Abstract
It is difficult to explain the differential rates of progression of premalignant colonic lesions and differences in behaviour of morphologically similar lesions. Heterogeneity for microsatellite instability (MSI) and promoter methylation in driving these phenomena forward may explain this; however, no previous analysis has examined this in detail at the gland level, the smallest unit of colorectal premalignant lesions. We aimed to carry out an analysis of gland level genomic instability for MSI and promoter methylation. MSI occurred significantly more frequently (20%) in colonic glands than has previously been observed in whole colorectal polyps. Significant promoter methylation was seen in MLH1, PMS2, MLH3 and MSH3 as well as significant heterogeneity for both MSI and promoter methylation. Methylation and MSI may have a significant role in driving forward colorectal carcinogenesis, although in the case of MSI, this association is less clear as it occurs significantly more frequently than previously thought, and may simply be a passenger in the adenoma-carcinoma sequence. Promoter methylation in MLH1, MLH3, MSH3 and PMS2 was also found to be significantly associated with MSI and should be investigated further. A total of 273 colorectal glands (126 hyperplastic, 147 adenomatous) were isolated via laser capture microdissection (targeted at regions of MLH1 loss) from 93 colonic polyps and tested for MSI, and promoter methylation of the DNA mismatch repair genes MLH1, MSH2, MLH3, MSH6, PMS2, MGMT and MLH3 via methylation specific multiplex ligation-dependent probe amplification. Logistic regression modelling was then used to identify significant associations between promoter methylation and gland histological type and MSI status.
Similar content being viewed by others
Introduction
The development of a colorectal polyp from a dysplastic adenoma has been well characterised. However, given these known pathways, it is difficult to explain different rates of progression of premalignant lesions and also differences in behaviour between morphologically similar types of polyp. This may be because the histological changes used to identify premalignant lesions are apparent at a certain stage, but the underlying changes driving them, that is, microsatellite instability (MSI) or promoter methylation may already exist, but be undetected due to polyclonality for these types of instability within a polyp. Currently, it cannot be predicted, which early lesions will develop into cancer. Early, but genomic instability at the level of the dysplastic or hyperplastic gland leading to heterogeneity for MSI and promoter methylation within premalignant lesions may go some way towards explaining this.1
Investigation of colorectal adenoma and carcinoma clonality has been performed previously in other, limited studies, which have found a range of results from monoclonality2 in colonic adenomas and carcinomas and polyclonality in familial adenomatous polyposis adenomas.
It is also unclear at what stage defective mismatch repair (MMR) and thus, MSI starts. Kloor et al.3 examined crypt foci microdissected from the normal mucosa of patients with Lynch syndrome. They identified a novel type of lesion, the MMR deficient crypt focus within normal mucosa. They hypothesised that this demonstrated that biallelic MMR gene mutation occurs at a high frequency in Lynch syndrome carriers and at an early stage, but it does not progress to malignancy, given the low frequency of cancers observed. Yurgelun et al.4 also noted a strong correlation between the loss of MMR protein and MSI in Lynch syndrome patients.
There is a variation in the observed rates of MSI in sporadic colorectal adenomas (probably due to variation in the definition of MSI and because of technical issues in the interpretation of MSI) with rates of 1–7% being reported.5, 6, 7, 8, 9 Another possibility is that the developing tumour remains heterogeneous for MSI+ status. It is also unclear whether this phenomenon would be different in hyperplastic polyps.
MSI has been observed in sporadic colorectal adenomas associated with loss of MLH1 expression10 as a consequence of MLH1 promoter hypermethylation.11 However, a potential issue with this mechanism is that the link between MSI and MLH1 expression in sporadic tumours is considerably weaker than that for Lynch syndrome and may be via an alternate mechanism. If using MLH1 protein expression as a guide to allow in-situ detection of regions that are microsatellite unstable, it is important to analyse the methylation status of the other MMR genes that could be associated with MSI within this gland. Several studies have found heterogeneity for methylation within glands in the same polyp sample.1213
In summary, there are a variety of disparate studies examining heterogeneity for both MSI and methylation within colonic lesions, using many different techniques, differing samples and differing clinical criteria. Clearly, a comprehensive study is needed with clear criteria to understand the true distribution of heterogeneity within colonic lesions.
