Abstract
The human protein methyltransferases (PMTs) constitute a large enzyme class composed of two families, the protein lysine methyltransferases (PKMTs) and the protein arginine methyltransferases (PRMTs). Examples have been reported of both PKMTs and PRMTs that are genetically altered in specific human cancers, and in several cases these alterations have been demonstrated to confer a unique dependence of the cancer cells on PMT enzymatic activity for the tumorigenic phenotype. Examples of such driver alterations in PMTs will be presented together with a review of current efforts towards the discovery and development of small-molecule inhibitors of these enzymes as personalized cancer therapeutics.
Similar content being viewed by others
Introduction
Genetic material is packaged within the nuclei of cells in the form of chromatin, a noncovalent complex of chromosomal DNA and the core nucleosomal histone proteins around which the DNA is wrapped (Figure 1a). The tightly condensed chromatin (referred to as heterochromatin) thus allows large amounts of chromosomal DNA to be packaged within the relatively small volume of the cellular nucleus. Activated gene transcription is enabled by conformational changes that permit the access of transcriptional machinery (that is, transcription factors, polymerases and the like) to specific gene promoters by de-condensing the chromatin (referred to as euchromatin; Figure 1b). The conformational transition between euchromatin and heterochromatin states is controlled by a collection of posttranslational modification of the histone proteins and covalent modifications of DNA.1, 2, 3 These covalent modifications of chromatin include DNA methylation (at CpG sites), lysine acetylation/deacetylation, lysine and arginine methylation/demethylation, lysine ubquitinylation/deubquitinylation, serine phosphorylation/dephosphorylation and others; each of these covalent modifications is catalyzed by specific group-transfer enzymes.4, 5 Alterations in the catalytic activity of any of these enzymes can lead to pathogenic alterations in the pattern of gene transcription within a particular cell type. For example, alterations in the enzymatic activity of specific protein methyltransferases (PMTs) can diminish the transcription of tumor suppressor genes or enhance the transcription of oncogenes—either mechanism leading to a hyperproliferative, tumorigenic phenotype6, 7 (vide infra).
(a) Covalent modifications of chromatin include methylation of the chromosomal DNA at CpG islands and posttranslational modifications of histone proteins. Among these posttranslational histone modifications are methylation reactions at the side chains of arginine and lysine residues that are catalyzed by the PMT enzyme families RMTs and KMTs, respectively. (b) Chromatin exists in two major conformational states: a relaxed, transcriptionally permissive state known as euchromatin, and a condensed, transcriptionally repressive state known as heterochromatin. The transition between these two conformational states is facilitated by the collection of posttranslational modifications of histone proteins, which are summarized in panel (a) of this figure. (c) The constituency of chromatin-modifying enzymes in humans. The numbers in parentheses indicate the number of enzymes of each family that are found in humans. (d) Methylation states of lysine and arginine that are catalyzed by KMTs and RMTs, respectively.
Surveying the constituency of chromatin-modifying enzymes in humans6 reveals that >50% of these enzymes are involved in the reversible transfer of methyl groups on to and off of lysine or arginine residues of histone proteins (Figure 1c). In particular, the PMTs constitute a large class of enzymes composed of two families:8 the protein lysine methyltransferases (PKMTs) and the protein arginine methyltransferases (PRMTs); a number of examples of genetic alterations in PMTs have been identified in specific human cancers, as will be discussed below. The reactions catalyzed by PMTs have a paramount role in controlling gene transcription, as they result in eight distinct chemical states of lysine and arginine (0, 1, 2, or 3 methyl groups on lysine and 0, 1, 2-symmetric and 2-asymmetric on arginine; Figure 1d), each of which can confer unique transcriptional, and hence phenotypic, effects on cells.9
In this review we shall focus our attention on the aberrant activity of PMTs that drive tumorigenesis in specific human cancers and current efforts to identify potent and selective small-molecule inhibitors of these enzymes as a basis for personalized cancer therapeutics.
