates and promotes lung tumor development. Interestingly, both Smad4−/− mouse tumors and human NSCLC samples with reduced Smad4 expression demonstrated increased DNA damage, whereas Smad4 knockdown in lung cancer cells reduced DNA repair and increased apoptosis after DNA damage. In addition, Smad4-deficient NSCLC cells demonstrated increased sensitivity to both chemotherapeutics that inhibit DNA topoisomerase and drugs that block double-strand DNA break repair by non-homologous end joining. In sum, these studies establish Smad4 as a lung tumor suppressor and suggest that the defective DNA repair phenotype of Smad4-deficient tumors can be exploited by specific therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ . Cancer statistics, 2009. CA Cancer J Clin 2009; 59: 225–249.
Goldstraw P, Ball D, Jett JR, Le Chevalier T, Lim E, Nicholson AG et al. Non-small-cell lung cancer. Lancet 2011; 378: 1727–1740.
Massague J . TGFbeta signalling in context. Nat Rev Mol Cell Biol 2012; 13: 616–630.
Malkoski SP, Wang XJ . Two sides of the story? Smad4 loss in pancreatic cancer versus head-and-neck cancer. FEBS Lett 2012; 586: 1984–1992.
Nagatake M, Takagi Y, Osada H, Uchida K, Mitsudomi T, Saji S et al. Somatic in vivo alterations of the DPC4 gene at 18q21 in human lung cancers. Cancer Res 1996; 56: 2718–2720.
Yanagisawa K, Uchida K, Nagatake M, Masuda A, Sugiyama M, Saito T et al. Heterogeneities in the biological and biochemical functions of Smad2 and Smad4 mutants naturally occurring in human lung cancers. Oncogene 2000; 19: 2305–2311.
Gemma A, Takenaka K, Hosoya Y, Matuda K, Seike M, Kurimoto F et al. Altered expression of several genes in highly metastatic subpopulations of a human pulmonary adenocarcinoma cell line. Eur J Cancer 2001; 37: 1554–1561.
Ke Z, Zhang X, Ma L, Wang L . Expression of DPC4/Smad4 in non-small-cell lung carcinoma and its relationship with angiogenesis. Neoplasma 2008; 55: 323–329.
Qiao W, Li AG, Owens P, Xu X, Wang XJ, Deng CX . Hair follicle defects and squamous cell carcinoma formation in Smad4 conditional knockout mouse skin. Oncogene 2006; 25: 207–217.
Teng Y, Sun AN, Pan XC, Yang G, Yang LL, Wang MR et al. Synergistic function of Smad4 and PTEN in suppressing forestomach squamous cell carcinoma in the mouse. Cancer Res 2006; 66: 6972–6981.
Bornstein S, White R, Malkoski S, Oka M, Han G, Cleaver T et al. Smad4 loss in mice causes spontaneous head and neck cancer with increased genomic instability and inflammation. J Clin Invest 2009; 119: 3408–3419.
Izeradjene K, Combs C, Best M, Gopinathan A, Wagner A, Grady WM et al. Kras(G12D) and Smad4/Dpc4 haploinsufficiency cooperate to induce mucinous cystic neoplasms and invasive adenocarcinoma of the pancreas. Cancer Cell 2007; 11: 229–243.
Kojima K, Vickers SM, Adsay NV, Jhala NC, Kim HG, Schoeb TR et al. Inactivation of Smad4 accelerates Kras(G12D)-mediated pancreatic neoplasia. Cancer Res 2007; 67: 8121–8130.
Bardeesy N, Cheng KH, Berger JH, Chu GC, Pahler J, Olson P et al. Smad4 is dispensable for normal pancreas development yet critical in progression and tumor biology of pancreas cancer. Genes Dev 2006; 20: 3130–3146.
Xu X, Kobayashi S, Qiao W, Li C, Xiao C, Radaeva S et al. Induction of intrahepatic cholangiocellular carcinoma by liver-specific disruption of Smad4 and Pten in mice. J Clin Invest 2006; 116: 1843–1852.
Xu X, Ehdaie B, Ohara N, Yoshino T, Deng CX . Synergistic action of Smad4 and Pten in suppressing pancreatic ductal adenocarcinoma formation in mice. Oncogene 2010; 29: 674–686.
Yang L, Mao C, Teng Y, Li W, Zhang J, Cheng X et al. Targeted disruption of Smad4 in mouse epidermis results in failure of hair follicle cycling and formation of skin tumors. Cancer Res 2005; 65: 8671–8678.