We hypothesised that genomic instability, specifically MMR deficiency, starts in very early colonic dysplasia, and is present in heterogeneous form in early tumours, resulting in MSI and this is associated with promoter methylation. We also hypothesised that the supposed hallmarks of genomic instability (that is, MSI+ or promoter methylation) occur in normal tissue and early tumours without gross evidence of underlying MMR deficiency.
Results
A total of 91 sporadic polyps from 91 patients were obtained, and a total of 273 glands were microdissected (range 2–4 per polyp). The average age of patients was 60.9 years (range 33–91 years), with 48/91 (52.7%) being male and 43/91 (47.3%) being female. Of the 91 polyps, 45/91 (49.5%) were adenomatous (30/45 tubular and 15/45 tubulovillous) and 46/91 (50.5%) were hyperplastic. There were no serrated adenoma samples in the group. Of the 45 adenomatous polyps, 27/45 (60.0%) were mildly dysplastic, 5/45 (11.1%) were moderately dysplastic and 17/45 (37.8%) were severely dysplastic. In terms of the size of polyps in the hyperplastic polyp group 2/46 (4.3%) were>1 cm in size, and in the adenomatous polyp group 8/45 (17.7%) were>1 cm in size. After microdissection of the polyps into 273 glands, there were 126/273 (46.2%) glands from classical hyperplastic polyps available for study and 147/273 (53.8%) glands that were adenomatous, of which 81/147 (55.1%) were mildly dysplastic, 15/147 (10.2%) moderately dysplastic and 51/147 (34.7%) severely dysplastic.
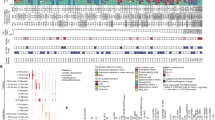
Within the 91 polyps, 154/273 glands (57%) had heterogeneous loss of MLH1 expression (that is, the polyp had some MLH1 expression, but this particular gland did not), with the remaining 119/273 glands (43%) having no loss of expression. There was no homogenous loss of MLH1 expression in any of the polyps studied (Figure 1).
The frequencies of BAT25 and BAT26 MSI in all glands were 21.0 and 21.5%, respectively. Results of PCR analysis by gland type for MSI is shown in Table 1, subdivided by dysplasia. In whole adenomatous polyps, 4/45 (8.9%) were MSI+ at the BAT25 locus and 3/45 (6.7%) were MSI+ at the BAT26 locus. For whole hyperplastic polyps, 4/46 (8.7%) were MSI+ at the BAT25 locus and 3/46 (6.5%) were MSI+ at the BAT 26 locus (see Table 1). On further analysis it was noted that 43/91 polyps were observed to be heterogeneous for gland level MSI within the sample polyp, that is, the glands studied within a single polyp were a mixture of MSI+ and microsatellite stable (MSS).
In order to examine the relationship between MLH1 expression status and MSI+ status, a 2 × 2 table of these variables was constructed (Table 2), for any one of BAT25 or BAT26. MSI status was significantly associated with the loss of MLH1 expression (Fishers exact, P=0.001), although a small proportion of glands had MSI without loss of MLH1 expression. The association between MLH1 expression and MSI+ status is most pronounced when looking at adenomatous and hyperplastic glands together. 155/273 (56.7%) glands were concordant for MLH1 loss and MSI, however, glands were preselected for MLH1 protein expression, which could bias this figure. However, not all glands with MLH1 loss identified in the prescreening were MSI.