Catalytic mechanism of PMTs
The PMTs catalyze the common chemical reaction of SN2 transfer of a methyl group from the universal methyl donor S-adenosylmethionine (SAM) to the nitrogen atom(s) of a lysine or arginine side chain7 and sometimes histidine or the amino terminus of the polypeptide chain10 (Figure 2a). In all PMTs that have thus far been studied, the reaction involves direct transfer of the methyl group from SAM to the nitrogen acceptor; thus, both SAM and the protein substrate must be simultaneously bound to the enzyme in such a way as to achieve correct molecular orbital alignment and proximity for methyl transfer.7 The reaction mechanism is thus referred to as a ternary complex bi–bi mechanism, indicating that the enzymes convert two substrates (SAM and the protein) to two products (SAH (S-adenosylhomocysteine) and the methylated peptide) through requisite formation of a ternary enzyme–SAM–protein complex.11 The crystal structures of a large number of PMTs have been solved, all showing a distinct SAM binding pocket adjacent to a well-defined lysine- or arginine-binding channel.12 The termini of the SAM and amino-acid binding sites intersect to create favorable positioning of the two substrates for facile methyl group transfer (Figure 2b). As will be discussed below, both the SAM and amino-acid binding pockets provide structural loci for potential inhibitor interactions.5, 6 Thus, the structural features of PMTs required for catalysis make these enzymes attractive targets for intervention by small, drug-like inhibitors.5, 7, 13, 14
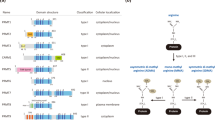
(a) Generic SN2 methyl transfer reaction from SAM to the amino-acid side-chain nitrogen of a lysine (or arginine) residue, as catalyzed by PMTs. The reaction shown is for methylation of a lysine residue. (b) Crystal structure of the PMT G9a illustrating the juxtaposition of the SAM binding site and the lysine binding site. In this structure the lysine binding channel is occupied by the small-molecule inhibitor UNC0224.73 (c) Family trees for the human protein lysine (left) and arginine (right) methyltransfersases. Data from Richon et al.8
Despite the commonality of chemical mechanism catalyzed, the PMTs display a remarkable degree of structural diversity within the SAM and amino-acid binding pockets; the relatedness of the SAM binding pockets of the human PMTs has been studied and mapped onto a set of family trees for the PKMTs and the PRMTs as illustrated in Figure 2c.8 This structural diversity portends the ability to design selective inhibitors that engage unique recognition elements within the natural ligand binding sites. Additionally, the PMTs display a considerable degree of substrate specificity with respect to the exact lysine or arginine residue that is methylated. Protein substrate specificity is likely conferred by recognition elements external to the SAM and amino-acid binding pockets that engage complementary elements within the amino-acid sequence of the histone. Thus, one finds generally that a limited number of PMT enzymes catalyze methylation at specific histone sites.2 For example, the enzyme DOT1L is the only human PMT known to catalyze methylation of histone H3 at lysine 79 (H3K79).15 Likewise, the multiprotein polycomb repressive complex 2 (PRC2), containing the catalytic subunit EZH2 or EZH1, is the only known catalyst of H3K27 methylation.16 Substrate specificity is somewhat more relaxed among the PRMTs, several of which have been demonstrated to methylate both nuclear and cytosolic protein substrates;17 nevertheless, within a particular protein substrate, PRMTs tend to catalyze methyl addition to a limited number (often only one) of specific amino-acid sites. This stringency of substrate specificity provides a direct mechanism for biochemical and biological fidelity with respect to control of gene transcription.
Genetic alterations of PMTs in cancer
A number of PMTs have been shown to be genetically altered in human cancers, and in several cases these alterations have been demonstrated to be drivers of tumorigenesis.6, 7 Among the types of alterations seen within PMTs are overexpression, gene amplification, chromosomal translocations and point mutations. In many, but not all cases, the genetic alteration leads to a gain- or change-of-function in catalytic activity that can be directly correlated with the tumorigenic phenotype of the disease. Three examples of PKMTs will be discussed here that exemplify the breadth of genetic alterations seen among PMTs in human cancers: DOT1L, EZH2 and WHSC1.
DOT1L
DOT1L is unique among the PKMTs in that it does not contain the otherwise universal catalytic domain of lysine methyltransferases known as the SET domain.15 In fact, from a structural and pharmacologic perspective, DOT1L is better associated with the PRMTs, despite the demonstrated biochemical activity of this enzyme as a lysine methyltransferase.8 The enzyme uniquely catalyzes the mono-, di- and tri-methylation of H3K79. Methylated H3K79 is associated with active gene transcription. DOT1L activity has been demonstrated to be a required driver of proliferation in mixed lineage leukemia 1 protein (MLL)-rearranged leukemia, a devastating disease that affects patients of all ages.18, 19
MLL-rearranged leukemia is universally associated with a chromosomal translocation affecting the MLL gene at chromosomal location 11q23.20, 21 The translocation results in a protein fusion between the MLL protein (itself a SET-domain PKMT that loses its SET domain as a result of the chromosomal translocation) and any of a number of protein partners, mainly of the AF and ENL families.20, 21, 22 The amino terminus contains domains that are responsible for targeting the MLL protein to specific target genes and these domains are retained within the context of the fusion protein.23, 24, 25 MLL fusion proteins recruit several complexes that increase the expression of their target genes and include polymerase-associated factor complex, positive transcription elongation factor and DOT1L.26, 27, 28, 29 Polymerase-associated factor complex is recruited via the amino terminus of MLL, which remains intact in the MLL fusion protein. Polymerase-associated factor complex has an important role in initiation, elongation and termination of gene transcription. Positive transcription elongation factor, consisting of cyclin T and cdk9, is recruited to the complex via the fusion partners (ENL, ELL and AF4) and phosphorylates the carboxy terminal domain of RNA polymerase II. Interestingly, the wild-type MLL protein is also essential for leukemogenesis in MLL-rearranged leukemia.30 The fusion partner proteins also provide the binding domain that recruits DOT1L to the MLL fusion.31 Thus, DOT1L is recruited to ectopic gene locations, where it catalyzes the methylation of H3K79 and thereby induces aberrant transcription of a number of leukemogenic genes, including HOXA9 and MEIS1.32 Studies using either RNA silencing methods19 or selective small-molecule inhibitors18 of DOT1L have demonstrated that the catalytic activity of this enzyme is required for proliferation of MLL-rearranged leukemic cells. Thus, the dependence of MLL-rearranged leukemia on DOT1L activity results from an indirect mechanism of genetic alteration. Although the enzyme itself is not altered in the disease, the chromosomal translocation indirectly results in a unique dependency of MLL-rearranged leukemic cells on DOT1L activity for transformation.