Glick AB, Weinberg WC, Wu IH, Quan W, Yuspa SH . Transforming growth factor beta 1 suppresses genomic instability independent of a G1 arrest, p53, and Rb [published erratum appears in Cancer Res 1997;57(10):2079]. Cancer Res 1996; 56: 3645–3650.
Kirshner J, Jobling MF, Pajares MJ, Ravani SA, Glick AB, Lavin MJ et al. Inhibition of transforming growth factor-beta1 signaling attenuates ataxia telangiectasia mutated activity in response to genotoxic stress. Cancer Res 2006; 66: 10861–10869.
Jalal S, Earley JN, Turchi JJ . DNA repair: from genome maintenance to biomarker and therapeutic target. Clin Cancer Res 17: 6973–6984.
Woods D, Turchi JJ . Chemotherapy induced DNA damage response: Convergence of drugs and pathways. Cancer Biol Therapy 2013; 14: 379–389.
Curtin NJ . DNA repair dysregulation from cancer driver to therapeutic target. Nat Rev Cancer 2012; 12: 801–817.
Shrivastav M, De Haro LP, Nickoloff JA . Regulation of DNA double-strand break repair pathway choice. Cell Res 2008; 18: 134–147.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 2012; 150: 1107–1120.
Cancer Genome Atlas Research N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 2012; 489: 519–525.
Caulin C, Nguyen T, Longley MA, Zhou Z, Wang XJ, Roop DR . Inducible activation of oncogenic K-ras results in tumor formation in the oral cavity. Cancer Res 2004; 64: 5054–5058.
Berton TR, Wang XJ, Zhou Z, Kellendonk C, Schutz G, Tsai S et al. Characterization of an inducible, epidermal-specific knockout system: differential expression of lacZ in different Cre reporter mouse strains. Genesis 2000; 26: 160–161.
Malkoski SP, Cleaver TG, Lu SL, Lighthall JG, Wang XJ . Keratin promoter based gene manipulation in the murine conducting airway. Int J Biol Sci 2010; 6: 68–79.
Malkoski SP, Haeger SM, Cleaver TG, Rodriguez KJ, Li H, Lu SL et al. Loss of transforming growth factor beta type II receptor increases aggressive tumor behavior and reduces survival in lung adenocarcinoma and squamous cell carcinoma. Clin Cancer Res 2012; 18: 2173–2183.
Crul M, van Waardenburg RC, Bocxe S, van Eijndhoven MA, Pluim D, Beijnen JH et al. DNA repair mechanisms involved in gemcitabine cytotoxicity and in the interaction between gemcitabine and cisplatin. Biochem Pharmacol 2003; 65: 275–282.
Bargonetti J, Champeil E, Tomasz M . Differential toxicity of DNA adducts of mitomycin C. J Nucleic Acids 2010; 2010.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Malkoski SP, Cleaver TG, Thompson JJ, Sutton WP, Haeger SM, Rodriguez KJ et al. Role of PTEN in basal cell derived lung carcinogenesis. Mol Carcinog 2013; 53: 841–846.
Kumar PA, Hu Y, Yamamoto Y, Hoe NB, Wei TS, Mu D et al. Distal airway stem cells yield alveoli in vitro and during lung regeneration following H1N1 influenza infection. Cell 2011; 147: 525–538.
Rock JR, Onaitis MW, Rawlins EL, Lu Y, Clark CP, Xue Y et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc Natl Acad Sci USA 2009; 106: 12771–12775.
Hong KU, Reynolds SD, Watkins S, Fuchs E, Stripp BR . Basal cells are a multipotent progenitor capable of renewing the bronchial epithelium. Am J Pathol 2004; 164: 577–588.
Papageorgis P, Cheng K, Ozturk S, Gong Y, Lambert AW, Abdolmaleky HM et al. Smad4 inactivation promotes malignancy and drug resistance of colon cancer. Cancer Res 2011; 71: 998–1008.
Yu SL, Lee DC, Son JW, Park CG, Lee HY, Kang J . Histone deacetylase 4 mediates SMAD family member 4 deacetylation and induces 5-fluorouracil resistance in breast cancer cells. Oncol Rep 2013; 30: 1293–1300.
Wyatt MD, Wilson DM . Participation of DNA repair in the response to 5-fluorouracil. Cell Mol Life Sci 2009; 66: 788–789.
Wachters FM, van Putten JW, Maring JG, Zdzienicka MZ, Groen HJ, Kampinga HH . Selective targeting of homologous DNA recombination repair by gemcitabine. Int J Radiat Oncol Biol Phys 2003; 57: 553–562.
Achanta G, Pelicano H, Feng L, Plunkett W, Huang P . Interaction of p53 and DNA-PK in response to nucleoside analogues: potential role as a sensor complex for DNA damage. Cancer Res 2001; 61: 8723–8729.