Twenty-one MSI+ glands were sequenced for BRAF codon 600 mutations. One out of eleven adenomatous glands (9%) and 1/10 (10%) hyperplastic glands had codon 600 mutations. The adenomatous gland had a c.1799 T>A, P.600 V>E mutation, and the hyperplastic gland a c.1798 G>C, P.600 V>L mutation. These mutations were confirmed by repeat sequencing in both the forward and reverse directions. Interestingly the two mutations seen were not observed in other MSI+ glands within the same polyp.
Methylation specific multiplex probe ligand amplification (MS-MLPA) of colonic glands
In total, 114 glands (56 adenomatous glands and 48 hyperplastic glands), and 10 normal glands from normal tissue biopsies (in patients without polyps or cancer) were used in this study. Of the adenomatous glands, 20 were MSI+ and of the hyperplastic glands, 12 were MSI+. In total 114 MS-MLPA reactions were performed, with a failure rate of 28/114 (24.5%) all due to failure of hybridisation.
In the 10 microdissected glands obtained from normal mucosal samples, promoter methylation was consistently observed in probes MLH1-a (10/10 samples), MSH2-b (10/10 samples), MGMT-c (8/10 samples) and MSH3-b (9/10 samples). There was sporadic methylation at three other probes in two other normal control samples—37% methylation in probe MSH6-a and MSH2-c in one sample and 67% methylation in probe MSH3-a in another sample. Because of these findings, it was felt that the analysis of 10 normal gland samples was adequate as the rate of background methylation in normal samples was very low.
The relationship between MLH1 promoter hypermethylation and MLH1 protein expression via immunohistochemistry was also examined. (Table 3), and found that there was a significant association between MLH1 promoter methylation and loss of expression of MLH1. The MLH1 promoter region was said to be methylated if any of the MLH1 promoter region probes (excepting probe MLH1a that was methylated in all normal samples) had a methylation percentage exceeding our defined methylation threshold (20%). There was a significant association between MLH1 promoter hypermethylation and MLH1 protein expression loss (Fishers exact, Bonferroni corrected P=0.009). A table showing the relationship between methylation at each probe and gland type is shown in Supplementary Table 1.
In order to analyse the complex relationship between CpG island methylation, MSI status and histological type, a multivariate logistic regression model was constructed (see Supplementary Table S2 for full details). All MS-MLPA probes were entered into the regression model. Three separate models were constructed to ascertain, which methylated genes were significantly associated with MSI status; adenomatous polyps and hyperplastic polyps.
In examining, which MS-MLPA probes correlated with the presence of MSI within colonic glands, univariate analysis showed that methylation in MLH1 (probe B, P=0.03), PMS2 (probe B, P=0.041) and demethylation in MSH3 (probe B, P=0.019) and in MLH3 (probe B, P=0.026) were all associated significantly with MSI+ status. In multivariate analysis only methylation of MLH1 (probe B, P=0.029) remained significant Table 4).
For adenomatous glands, univariate analysis showed a significant association with methylation in PMS2 (probe C, P=0.013), however, after multivariate analysis demethylation of MLH1 (probe E, P=0.048) and methylation of MLH3 (probe A, P=0.048) became significant (Table 4).
For hyperplastic glands, on univariate analysis methylation in PMS2 (probe C, P=0.034) and MLH3 (probe B, P=0.04) was significant. On multivariate analysis demethylation in MSH3 (probe A, P=0.026) and MSH3 (probe B, P=0.003), and methylation in MLH3 (probe B, P=0.024) and PMS2 (probe B, P=0.029) were found to be significantly associated with hyperplastic glands (Table 4).
In order to explain the MSI positivity in glands that were not methylated at MLH1, the regression model was repeated, this time omitting the probes that were observed to be relevant previously (MLH1 probe B) or known to be methylated in normal tissue (MLH1 probe a, MLH1 probe f, MSH2 probe b and MLH3 probe a). In a univariate model (Supplementary Table S3), several probes were significant including methylation of PMS2 (probe A, P=0.024), de-methylation of MGMT (probe A, P=0.037) and MLH3 (probe B, P=0.022).