In addition to having a role in MLL-rearranged leukemia, DOT1L may also have a role in colon cancer. Mahmoudi et al.33 identified AF10 and DOT1L as β-catenin-associated proteins in intestinal crypts and showed that DOT1L is essential for Wnt-dependent transcription.
WHSC1
WHSC1 (also referred to as MMSET or NSD2) is a SET-domain PKMT that catalyzes the dimethylation of H3K36.34, 35 Methylation of this histone site is associated with regions that are transcriptionally active. The t(4;14) chromosomal translocation occurs in a subset of about 15% of multiple myeloma cases36, 37 and patients with this translocation represent one of the worst prognostic subgroups of multiple myeloma.38 The t(4;14) translocation results in massive overexpression of WHSC1 and of fibroblast growth factor receptor 3 (FGFR3) due to the placement of the strong immunoglobulin H intronic Eμ enhancer and 3′ enhancer in the promoter regions of WHSC1 and FGFR3 genes, respectively. Although ca 30% of t(4;14) patients have lost expression of FGFR3, 100% retain overexpression of WHSC1, suggesting that WHSC1, rather than FGFR3, is the primary driver of the disease.38 The overexpression of WHSC1 in t(4;14) translocated cells results in significantly elevated levels of dimethylated H3K36, as would be expected from elevation of catalytic enzyme levels.39 Genetic knockdown of WHSC1 or disruption of the translocated allele in t(4;14) myeloma cells results in inhibition of cellular proliferation and of tumorigenicity. As expected, genetic knockdown of WHSC1 demonstrates an accompanying reduction in global levels of H3K36me2.39
EZH2
EZH2, or the closely related EZH1, is a SET-domain PKMT that represents the catalytic subunit of a multiprotein complex referred to as PRC2. At least four protein subunits are required for PKMT activity (EZH2/1, EED, SUZ12 and RbAp48) and the active PRC2 complex demonstrates a high degree of specificity for methylation of H3K27.16 PRC2 catalyzes three sequential methylation reactions at H3K27, resulting in mono-, di- and tri-methylated H3K27. The tri-methyl H3K27 mark has been shown to be associated with transcriptional silencing; among the genes whose transcription is silenced are several known tumor suppressors. Not surprisingly, then, mechanisms that lead to a hyper-trimethylated state of H3K27 have been found to be universally tumorigenic for a number of hematologic and solid tumors.40
Several mechanisms have been reported to lead to a hyper-trimethylated state of H3K27.40 These include overexpression of EZH2, amplification of EZH2 and/or other PRC2 subunits, and loss-of-function mutations in the corresponding H3K27 demethylase, UTX (also known as KDM6A). Recently, recurrent somatic mutations at tyrosine 641 of EZH2 (Y641F, Y641N, Y641S and Y641H) have been reported in a subgroup of patients with non-Hodgkin’s lymphoma (NHL).41 These point mutations have been demonstrated to also lead to a hyper-trimethylated state of H3K27 by a novel mechanism. The Y641 mutations were found to be heterozygous in NHL patients where equal amounts of wild-type and mutant enzyme were found (at both the mRNA and protein level).41 The wild-type enzyme was found to be a very efficient catalyst of H3K27 mono-methylation, but to wane in activity for the dimethylation and especially the trimethylation reaction. In direct contrast, the Y641 mutant EZH2 proteins showed the exact opposite substrate specificity; they were essentially inactive as H3K27 mono-methyltransferases, but active in taking the preformed H3K27me1 species to the H3K27me2 form and especially efficient at taking the H3K27me2 state to the H3K27me3 form.42 Thus, hyper-trimethylation of H3K27 in the mutant-bearing NHL cells is the result of requisite coupling of enzymatic activity between the wild-type and mutant forms of the enzyme. Hence, pathogenesis is only conferred in this situation when the wild-type and mutant enzymes are present in the context of heterozygosity. Another mutation of EZH2 has also been found to occur heterozygously in a subset of NHL patients. This more rare mutation occurs at alanine 677 (A677G).41 Recently McCabe et al.43 have shown that this mutation results in nearly equal efficiency for all three substrates and increased trimethylation of H3K27.
Genetic alterations of EZH2 and other PRC2 proteins are not limited to the Y641 and A677 mutations observed in lymphoma. A spectrum of genetic alterations of PRC2 has been documented in a variety of hematologic and solid tumors. Interestingly, in myeloid malignancies and T-cell leukemia, mutations in PRC2 subunits lead to loss of enzymatic function.44, 45, 46 The fact that both activating and inactivating mutations of EZH2 are associated with malignancy is remarkable and indicates the complex role of PRC2 target genes in cell fate decisions.
A recent study by Mochizuki-Kashio et al.47 showed that EZH2 is essential for fetal hematopoiesis but is not required for hematopoietic stem cells in adult bone marrow. Rather EZH1 is highly expressed in hematopoietic stems cells from adult bone marrow. Therefore, the development of compounds that selectively inhibit EZH2 with more limited impact on EZH1 could spare any potential effect on hematopoietic stem cells in the bone marrow.