Adachi N, So S, Koyama H . Loss of nonhomologous end joining confers camptothecin resistance in DT40 cells. Implications for the repair of topoisomerase I-mediated DNA damage. J Biol Chem 2004; 279: 37343–37348.
Jin S, Inoue S, Weaver DT . Differential etoposide sensitivity of cells deficient in the Ku and DNA-PKcs components of the DNA-dependent protein kinase. Carcinogenesis 1998; 19: 965–971.
Treszezamsky AD, Kachnic LA, Feng Z, Zhang J, Tokadjian C, Powell SN . BRCA1- and BRCA2-deficient cells are sensitive to etoposide-induced DNA double-strand breaks via topoisomerase II. Cancer Res 2007; 67: 7078–7081.
Neal JA, Meek K . Choosing the right path: does DNA-PK help make the decision? Mutat Res 2011; 711: 73–86.
Kass EM, Jasin M . Collaboration and competition between DNA double-strand break repair pathways. FEBS Lett 2010; 584: 3703–3708.
Li W, Qiao W, Chen L, Xu X, Yang X, Li D et al. Squamous cell carcinoma and mammary abscess formation through squamous metaplasia in Smad4/Dpc4 conditional knockout mice. Development 2003; 130: 6143–6153.
Johnson L, Mercer K, Greenbaum D, Bronson RT, Crowley D, Tuveson DA et al. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 2001; 410: 1111–1116.
Wunderlich FT, Wildner H, Rajewsky K, Edenhofer F . New variants of inducible Cre recombinase: a novel mutant of Cre-PR fusion protein exhibits enhanced sensitivity and an expanded range of inducibility. Nucleic Acids Res 2001; 29: E47.
Nikitin AY, Alcaraz A, Anver MR, Bronson RT, Cardiff RD, Dixon D et al. Classification of proliferative pulmonary lesions of the mouse: recommendations of the mouse models of human cancers consortium. Cancer Res 2004; 64: 2307–2316.
Pan D, Schomber T, Kalberer CP, Terracciano LM, Hafen K, Krenger W et al. Normal erythropoiesis but severe polyposis and bleeding anemia in Smad4 deficient mice. Blood 2007; 110: 3049–3055.
Jackson EL, Olive KP, Tuveson DA, Bronson R, Crowley D, Brown M et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res 2005; 65: 10280–10288.
Jazag A, Kanai F, Ijichi H, Tateishi K, Ikenoue T, Tanaka Y et al. Single small-interfering RNA expression vector for silencing multiple transforming growth factor-beta pathway components. Nucleic Acids Res 2005; 33: e131.
Acknowledgements
This work was supported by the NIH/NCI (K08 CA131483 to SPM) and NIH/NIDCR (R01 DE015953 to XJW). SPM was also supported by the National Lung Cancer Partnership and a Career Development Award from the Colorado Lung Cancer SPORE (P50 CA058187) and an American Cancer Society Institutional Research Grant (ACS IRG 57-001-53). The OHSU BioLibrary is supported by P30 CA069533. The Colorado Lung Cancer SPORE Tissue Bank is supported by P30 CA046934 and P50 CA058187. We thank OHSU BioLibrary staff for their assistance with patient samples. We also thank Jerry Haney, Christina Nall, Marina Lewis and Wilbur Franklin at the Colorado Lung SPORE for assistance acquiring patient samples and data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
This work has been funded by the NIH (to Drs Malkoski and Wang), the National Lung Cancer Partnership (to Dr Malkoski) and the American Cancer Society (to Dr Malkoski). The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Haeger, S., Thompson, J., Kalra, S. et al. Smad4 loss promotes lung cancer formation but increases sensitivity to DNA topoisomerase inhibitors. Oncogene 35, 577–586 (2016). https://doi.org/10.1038/onc.2015.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.112
This article is cited by
-
HPV-mediated regulation of SMAD4 modulates the DNA damage response in head and neck cancer
Journal of Experimental & Clinical Cancer Research (2022)
-
Correlation between loss of Smad4 and clinical parameters of non-small cell lung cancer: an observational cohort study
BMC Pulmonary Medicine (2021)
-
Preoperative clinical and tumor genomic features associated with pathologic lymph node metastasis in clinical stage I and II lung adenocarcinoma
npj Precision Oncology (2021)
-
Development of syngeneic murine cell lines for use in immunocompetent orthotopic lung cancer models
Cancer Cell International (2020)
-
miRNA-558 promotes gastric cancer progression through attenuating Smad4-mediated repression of heparanase expression
Cell Death & Disease (2016)