Discussion
This study has examined the presence and rates of MSI in isolated colorectal glands, showing that MSI does exist in isolated glands, and at a rate (20%) higher than has been previously reported in whole polyps tested for MSI.
Previous studies have shown the rates of MSI of ∼2–15%.14, 15 Previous attempts to analyse MSI in polyps may have reported14 lower rates of MSI+ because only a few glands within a polyp may be MSI+ and their signal is overwhelmed by the majority MSS DNA present within the polyp sample.
Of note is the level of heterogeneity for MSI seen in colorectal glands. In this study, just under 50% of all polyps showed heterogeneity for MSI (Table 1b). This suggests that MSI may exist at a higher rate than previously observed. Accelerated progression to cancer has been observed in tumours where MSI exists compared with MSS tumours, although this has not been observed consistently, and one possible explanation for this phenomenon is the presence of a clone of microsatellite unstable glands within an apparently MSS polyp. This may confer a selective growth advantage towards neoplasia (because the MSI crypt has a survival advantage of MSS crypts, leading to it becoming the dominant clone) even if the eventual tumour does not appear to be microsatellite unstable. It is also possible that MSI is a background phenomenon and is only present in but does not participate in the progression to carcinogenesis. Hypermethylation of the MLH1 promoter region has been seen in normal colonic mucosa at a very low level,16 but has not been associated with MSI. A possible further study would be to carry out MS-MLPA analysis of the whole colonic polyps to ascertain whether methylation status at the crypt level differs to that of the whole polyp.
The concordance between MLH1 expression and methylation is only ∼60% for all glands in our study, lower than would be expected if MLH1 methylation was solely responsible for MSI. This differs from the findings of Kloor et al.3 and Yurgelun et al.,4 however, their studies examined normal mucosa and polyps from Lynch syndrome carriers only. The mechanism by which this occurs in sporadic lesions may be different and may be related to methylation of other MMR gene CpG islands within these glands. We also noted heterogeneity for BRAF mutation status within the same polyp. This could represent the heterogeneity already seen for MSI within the polyp, or these BRAF mutations could simply be random passenger mutations occurring during polyp progression.
The hyperplastic glands in this study demonstrate an extensive pattern of methylation in the DNA repair genes, specifically with methylation occurring within the CpG islands of PMS2, MLH3 and most strongly, MSH3. Loss of MLH3 expression has been shown to lead to MSI,17 and MSH3 is known to form a heterodimer with MSH218 and participate in DNA repair.
These findings suggest that there may be other loci that influence MSI+ status and it is possible that these may be influenced by methylation in other MMR genes. Statistical analysis of this demonstrates a complex relationship, with promoter methylation of PMS2, MGMT and MLH3 being potentially significant in initiating MSI. The heterogeneity for methylation observed within glands in our study is widespread, leading to the possibility that the methylation seen is a random phenomenon that may or may not trigger MSI leading to accelerated polyp development.
A weakness of this study is that replicates were not obtained for every sample analysed via MS-MLPA, and that the MS-MLPA results were not verified by another technique such as bisulphite sequencing or pyrosequencing. This was not done because of the limitations of the small amounts of DNA inherent to laser capture microdissection. MS-MLPA has also been shown to be robust and reproducible in prior studies,19, 20 and thus was considered sufficiently reliable to be used in this manner. In addition, the use of both hyperplastic and adenomatous polyps in this study could potentially be criticised as biasing the results, as two separate pathways are thought to be involved in the initiation of these polyps. However, we feel that the relationship between MMR promoter methylation and MSI is likely to be ‘polyp independent’ as this is the only biologically plausible way in which this could occur.
In conclusion, MSI is present in individual colonic glands, and at a higher rate than previously demonstrated for whole polyps. The mechanism for this is not completely clear, but may partly be caused by MLH1 promoter methylation leading to inactivation of the gene as demonstrated in colorectal cancer cells in previous studies.21 This may be of clinical importance, as there is an increasing use of molecular stratification of tumours into MSI/MSS, and this study suggests that this phenomenon may be heterogenous. If this is the case, this will need to be carefully accounted for in any study of the link between MSI and clinical outcome.
