Abstract
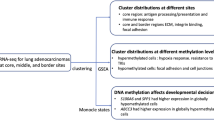
Aberrant DNA hypermethylation of tumor suppressor genes is thought to be an early event in tumorigenesis. Many studies have reported the methylation status of individual genes with known involvement in cancer, but an unbiased assessment of the biological function of the collective of hypermethylated genes has not been conducted so far. Based on the observation that a variety of human cancers recapitulate developmental gene expression patterns (that is activate genes normally expressed in early development and suppress late developmental genes), we hypothesized that the silencing of differentiation-associated genes in cancer could be attributed in part to DNA hypermethylation. To this end, we investigated the developmental expression patterns of genes with hypermethylated CpG islands in primary human lung carcinomas and lung cancer cell lines. We found that DNA hypermethylation primarily affects genes that are expressed in late stages of murine lung development. Gene ontology characterization of these genes shows that they are almost exclusively involved in morphogenetic differentiation processes. Our results indicate that DNA hypermethylation in cancer functions as a selective silencing mechanism of genes that are required for the maintenance of a differentiated state. The process of cellular de-differentiation that is evident on both the microscopic and transcriptional level in cancer might at least partly be mediated by these epigenetic events. Our observations provide a mechanistic explanation for induction of differentiation upon treatment with DNA methyltransferase inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Antequera F, Boyes J, Bird A . (1990). High levels of de novo methylation and altered chromatin structure at CpG islands in cell lines. Cell 62: 503–514.
Armstrong L, Lako M, Dean W, Stojkovic M . (2006). Epigenetic modification is central to genome reprogramming in somatic cell nuclear transfer. Stem Cells 24: 805–814.
Banerjee S, Bacanamwo M . (2010). DNA methyltransferase inhibition induces mouse embryonic stem cell differentiation into endothelial cells. Exp Cell Res 316: 172–180.
Baylin SB, Herman JG . (2000). DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet 16: 168–174.
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP . (1998). Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res 72: 141–196.
Belinsky SA . (2004). Gene-promoter hypermethylation as a biomarker in lung cancer. Nat Rev Cancer 4: 707–717.
Belinsky SA, Nikula KJ, Palmisano WA, Michels R, Saccomanno G, Gabrielson E et al. (1998). Aberrant methylation of p16INK4a is an early event in lung cancer and a potential biomarker for early diagnosis. Proc Natl Acad Sci U S A 95: 11891–11896.
Borczuk AC, Gorenstein L, Walter KL, Assaad AA, Wang L, Powell CA . (2003). Non-small-cell lung cancer molecular signatures recapitulate lung developmental pathways. Am J Pathol 163: 1949–1960.
Costello JF, Frühwald MC, Smiraglia DJ, Rush LJ, Robertson GP, Gao X et al. (2000). Aberrant CpG-island methylation has non-random and tumor-type-specific patterns. Nat Genet 24: 132–138.
Delgado S, Gómez M, Bird A, Antequera F . (1998). Initiation of DNA replication at CpG islands in mammalian chromosomes. EMBO J 17: 2426–2435.
Esteller M . (2002). CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene 21: 5427–5440.
Esteller M . (2005). Aberrant DNA methylation as a cancer-inducing mechanism. Annu Rev Pharmacol Toxicol 45: 629–656.
Esteller M, Corn PG, Baylin SB, Herman JG . (2001). A gene hypermethylation profile of human cancer. Cancer Res 61: 3225–3229.
Falcon S, Gentleman R . (2007). Using GOstats to test gene lists for GO term association. Bioinformatics 23: 257–258.
Feinberg AP, Vogelstein B . (1983). Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301: 89–92.
Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML et al. (2005). Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci USA 102: 10604–10609.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S et al. (2004). Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5: R80.
Gonzalez-Zulueta M, Bender CM, Yang AS, Nguyen T, Beart RW, Van Tornout JM et al. (1995). Methylation of the 5′ CpG island of the p16/CDKN2 tumor suppressor gene in normal and transformed human tissues correlates with gene silencing. Cancer Res 55: 4531–4535.
Greger V, Passarge E, Höpping W, Messmer E, Horsthemke B . (1989). Epigenetic changes may contribute to the formation and spontaneous regression of retinoblastoma. Hum Genet 83: 155–158.
Hajkova P, Erhardt S, Lane N, Haaf T, El-Maarri O, Reik W et al. (2002). Epigenetic reprogramming in mouse primordial germ cells. Mech Dev 117: 15–23.
Hayslip J, Montero A . (2006). Tumor suppressor gene methylation in follicular lymphoma: a comprehensive review. Mol Cancer 5: 44.
Heller G, Zielinski CC, Zöchbauer-Müller S . (2010). Lung cancer: from single-gene methylation to methylome profiling. Cancer Metastasis Rev 29: 95–107.
Herman JG, Merlo A, Mao L, Lapidus RG, Issa JP, Davidson NE et al. (1995). Inactivation of the CDKN2/p16/MTS1 gene is frequently associated with aberrant DNA methylation in all common human cancers. Cancer Res 55: 4525–4530.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al. (2003). Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4: 249–264.
Keshet I, Lieman-Hurwitz J, Cedar H . (1986). DNA methylation affects the formation of active chromatin. Cell 44: 535–543.
Kho AT, Zhao Q, Cai Z, Butte AJ, Kim JY, Pomeroy SL et al. (2004). Conserved mechanisms across development and tumorigenesis revealed by a mouse development perspective of human cancers. Genes Dev 18: 629–640.
