Abstract
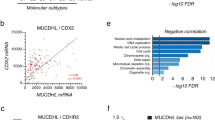
Desmosomal cadherins mediate cell–cell adhesion in epithelial tissues and have been known to be altered in cancer. We have previously shown that one of the two intestinal epithelial desmosomal cadherins, desmocollin-2 (Dsc2) loss promotes colonic epithelial carcinoma cell proliferation and tumor formation. In this study we show that loss of the other intestinal desmosomal cadherin, desmoglein-2 (Dsg2) that pairs with Dsc2, results in decreased epithelial cell proliferation and suppressed xenograft tumor growth in mice. Dsg2-deficient cells demonstrated a compensatory increase in Dsc2 expression, and small interfering RNA-mediated loss of Dsc2 restored proliferation in Dsg2-deficient cells. Dsg2 downregulation inhibited epidermal growth factor receptor (EGFR) signaling and cell proliferation through altered phosphorylation of EGFR and downstream extracellular signal-regulated kinase activation in parallel with inhibited EGFR receptor internalization. Additionally, we demonstrated a central role of Dsc2 in controlling EGFR signaling and cell proliferation in intestinal epithelial cells. Consistent with these findings, analyses of human colon cancers demonstrated increased Dsg2 protein expression. Taken together, these data demonstrate that partner desmosomal cadherins Dsg2 and Dsc2 play opposing roles in controlling colonic carcinoma cell proliferation through differential effects on EGFR signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Holthofer B, Windoffer R, Troyanovsky S, Leube RE . Structure and function of desmosomes. Int Rev Cytol 2007; 264: 65–163.
Getsios S, Amargo EV, Dusek RL, Ishii K, Sheu L, Godsel LM et al. Coordinated expression of desmoglein 1 and desmocollin 1 regulates intercellular adhesion. Differentiation 2004; 72: 419–433.
Tselepis C, Chidgey M, North A, Garrod D . Desmosomal adhesion inhibits invasive behavior. Proc Natl Acad Sci USA 1998; 95: 8064–8069.
Schlegel N, Meir M, Heupel WM, Holthofer B, Leube RE, Waschke J . Desmoglein 2-mediated adhesion is required for intestinal epithelial barrier integrity. Am J Physiol Gastrointest Liver Physiol 2010; 298: G774–G783.
Brennan D, Hu Y, Joubeh S, Choi YW, Whitaker-Menezes D, O'Brien T et al. Suprabasal Dsg2 expression in transgenic mouse skin confers a hyperproliferative and apoptosis-resistant phenotype to keratinocytes. J Cell Sci 2007; 120: 758–771.
Mannan T, Jing S, Foroushania SH, Fortune F, Wan H . RNAi-mediated inhibition of the desmosomal cadherin (desmoglein 3) impairs epithelial cell proliferation. Cell Prolif 2011; 44: 301–310.
Kolegraff K, Nava P, Helms MN, Parkos CA, Nusrat A . Loss of desmocollin-2 confers a tumorigenic phenotype to colonic epithelial cells through activation of Akt/β-catenin signaling. Mol Biol Cell 2011; 22: 1121–1134.
Biedermann K, Vogelsang H, Becker I, Plaschke S, Siewert JR, Hofler H et al. Desmoglein 2 is expressed abnormally rather than mutated in familial and sporadic gastric cancer. J Pathol 2005; 207: 199–206.
Brennan D, Mahoney MG . Increased expression of Dsg2 in malignant skin carcinomas: a tissue-microarray based study. Cell Adh Migr 2009; 3: 148–154.
Wex T, Kuester D, Monkemuller K, Stahr A, Fry LC, Kandulski A et al. Assessment of desmosomal components (desmoglein 1-3, plakoglobin) in cardia mucosa in relation to gastroesophageal reflux disease and Helicobacter pylori infection. Hum Pathol 2012; 43: 1745–1754.
Baron S, Hoang A, Vogel H, Attardi LD . Unimpaired skin carcinogenesis in Desmoglein 3 knockout mice. PLoS One 2012; 7: e50024.