Direct inhibitors of PMTs
The cumulative data suggesting that genetic alterations in PMTs drive tumorigenesis in a number of human cancers has generated considerable interest in the discovery and development of selective inhibitors of these enzymes as potential therapeutic agents for genetically-defined cancers.6, 7 For example, the overarching objective of our own group is to identify driver alterations of PMTs in human cancers that confer to the cancer cell a unique dependence on the enzymatic activity of the altered PMT for proliferation, and to then discover and develop selective inhibitors against that PMT. In this manner, the selective inhibition of the altered PMT can provide a basis for therapeutic intervention with a reasonable therapeutic index, owing to the unique dependence of the genetically-defined cancer on PMT activity.
Early reports of small-molecule inhibitors of PMTs presented either non-specific SAM analogs, such as SAH (the universal product of SAM hydrolysis) and sinefungin (Figure 3), or indirect inhibitors, such as 3-deazaneplanocin, which does not inhibit PMTs per se, but rather inhibits the enzyme SAH hydrolase and thereby increases cellular levels of SAH, resulting in non-selective inhibition of PMT (and other SAM-utilizing enzymes) activity.48
Over the past 5 years a number of potent, selective inhibitors of PMTs have been published; a representative sampling of these inhibitors is illustrated in Figure 3, where we also provide the reported affinity of the compound for its primary target PMT. These compounds generally bind their enzyme targets in one of two ways: either by competing with SAM for its binding pocket, or by competing with the methyl-accepting amino-acid binding pocket.
Compounds that are competitive with SAM include the aminonucleoside analogs reported by Mori et al.,49 Yao et al.50 and Daigle et al.18 These compounds were designed as SAM/SAH mimetics and display a range of target affinity and selectivity. The compound EPZ004777, for example, is a 300-pM, SAM-competitive inhibitor of DOT1L that demonstrates>1200-fold selectivity for this enzyme over all other tested PMTs.18 The high affinity of EPZ004777 and related analogs derives from a conformational adaptation mechanism in which the enzyme changes conformation to close down the ligand binding pocket around the inhibitor.51 This mechanism drives compound affinity largely by reducing the rate of dissociation for the enzyme-inhibitor binary complex, thus resulting in very long drug-target residence time.52, 53
Consistent with the idea that the chromosomal translocation confers a unique dependence on DOT1L enzymatic activity in MLL-rearranged leukemia, EPZ004777 was found to selectively kill MLL-rearranged leukemia cells in vitro, with little antiproliferative effect on non-rearranged cells. EPZ004777 causes a decrease in celluar histone H3K79 methylation followed by a decrease in MLL fusion target gene expression and induction of apoptosis in MLL-rearranged leukemia cell lines. The induction of apoptosis is preceded by accumulation of cells in the G1 phase of the cell cycle and gene expression changes consistent with hematopoietic differentiation.18 Additionally, EPZ004777 is the first PMT inhibitor to demonstrate anticancer effects and survival benefit in an animal model of MLL-rearranged leukemia18 (vide infra).
Small-molecule inhibition of DOT1L has therapeutic potential beyond oncology. Recently, Onder et al.54 found that inhibition of DOT1L activity results in acceleration of induced pluripotent stem cell generation. Importantly they demonstrate that DOT1L inhibition causes a decrease in H3K79 methylation in genes that are fated to be repressed in the pluripotent state.
The aminonucleoside inhibitors of DOT1L and SET7/9 bear a clear structural resemblance to SAM and SAH. It is therefore not surprising that these compounds act in a competitive manner with respect to SAM. In contrast, other SAM-competitive inhibitors of PMTs have been recently reported that do not retain any chemical resemblance to the nucleoside substrate. The indazole EPZ005687, for example, is a potent and selective SAM-competitive inhibitor of EZH2-containing PRC2 that has been shown to inhibit wild-type and NHL-associated mutant EZH2 with nanomolar affinity.55, 56 Although this compound inhibits both wild-type and mutant EZH2, one would expect the mutant-bearing NHL cells would have a unique dependence on EZH2 activity for proliferation, such that compounds like EPZ005687 would demonstrate selective killing of these cells. Indeed, EPZ005687 has been shown to selectively kill NHL cells that are heterozygous for EZH2 mutations at either Y641 or A677G, with limited impact on the growth of homozygous wild-type EZH2-containing NHL cells.55 Compounds of similar structure, potency and EZH2 selectivity have also been reported in a set of patent applications from the group at GlaxoSmithKline57, 58, 59, 60 (a representative example is shown in Figure 3).
A large number of PMT inhibitors act by binding not to the SAM pocket but instead to the protein-substrate binding site and engaging recognition elements within the amino-acid channel. Examples of compounds with this binding modality have been identified for both PKMT and PRMT targets. For example, the compound BIX-01294 was among the first PMT inhibitors to be reported and is a potent and selective inhibitor of the SET-domain PKMT EHMT2 (also known as G9a).61 Structural analogs of this compound were subsequently designed as dual EHMT1 and EHMT2 inhibitors with much greater target affinity and cell permeability. The compound UNC-0638, for example, was shown to reduce levels of H3K9 methylation in MDA-MB231 cells with an IC50 of 81 nM.62 Likewise, the compound AZ505 has been shown crystallographically to bind to the SET-domain PKMT SMYD2 within the lysine binding channel.63 Similarly, the structurally related compounds from the Methylgene and Bristol-Myers Squibb groups are nanomolar inhibitors of the PRMT CARM1 (also known as PRMT4) that have been shown crystallographically to bind within the arginine channel of this enzyme.64, 65, 66
A final modality of enzyme interaction is exemplified by the PRMT1 inhibitor Compound 5, which is presumed to act as a bisubstrate inhibitor, engaging recognition elements both within the SAM and within the arginine binding pockets. This compound is a relatively modest inhibitor of PRMT1, but demonstrates cellular activity in reducing methylation in HepG2 cells.67 Hence, this may be an interesting starting point for further compound optimization.