This study also raises issues regarding the clonality of colorectal polyps. Our results would suggest that polyclonality exists in colorectal polyps, at least for MSI and for methylation. Two possible scenarios exist with regards to this finding, first that multiple pathways, in which a polyp arises can exist within the same polyp and contribute to the progression of the polyp, and that a single pathway becomes dominant towards the end of progression of the polyp. Alternatively, it is possible that this observed heterogeneity is of no consequence and that a single pathway is still dominant and the observed heterogeneity are merely ‘passenger’ events. Clearly further investigation is needed of this intriguing phenomenon.
There is a clear link between MMR gene promoter methylation and MSI status; however, it is unclear, excluding MLH1 methylation, which promoters cause a tendency to MSI+. Other MMR genes may have a role as suggested by this study. Little is known about the functional relevance of these genes and further study is needed to ascertain their relevance to colorectal cancer.
Materials and methods
Patients were recruited as from the SW London Colonoscopic Surveillance study (MREC 07/H0806/09). Patients who were >18 years old, were of Caucasian background, had at least one polyp at colonoscopy and with no family history or personal history of colorectal cancer or endometrial cancer (to the second degree level) were chosen.
Formalin-fixed, paraffin-embedded blocks were obtained and serial sections cut for H&E staining and immunohistochemistry and onto PEN laser capture slides for laser capture microdissection. Laser capture slides were stained with a 0.5% solution of methyl green dye (Sigma-Aldrich, Dorset, UK) to highlight cellular structure for microdissection. Immunohistochemistry against MLH1 was carried out using an indirect secondary method using mouse antihuman MLH1 monoclonal antibody at 1:50 dilution for 1 h at room temperature (BD Pharmingen G168-15, no.550838, Oxford, UK). MLH1 expression was scored as present or lost in either individual glands or the whole polyp when compared with the control tissue for consistency. Laser capture microdissection was targeted at glands where there was loss of MLH1 expression, on the basis that they were more likely to possess MSI. At least two separate glands with loss of MLH1 expression were microdissected per tumour, and one gland with normal MLH1 expression as a reference. If there was no loss of MLH1 expression, three randomly selected glands were microdissected. For comparison with glands, sections from the whole polyp were taken and DNA extracted. Normal control glands were microdissected from patients undergoing colonoscopy and who had no pathology present on colonoscopy (that is, those with a change in bowel habit and normal biopsies). All normal biopsy specimens were stained for MLH1 expression to confirm that there was no loss of expression.
DNA was extracted from laser capture microdissected glands using a PicoPure DNA (Molecular Devices Inc., Sunnyvale, CA, USA), kit was added and DNA was extracted according to the manufacturer’s instructions. An ethanol co-precipitation method was used to purify DNA using a glycogen carrier.
Analysis of MSI was carried out using BAT25 and BAT26 primers (sequences available on request) in a standard PCR reaction. MSI analysis was not carried out for dinucleotide repeats owing to insufficient sample DNA. Consequently this MSI panel did not make up the standard National Institutes of Health (NIH) panel for detection of MSI. For this study, MSI+ was defined as additional alleles at either of or both of the BAT25 and BAT26 loci compared with DNA from a known MSS normal colonic tissue donor. For BRAF V600E sequencing PCR, 1 μl of purified gland DNA was made up for a PCR reaction using a Qiagen Multiplex PCR kit under standard conditions (available on request). Negative controls consisted of a water well and tissue derived DNA from a patient known to be negative for somatic mutations in BRAF at V600E.