Laird PW . (2003). The power and the promise of DNA methylation markers. Nat Rev Cancer 3: 253–266.
Liu H, Kho AT, Kohane IS, Sun Y . (2006). Predicting survival within the lung cancer histopathological hierarchy using a multi-scale genomic model of development. PLoS Med 3: e232.
Marchevsky AM, Tsou JA, Laird-Offringa IA . (2004). Classification of individual lung cancer cell lines based on DNA methylation markers: use of linear discriminant analysis and artificial neural networks. J Mol Diagn 6: 28–36.
McLaren A . (2000). Germ and somatic cell lineages in the developing gonad. Mol Cell Endocrinol 163: 3–9.
Meissner A, Mikkelsen TS, Gu H, Wernig M, Hanna J, Sivachenko A et al. (2008). Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454: 766–770.
Merlo A, Herman JG, Mao L, Lee DJ, Gabrielson E, Burger PC et al. (1995). 5' CpG island methylation is associated with transcriptional silencing of the tumor suppressor p16/CDKN2/MTS1 in human cancers. Nat Med 1: 686–692.
Momparler RL, Bouchard J, Samson J . (1985). Induction of differentiation and inhibition of DNA methylation in HL-60 myeloid leukemic cells by 5-AZA-2′-deoxycytidine. Leuk Res 9: 1361–1366.
Naxerova K, Bult CJ, Peaston A, Fancher K, Knowles BB, Kasif S et al. (2008). Analysis of gene expression in a developmental context emphasizes distinct biological leitmotifs in human cancers. Genome Biol 9: R108.
Nephew K . (2003). Epigenetic gene silencing in cancer initiation and progression. Cancer Lett 190: 125–133.
Nuovo GJ, Plaia TW, Belinsky SA, Baylin SB, Herman JG . (1999). In situ detection of the hypermethylation-induced inactivation of the p16 gene as an early event in oncogenesis. Proc Natl Acad Sci USA 96: 12754–12759.
Pfeifer GP, Rauch TA . (2009). DNA methylation patterns in lung carcinomas. Semin Cancer Biol 19: 181–187.
Rauch T, Wang Z, Zhang X, Zhong X, Wu X, Lau SK et al. (2007). Homeobox gene methylation in lung cancer studied by genome-wide analysis with a microarray-based methylated CpG island recovery assay. Proc Natl Acad Sci USA 104: 5527–5532.
Rauch TA, Zhong X, Wu X, Wang M, Kernstine KH, Wang Z et al. (2008). High-resolution mapping of DNA hypermethylation and hypomethylation in lung cancer. Proc Natl Acad Sci USA 105: 252–257.
Safar AM, Spencer III H, Su X, Coffey M, Cooney CA, Ratnasinghe LD et al. (2005). Methylation profiling of archived non-small cell lung cancer: a promising prognostic system. Clin Cancer Res 11: 4400–4405.
Sakai T, Toguchida J, Ohtani N, Yandell DW, Rapaport JM, Dryja TP et al. (1991). Allele-specific hypermethylation of the retinoblastoma tumor-suppressor gene. Am J Hum Genet 48: 880–888.
Sasaki H, Matsui Y . (2008). Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat Rev Genet 9: 129–140.
Shames DS, Girard L, Gao B, Sato M, Lewis CM, Shivapurkar N et al. (2006). A genome-wide screen for promoter methylation in lung cancer identifies novel methylation markers for multiple malignancies. PLoS Med 3: e486.
Taylor SM, Jones PA . (1979). Multiple new phenotypes induced in 10T1/2 and 3T3 cells treated with 5-azacytidine. Cell 17: 771–779.
Tsou JA, Hagen JA, Carpenter CL, Laird-Offringa IA . (2002). DNA methylation analysis: a powerful new tool for lung cancer diagnosis. Oncogene 21: 5450–5461.
Ushijima T . (2005). Detection and interpretation of altered methylation patterns in cancer cells. Nat Rev Cancer 5: 223–231.
Wang Y, Leung FCC . (2004). An evaluation of new criteria for CpG islands in the human genome as gene markers. Bioinformatics 20: 1170–1177.
Xu C, Police S, Rao N, Carpenter MK . (2002). Characterization and enrichment of cardiomyocytes derived from human embryonic stem cells. Circ Res 91: 501–508.
Yoon BS, Yoo SJ, Lee JE, You S, Lee HT, Yoon HS . (2006). Enhanced differentiation of human embryonic stem cells into cardiomyocytes by combining hanging drop culture and 5-azacytidine treatment. Differentiation 74: 149–159.
Acknowledgements
We would like to thank David Page, Simon Kasif and Annina Deleo for valuable discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Helman, E., Naxerova, K. & Kohane, I. DNA hypermethylation in lung cancer is targeted at differentiation-associated genes. Oncogene 31, 1181–1188 (2012). https://doi.org/10.1038/onc.2011.307
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.307
Keywords
This article is cited by
-
Differences in MWCNT- and SWCNT-induced DNA methylation alterations in association with the nuclear deposition
Particle and Fibre Toxicology (2018)
-
High-frequency aberrantly methylated targets in pancreatic adenocarcinoma identified via global DNA methylation analysis using methylCap-seq
Clinical Epigenetics (2014)
-
Genome-wide DNA methylation profiling of non-small cell lung carcinomas
Epigenetics & Chromatin (2012)
-
Reversal of Aberrant Cancer Methylome and Transcriptome upon Direct Reprogramming of Lung Cancer Cells
Scientific Reports (2012)