In vivo anticancer activity of PMT inhibitors
A critical step towards the eventual clinical use of PMT inhibitors is the demonstration of antitumor effects in appropriate animal models of cancer. In this regard, the field of PMT inhibitors is still quite early, with only one published report of in vivo activity for any compound.
In 2011, Daigle et al.18 reported the first clear evidence that inhibition of a specific PMT could lead to selective cancer cell killing and resultant in vivo efficacy in an animal model of cancer. These workers reported studies of EPZ004777, a potent (Ki 300 pM) and selective (⩾1200-fold over other PMTs) DOT1L inhibitor. The compound selectively blocked global H3K79 methylation in leukemia cells regardless of whether or not the cells harbored MLL gene translocations. However, despite the ability to affect intracellular H3K79 methylation in all cell types, the compound was selectively cytotoxic for leukemia cells containing the MLL gene translocations, with minimal effects on non-translocated cells. Daigle et al.18 went on to test the effects of EPZ004777 in a highly aggressive, disseminated mouse model of mixed lineage leukemia. The compound was administered by continuous administration at three concentrations of 50, 100 and 150 mg/ml for 14 days, and survival was monitored over the course of 30 days. At all doses administered, statistically significant survival advantages over the vehicle-dosed animals were achieved; the extent of survival increased with increasing dose of the compound. These data represent the first in vivo evidence that selective PMT inhibition can lead to antitumor efficacy.
Jo et al.68 showed that DOT1L has a crucial role in hematopoiesis using a postnatal conditional knockout technique. Importantly, overt toxicity in the hematopoietic compartment was not detected for the first 7–8 weeks after Dot1L excision. The authors thus suggest that this potential toxicity may be managed using noncontinuous dosing paradigms and transfusions. Whether or not reversible inhibition of DOT1L by a small-molecule drug will result in any hematopoietic liability remains to be determined.
Recently, the GlaxoSmithKline and Epizyme groups have reported at scientific meetings that some of their selective EZH2 inhibitors have demonstrated antitumor effects in subcutaneous xenograft mouse models. No peer-reviewed publications of this work have yet appeared, but these preliminary verbal reports are of significant interest to this emerging field of drug discovery and development.
Future directions
The interest in PMT inhibitors as potential cancer therapeutics has grown considerable over the past 4−5 years, to the point that today many academic groups and most pharmaceutical companies have efforts ongoing against this target class. On the basis of recent presentations at scientific meetings it seems very likely that selective PMT inhibitors will enter human clinical trials within the next year or two. Specifically, inhibitors of DOT1L for MLL-rearranged leukemia and EZH2 for mutant-bearing NHL appear to be the most advanced programs among industrial groups that have been presented publically. Thus, the near future is likely to see the clinical testing of PMT inhibitors for human cancer indications. This is an exciting development for the field and the demonstration of clinical response to these inhibitors will be a key milestone to ensure continued enthusiasm for these novel therapeutic targets. Although it is still early in the development of PMT inhibitors for clinical application, a common concern of all targeted cancer drugs is the emergence of drug resistance. Resistance to therapy may arise through several potential mechanisms, including direct effect on the targeted methyltransferase, activation of compensatory pathways (for example, by increased expression of a redundant methyltransferase), altered metabolism or efflux of the inhibitor. Whether or not such resistance mechanisms will constitute a major impediment to the clinical use of PMT inhibitors remains to be determined.
The development of potent, selective inhibitors of DOT1L and EZH2 reflect merely the tip of the proverbial iceberg for PMT targets that are genetically altered in human cancers. Beyond DOT1L, EZH2 and WHSC1, surveys of public cancer genome databases and published literature suggest that many other PMTs are genetically altered in human cancers through translocation, gene amplification and mutation. Likewise, there have been several reports of loss-of-function mutations or deletions in protein demethylases in cancers that may, in an indirect manner, confer sensitivity to inhibition of the corresponding PMT for specific amino-acid methylation/demethylation events. For example, loss-of-function mutations in the H3K27 demethylase UTX, with an accompanying hyper-trimethylation of H3K27, have been reported for subsets of myeloma, esophageal and renal cancers69 and medulloblastoma.70, 71 Hence, inhibition of PRC2-catalyzed methylation of the H3K27 site may be of value for these indications. Additionally, the H3K4 demethylase KDM1A has recently been shown to have an essential role in maintaining MLL fusion protein target gene expression in MLL-rearranged leukemia.72 Inhibition of this demethylase with small-molecule inhibitors induced cell death. Whether all of these alterations of epigenetic enzymes will prove to be tumorigenic drivers remains to be determined. Nevertheless, even if only a subset of these enzymatic activities are required for the tumorigenic phenotype, the PMTs would still constitute a large pool of targets for drug discovery.