In order to carry out MS-MLPA, the quantity of DNA used in each reaction was standardised as 50 ng in 5 μl giving a concentration of 10 ng/μl. A ME-011 MS-MLPA kit (MRC Holland, Amsterdam, The Netherlands) was used to carry out MS-MLPA. This analyses MMR gene promoter methylation at MLH1, MSH2, MLH3, MSH6, PMS2, MGMT and MLH3 (Genomic locations of the probes can be found at http://www.mlpa.com). Controls consisted of a blank water well, a 100% unmethylated human pooled DNA control and a 100% methylated human pooled DNA control. PCR products were analysed on an ABI 3730xl automated capillary sequencer. Peak patterns from fragment analysis were inspected visually to ensure that they passed quality control according to the manufacturer’s instructions. If any peaks were present in the blank water well ,the sample plate was said to have failed. A custom-made Microsoft Excel spreadsheet (available on request) was used to quantify methylation at each allele by carrying out intrasample normalisation of each informative probe to several reference probes. Methylation was expressed as a percentage at each allele.
All statistical analyses were performed with STATA 11.1 (StataCorp, TX, USA). A observed frequency of methylation in normal tissue of 5% was assumed, based on the literature with a change in methylation rate in polyp samples to 50%, therefore it was calculated a minimum sample size of 23 would be needed in the adenoma and hyperplastic polyp groups each, assuming 90% power and a significance level of 0.05. In order to analyse rates vs levels of dysplasia, analysis of variance analysis was chosen, and for MSI+ vs MLH1 expression status, Fisher’s exact testing was carried out, owing to the small numbers in some cells of the 2 × 2 table.
In analysis of the MS-MLPA results, samples were called methylated if they exceeded the methylation threshold, which was set at 20% based on previous studies.19 In order to correct for interactions between each methylation probe, a reverse stepwise multivariate logistic regression model was carried out using either adenoma, HP, serrated or MSI+ as the independent variable and the methylation status of all probes as the dependent variables. In all situations where multiple testing was carried out, the Bonferroni correction was used unless otherwise stated.
References
Sieber OM, Heinimann K, Tomlinson IP . Genomic instability—the engine of tumorigenesis? Nat Rev Cancer 2003; 3: 701–708.
Fearon ER, Hamilton SR, Vogelstein B . Clonal analysis of human colorectal tumors. Science 1987; 238: 193–197.
Kloor M, Huth C, Voigt AY, Benner A, Schirmacher P, von Knebel Doeberitz M et al. Prevalence of mismatch repair-deficient crypt foci in Lynch syndrome: a pathological study. Lancet Oncol 2012; 13: 598–606.
Yurgelun MB, Goel A, Hornick JL, Sen A, Turgeon DK, MTt Ruffin et al. Microsatellite instability and DNA mismatch repair protein deficiency in Lynch syndrome colorectal polyps. Cancer prevention research 2012; 5: 574–582.
Brueckl WM, Jung A, Wein A, Brabletz T, Guenther K, Nusko G et al. Microsatellite instability in colorectal adenomas: relevance and clinical importance. Int J Colorectal Dis 2000; 15: 189–196.
Rashid A, Zahurak M, Goodman SN, Hamilton SR . Genetic epidemiology of mutated K-ras proto-oncogene, altered suppressor genes, and microsatellite instability in colorectal adenomas. Gut 1999; 44: 826–833.
Samowitz WS, Slattery ML . Microsatellite instability in colorectal adenomas. Gastroenterology 1997; 112: 1515–1519.
Samowitz WS, Slattery ML . Transforming growth factor-beta receptor type 2 mutations and microsatellite instability in sporadic colorectal adenomas and carcinomas. Am J Pathol 1997; 151: 33–35.
Lothe RA, Andersen SN, Hofstad B, Meling GI, Peltomaki P, Heim S et al. Deletion of 1p loci and microsatellite instability in colorectal polyps. Genes Chromosomes Cancer 1995; 14: 182–188.
Lanza G, Gafa R, Maestri I, Santini A, Matteuzzi M, Cavazzini L . Immunohistochemical pattern of MLH1/MSH2 expression is related to clinical and pathological features in colorectal adenocarcinomas with microsatellite instability. Mod Pathol 2002; 15: 741–749.