Lock JG, Stow JL . Rab11 in recycling endosomes regulates the sorting and basolateral transport of E-cadherin. Mol Biol Cell 2005; 16: 1744–1755.
Getsios S, Simpson CL, Kojima S, Harmon R, Sheu LJ, Dusek RL et al. Desmoglein 1-dependent suppression of EGFR signaling promotes epidermal differentiation and morphogenesis. J Cell Biol 2009; 185: 1243–1258.
Biscardi JS, Maa MC, Tice DA, Cox ME, Leu TH, Parsons SJ . c-Src-mediated phosphorylation of the epidermal growth factor receptor on Tyr845 and Tyr1101 is associated with modulation of receptor function. J Biol Chem 1999; 274: 8335–8343.
Oksvold MP, Thien CB, Widerberg J, Chantry A, Huitfeldt HS, Langdon WY . Serine mutations that abrogate ligand-induced ubiquitination and internalization of the EGF receptor do not affect c-Cbl association with the receptor. Oncogene 2003; 22: 8509–8518.
Hynes NE, Lane HA . ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer 2005; 5: 341–354.
Ishii K, Norvell SM, Bannon LJ, Amargo EV, Pascoe LT, Green KJ . Assembly of desmosomal cadherins into desmosomes is isoform dependent. J Invest Dermatol 2001; 117: 26–35.
Nava P, Laukoetter MG, Hopkins AM, Laur O, Gerner-Smidt K, Green KJ et al. Desmoglein-2: a novel regulator of apoptosis in the intestinal epithelium. Mol Biol Cell 2007; 18: 4565–4578.
Resnik N, Sepcic K, Plemenitas A, Windoffer R, Leube R, Veranic P . Desmosome assembly and cell-cell adhesion are membrane raft-dependent processes. J Biol Chem 2011; 286: 1499–1507.
Chen YJ, Chang JT, Lee L, Wang HM, Liao CT, Chiu CC et al. DSG3 is overexpressed in head neck cancer and is a potential molecular target for inhibition of oncogenesis. Oncogene 2007; 26: 467–476.
Kolegraff K, Nava P, Laur O, Parkos CA, Nusrat A . Characterization of full-length and proteolytic cleavage fragments of desmoglein-2 in native human colon and colonic epithelial cell lines. Cell Adh Migr 2011; 5: 306–314.
Shimoyama Y, Hirohashi S, Hirano S, Noguchi M, Shimosato Y, Takeichi M et al. Cadherin cell-adhesion molecules in human epithelial tissues and carcinomas. Cancer Res 1989; 49: 2128–2133.
Acknowledgements
This work was supported by the 12th Glaxo-SmithKline International Award from Japan Society of Immunology and Allergology in Otolaryngology (RK), an American Gastroenterological Association Research Scholar Award (PN), a Crohn’s and Colitis Foundation of America Career Development Award (PN), a CONACyT grant 175854 (PN) and National Institutes of Health grants DK061379, DK072564 (CAP), DK055679, DK059888 (AN) and DDRDC DK064399.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Kamekura, R., Kolegraff, K., Nava, P. et al. Loss of the desmosomal cadherin desmoglein-2 suppresses colon cancer cell proliferation through EGFR signaling. Oncogene 33, 4531–4536 (2014). https://doi.org/10.1038/onc.2013.442
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2013.442
Keywords
This article is cited by
-
DSG2 expression is low in colon cancer and correlates with poor survival
BMC Gastroenterology (2021)
-
Loss of desmoglein-2 promotes gallbladder carcinoma progression and resistance to EGFR-targeted therapy through Src kinase activation
Cell Death & Differentiation (2021)
-
DSG2 expression is correlated with poor prognosis and promotes early-stage cervical cancer
Cancer Cell International (2020)
-
VPS33B interacts with NESG1 to modulate EGFR/PI3K/AKT/c-Myc/P53/miR-133a-3p signaling and induce 5-fluorouracil sensitivity in nasopharyngeal carcinoma
Cell Death & Disease (2019)
-
Intercalated discs: cellular adhesion and signaling in heart health and diseases
Heart Failure Reviews (2019)