The concept of personalized cancer therapeutics rests on the ability to (1) define unique driver alterations in a specific cancer, (2) discover and develop potent and selective drugs against that uniquely altered target and (3) simultaneously develop screening modalities with which to identify patients harboring the alteration, who are most likely to benefit from treatment with the targeted drug (that is, companion diagnostics). Recent examples of the success of this approach with selective kinase inhibitors portend a greater future reliance on personalized cancer therapeutics for clinical oncology and perhaps in other areas of medicine as well. As described in this brief review, the PMTs seem ideally suited as targets for personalized cancer therapeutics, both because of the demonstrated reliance of specific cancers on the enzymatic activity of these enzymes and because of the general susceptibility of this target class to inhibition by small, drug-like compounds. Further development of selective inhibitors against these enzymes thus offers considerable potential for augmenting the current armamentarium of personalized cancer therapies.
References
Kouzarides T . Chromatin modifications and their function. Cell 2007; 128: 693–705.
Strahl BD, Allis CD . The language of covalent histone modifications. Nature 2000; 403: 41–45.
Allis CD, Berger S, Cote G, Dent SYR, Jenuwien T, Kouzarides T et al. New nomenclature for chromatin-modifying enzymes. Cell 2007; 131: 633–636.
Copeland RA, Olhava EJ, Scott MP . Targeting epigenetic enzymes for drug discovery. Curr Opin Chem Biol 2010; 14: 505–510.
Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M . Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov 2012; 11: 384–400.
Copeland RA . Protein methyltransferase inhibitors as personalized cancer therapeutics. Drug Discov Today Ther Strategies 2011, (doi:10.1016/j.ddstr.2011.08.001).
Copeland RA, Solomon ME, Richon VM . Protein methyltransferases as a target class for drug discovery. Nat Rev Drug Discov 2009; 8: 724–732.
Richon VM, Johnston D, Sneeringer CJ, Jin L, Majer CR, Elliston K et al. Chemogenetic analysis of human protein methyltransferases. Chem Biol Drug Des 2011; 78: 199–210.
Barski A, Cuddapah S, Cui K, Roh T-Y, Schones DE, Wang Z et al. High-resolution profiling of histone methylations in the human genome. Cell 2007; 129: 823–837.
Webb KJ, Lipson RS, Al-Hadid Q, Whitelegge JP, Clarke SG . Identification of protein N-terminal methyltransferases in yeast and humans. Biochemistry 2010; 49: 5225–5235.
Copeland RA . Enzymes: A Practical Introduction to Structure, Mechanism and Data Analysis 2nd edn. New York, Wiley, 2000.
Cheng X, Collins RE, Zhang X . Structural and sequence motifs of protein (histone) methylation enzymes. Annu Rev Biophys Biomol Struct 2005; 34: 267–294.
Kelly TK, De Carvalho DD, Jones PA . Epigenetic modifications as therapeutic targets. Nat Biotechnol 2010; 28: 1069–1078.
Luo M . Current chemical biology approaches to interrogate protein methyltransferases. ACS Chem Biol 2012; 7: 443–463.
Feng Q, Wang H, Ng HH, Erdjument-Bromage H, Tempst P, Struhl K et al. Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr Biol 2002; 12: 1052–1058.
Cao R, Zhang Y . The functions of E(Z)/EZH2-mediated methylation of lysine 27 in histone H3. Curr Opin Genet Dev 2004; 14: 155–164.
Lee YH, Stallcup MR . Minireview: protein arginine methylation of nonhistone proteins in transcriptional regulation. Mol Endocrinol 2009; 23: 425–433.
Daigle SR, Olhava EJ, Therkelsen CA, Majer CR, Sneeringer CJ, Song J et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule dot1L inhibitor. Cancer Cell 2011; 20: 53–65.
Bernt KM, Zhu N, Sinha AU, Vempati S, Faber J, Krivtsov AV et al. MLL-rearranged leukemia is dependent on aberrant h3k79 methylation by dot1L. Cancer Cell 2011; 20: 66–78.
Hess JL, Hug BA . Fusion-protein truncation provides new insights into leukemogenesis. Proc Natl Acad Sci USA 2004; 101: 16985–16986.
Krivtsov AV, Armstrong SA . MLL translocations, histone modifications and leukaemia stem-cell development. Nat Rev Cancer 2007; 7: 823–833.
Slany RK . The molecular biology of mixed lineage leukemia. Haematologica 2009; 94: 984–993.
Ayton PM, Chen EH, Cleary ML . Binding to nonmethylated CpG DNA is essential for target recognition, transactivation, and myeloid transformation by an MLL oncoprotein. Mol Cell Biol 2004; 24: 10470–10478.
Slany RK, Lavau C, Cleary ML . The oncogenic capacity of HRX-ENL requires the transcriptional transactivation activity of ENL and the DNA binding motifs of HRX. Mol Cell Biol 1998; 18: 122–129.
Zeleznik-Le NJ, Harden AM, Rowley JD . 11q23 translocations split the ‘AT-hook’ cruciform DNA-binding region and the transcriptional repression domain from the activation domain of the mixed-lineage leukemia (MLL) gene. Proc Natl Acad Sci USA 1994; 91: 10610–10614.