Cunningham JM, Christensen ER, Tester DJ, Kim CY, Roche PC, Burgart LJ et al. Hypermethylation of the hMLH1 promoter in colon cancer with microsatellite instability. Cancer Res 1998; 58: 3455–3460.
Umetani N, de Maat MF, Sunami E, Hiramatsu S, Martinez S, Hoon DS . Methylation of p16 and Ras association domain family protein 1a during colorectal malignant transformation. Mol Cancer Res 2006; 4: 303–309.
Belshaw NJ, Pal N, Tapp HS, Dainty JR, Lewis MP, Williams MR et al. Patterns of DNA methylation in individual colonic crypts reveal aging and cancer-related field defects in the morphologically normal mucosa. Carcinogenesis 2010; 31: 1158–1163.
Aaltonen LA, Peltomaki P, Leach FS, Sistonen P, Pylkkanen L, Mecklin JP et al. Clues to the pathogenesis of familial colorectal cancer. Science 1993; 260: 812–816.
Koh DC, Luchtefeld MA, Kim DG, Attal H, Monroe T, Ingersoll K . Microsatellite instability and MLH1 hypermethylation - incidence and significance in colorectal polyps in young patients. Colorectal Dis 2007; 9: 521–526.
Ahlquist T, Lind GE, Costa VL, Meling GI, Vatn M, Hoff GS et al. Gene methylation profiles of normal mucosa, and benign and malignant colorectal tumors identify early onset markers. Mol Cancer 2008; 7: 94.
Lipkin SM, Wang V, Jacoby R, Banerjee-Basu S, Baxevanis AD, Lynch HT et al. MLH3: a DNA mismatch repair gene associated with mammalian microsatellite instability. Nat Genet 2000; 24: 27–35.
Guerrette S, Wilson T, Gradia S, Fishel R . Interactions of human hMSH2 with hMSH3 and hMSH2 with hMSH6: examination of mutations found in hereditary nonpolyposis colorectal cancer. Mol Cell Biol 1998; 18: 6616–6623.
Jeuken JW, Cornelissen SJ, Vriezen M, Dekkers MM, Errami A, Sijben A et al. MS-MLPA: an attractive alternative laboratory assay for robust, reliable, and semiquantitative detection of MGMT promoter hypermethylation in gliomas. Lab Invest 2007; 87: 1055–1065.
Park CK, Kim J, Yim SY, Lee AR, Han JH, Kim CY et al. Usefulness of MS-MLPA for detection of MGMT promoter methylation in the evaluation of pseudoprogression in glioblastoma patients. Neuro Oncol 2011; 13: 195–202.
Poynter JN, Siegmund KD, Weisenberger DJ, Long TI, Thibodeau SN, Lindor N et al. Molecular characterization of MSI-H colorectal cancer by MLHI promoter methylation, immunohistochemistry, and mismatch repair germline mutation screening. Cancer Epidemiol Biomarkers Prev 2008; 17: 3208–3215.
Acknowledgements
ADB was funded by a Mason Medical Foundation Research Fellowship and grants from the Peel Medical Research Trust and St George’s Hospital Charity; IPMT acknowledges funding from Cancer Research UK and the Wellcome Trust 090532/Z/09/Z.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Beggs, A., Domingo, E., Abulafi, M. et al. A study of genomic instability in early preneoplastic colonic lesions. Oncogene 32, 5333–5337 (2013). https://doi.org/10.1038/onc.2012.584
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.584
Keywords
This article is cited by
-
Mutational analysis of driver genes defines the colorectal adenoma: in situ carcinoma transition
Scientific Reports (2022)
-
Update on Lynch syndrome genomics
Familial Cancer (2016)
-
DNA hypermethylation appears early and shows increased frequency with dysplasia in Lynch syndrome-associated colorectal adenomas and carcinomas
Clinical Epigenetics (2015)
-
Genomic instability in pre-neoplastic colonic lesions
Oncogene (2013)