Muntean AG, Tan J, Sitwala K, Huang Y, Bronstein J, Connelly JA et al. The PAF complex synergizes with MLL fusion proteins at HOX loci to promote leukemogenesis. Cancer Cell 2010; 17: 609–621.
He N, Chan CK, Sobhian B, Chou S, Xue Y, Liu M et al. Human polymerase-associated factor complex (PAFc) connects the super elongation complex (SEC) to RNA polymerase II on chromatin. Proc Natl Acad Sci USA 2011; 108: E636–E645.
Lin C, Smith ER, Takahashi H, Lai KC, Martin-Brown S, Florens L et al. AFF4, a component of the ELL/P-TEFb elongation complex and a shared subunit of MLL chimeras, can link transcription elongation to leukemia. Mol Cell 2010; 37: 429–437.
Benedikt A, Baltruschat S, Scholz B, Bursen A, Arrey TN, Meyer B et al. The leukemogenic AF4-MLL fusion protein causes P-TEFb kinase activation and altered epigenetic signatures. Leukemia 2011; 25: 135–144.
Thiel AT, Blessington P, Zou T, Feather D, Wu X, Yan J et al. MLL-AF9-induced leukemogenesis requires coexpression of the wild-type mll allele. Cancer Cell 2010; 17: 148–159.
Okada Y, Feng Q, Lin Y, Jiang Q, Li Y, Coffield VM et al. hDOT1L links histone methylation to leukemogenesis. Cell 2005; 121: 167–178.
Krivtsov AV, Feng Z, Lemieux ME, Faber J, Vempati S, Sinha AU et al. H3K79 methylation profiles define murine and human mll-af4 leukemias. Cancer Cell 2008; 14: 355–368.
Mahmoudi T, Boj SF, Hatzis P, Li VSW, Taouatas N, Vries RGJ et al. The leukemia-associated mllt10/af10-Dot1L are TCF4/β-catenin coactivators essential for intestinal homeostasis. PLoS Biol 2010; 8: e1000539.
Marango J, Shimoyama M, Nishio H, Meyer JA, Min DJ, Sirulnik A et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood 2008; 111: 3145–3154.
Li B, Jackson J, Simon MD, Fleharty B, Gogol M, Seidel C et al. Histone H3 lysine 36 dimethylation (H3K36me2) is sufficient to recruit the Rpd3s histone deacetylase complex and to repress spurious transcription. J Biol Chem 2009; 284: 7970–7976.
Chesi M, Nardini E, Lim RS, Smith KD, Kuehl WM, Bergsagel PL . The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 1998; 92: 3025–3034.
Keats JJ, Maxwell CA, Taylor BJ, Hendzel MJ, Chesi M, Bergsagel PL et al. Overexpression of transcripts originating from the MMSET locus characterizes all t(4;14)(p16;q32)-positive multiple myeloma patients. Blood 2005; 105: 4060–4069.
Keats JJ, Reiman T, Maxwell CA, Taylor BJ, Larratt LM, Mant MJ et al. In multiple myeloma, t(4;14)(p16;q32) is an adverse prognostic factor irrespective of FGFR3 expression. Blood 2003; 101: 1520–1529.
Martinez-Garcia E, Popovic R, Min D-J, Sweet SMM, Thomas PM, Zamdborg L et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood 2011; 117: 211–220.
Simon JA, Lange CA . Roles of the EZH2 histone methyltransferase in cancer epigenetics. Mutat Res 2008; 647: 21–29.
Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet 2010; 42: 181–185.
Sneeringer C, Scott MP, Kuntz K, Knutson SK, Pollock RM, Richon V et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. ProceNatl Acad Sci USA 2010; 107: 20980–20985.
McCabe MT, Graves AP, Ganji G, Diaz E, Halsey WS, Jiang Y et al. Mutation of A677 in histone methyltransferase EZH2 in human B-cell lymphoma promotes hypertrimethylation of histone H3 on lysine 27 (H3K27). Proc Natl Acad Sci USA 2012; 109: 2989–2994.
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet 2010; 42: 722–726.
Nikoloski G, Langemeijer SM, Kuiper RP, Knops R, Massop M, Tonnissen ER et al. Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet 2010; 42: 665–667.
Ntziachristos P, Tsirigos A, Vlierberghe PV, Nedjic J, Trimarchi T, Flaherty MS et al. Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat Med 2012; 18: 298–303.
Mochizuki-Kashio M, Mishima Y, Miyagi S, Negishi M, Saraya A, Konuma T et al. Dependency on the polycomb gene Ezh2 distinguishes fetal from adult hematopoietic stem cells. Blood 2011; 118: 6553–6561.
Chiang PK . Biological effects of inhibitors of S-adenosylhomocysteine hydrolase. Pharmacol Ther 1998; 77: 115–134.
Mori S, Iwase K, Iwanami N, Tanaka Y, Kagechika H, Hirano T . Development of novel bisubstrate-type inhibitors of histone methyltransferase SET7/9. Bioorg Med Chem 2010; 18: 8158–8166.
Yao Y, Chen P, Diao J, Cheng G, Deng L, Anglin JL et al. Selective inhibitors of histone methyltransferase DOT1L: design, synthesis, and crystallographic studies. J Am Chem Soc 2011; 133: 16746–16749.
Basavapathruni A, Jin L, Daigle SR, Majer CR, Therkelsen CA, Wigle TJ et al. Conformational adaption drives potent, selective and durable inhibition of the human protein methyltransferase DOT1L. Chem Biol Drug Des 2012; 80: 971–980.
Copeland RA, Pompliano DL, Meek TD . Drug-target residence time and its implications for lead optimization. Nat Rev Drug Discov 2006; 5: 730–739.
Copeland RA . Conformational adaptation in drug-target interactions and residence time. Future Med Chem 2011; 3: 1491–1501.
Onder TT, Kara N, Cherry A, Sinha AU, Zhu N, Bernt KM et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 2012; 483: 598–602.
Knutson SK, Wigle TJ, Warholic NM, Sneeringer CJ, Allain CJ, Klaus CR et al. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nat Chem Biol 2012; 8: 890–896.
Copeland RA, Richon VM, Scott MD, Sneeringer CJ, Kuntz KW, Knutson SK et alInhibitors of human EZH2, and methods of use thereof. Patent Application WO2012034132, 2012.
Brackley J, Burgess JL, Grant S, Johnson N, Knight SD, LaFrance L et alIndoles WO2011140324, 2011.
Duquenne C, Johnson N, Knight SD, LaFrance L, Miller WH, Newlander K et alIndoles WO2011140325, 2011.
Burgess JL, Johnson N, Knight SD, LaFrance L, Miller WH, Newlander K et alAzaindazoles WO2012005805, 2012.
Knight SD, Miller WH, Newlander KA, Verma SK Indoles WO2012075080, 2012.
Kubicek S, O'Sullivan RJ, August EM, Hickey ER, Zhang Q, Teodoro ML et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell 2007; 25: 473–481.
Vedadi M, Barsyte-Lovejoy D, Liu F, Rival-Gervier S, Allali-Hassani A, Labrie V et al. A chemical probe selectively inhibits G9a and GLP methyltransferase activity in cells. Nat Chem Biol 2011; 7: 566–574.
Ferguson ALN, Howard T, Pollard H, Green I, Grande C, Cheung T et al. structural basis of substrate methylation and inhibition of SMYD2. Structure 2011.
Purandare AV, Chen Z, Huynh T, Pang S, Geng J, Vaccaro W et al. Pyrazole inhibitors of coactivator associated arginine methyltransferase 1 (CARM1). Bioorg Med Chem Lett 2008; 18: 4438–4441.
Allan M, Manku S, Therrien E, Nguyen N, Styhler S, Robert MF et al. N-benzyl-1-heteroaryl-3-(trifluoromethyl)-1H-pyrazole-5-carboxamides as inhibitors of co-activator associated arginine methyltransferase 1 (CARM1). Bioorg Med Chem Lett 2009; 19: 1218–1223.
Huynh T, Chen Z, Pang S, Geng J, Bandiera T, Bindi S et al. Optimization of pyrazole inhibitors of coactivator associated arginine methyltransferase 1 (CARM1). Bioorg Med Chem Lett 2009; 19: 2924–2927.
Spannhoff A, Machmur R, Heinke R, Trojer P, Bauer I, Brosch G et al. A novel arginine methyltransferase inhibitor with cellular activity. Bioorg Med Chem Lett 2007; 17: 4150–4153.
Jo SY, Granowicz EM, Maillard I, Thomas D, Hess JL . Requirement for Dot1L in murine postnatal hematopoiesis and leukemogenesis by MLL translocation. Blood 2011; 117: 4759–4768.
van Haaften G, Dalgliesh GL, Davies H, Chen L, Bignell G, Greenman C et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat Genet 2009; 41: 521–523.
Parsons DW, Li M, Zhang X, Jones S, Leary RJ, Lin JC et al. The genetic landscape of the childhood cancer medulloblastoma. Science 2011; 331: 435–439.
Robinson G, Parker M, Kranenburg TA, Lu C, Chen X, Ding L et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 2012; 488: 43–48.
Harris WJ, Huang X, Lynch JT, Spencer GJ, Hitchin JR, Li Y et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 2012; 21: 473–487.
Liu F, Chen X, Allali-Hassani A, Quinn AM, Wasney GA, Dong A et al. Discovery of a 2,4-diamino-7-aminoalkoxyquinazoline as a potent and selective inhibitor of histone lysine methyltransferase G9a. J Med Chem 2009; 52: 7950–7953.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors of the paper are employees of Epizyme.
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Copeland, R., Moyer, M. & Richon, V. Targeting genetic alterations in protein methyltransferases for personalized cancer therapeutics. Oncogene 32, 939–946 (2013). https://doi.org/10.1038/onc.2012.552
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.552
Keywords
This article is cited by
-
Charakterisierung der Substratspezifität von Protein-Methyltransferasen
BIOspektrum (2023)
-
Protein kinase A-mediated phosphorylation regulates STAT3 activation and oncogenic EZH2 activity
Oncogene (2018)
-
Detection of PRMT1 inhibitors with stopped flow fluorescence
Signal Transduction and Targeted Therapy (2018)
-
The emerging role of lysine methyltransferase SETD8 in human diseases
Clinical Epigenetics (2016)
-
Structural insights into binding of small molecule inhibitors to Enhancer of Zeste Homolog 2
Journal of Computer-Aided Molecular Design (2014)