Abstract
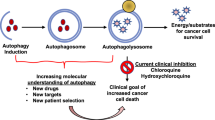
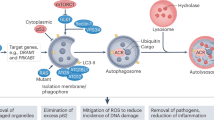
Autophagy, hallmarked by the formation of double-membrane bound organelles known as autophagosomes, is a lysosome-dependent pathway for protein degradation. The role of autophagy in carcinogenesis is context dependent. As a tumor-suppressing mechanism in early-stage carcinogenesis, autophagy inhibits inflammation and promotes genomic stability. Moreover, disruption of autophagy-related genes accelerates tumorigenesis in animals. However, autophagy may also act as a pro-survival mechanism to protect cancer cells from various forms of cellular stress. In cancer therapy, adaptive autophagy in cancer cells sustains tumor growth and survival in face of the toxicity of cancer therapy. To this end, inhibition of autophagy may sensitize cancer cells to chemotherapeutic agents and ionizing radiation. Nevertheless, in certain circumstances, autophagy mediates the therapeutic effects of some anticancer agents. Data from recent studies are beginning to unveil the apparently paradoxical nature of autophagy as a cell-fate decision machinery. Taken together, modulation of autophagy is a novel approach for enhancing the efficacy of existing cancer therapy, but its Janus-faced nature may complicate the clinical development of autophagy modulators as anticancer therapeutics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Abida WM, Gu W . (2008). p53-Dependent and p53-independent activation of autophagy by ARF. Cancer Res 68: 352–357.
Ahn CH, Jeong EG, Lee JW, Kim MS, Kim SH, Kim SS et al. (2007). Expression of beclin-1, an autophagy-related protein, in gastric and colorectal cancers. APMIS 115: 1344–1349.
Akar U, Chaves-Reyez A, Barria M, Tari A, Sanguino A, Kondo Y et al. (2008). Silencing of Bcl-2 expression by small interfering RNA induces autophagic cell death in MCF-7 breast cancer cells. Autophagy 4: 669–679.
Alexander A, Cai SL, Kim J, Nanez A, Sahin M, MacLean KH et al. (2010). ATM signals to TSC2 in the cytoplasm to regulate mTORC1 in response to ROS. Proc Natl Acad Sci USA 107: 4153–4158.
Altman BJ, Jacobs SR, Mason EF, Michalek RD, Macintyre AN, Coloff JL et al. (2011). Autophagy is essential to suppress cell stress and to allow BCR-Abl-mediated leukemogenesis. Oncogene 30: 1855–1867.
Altman BJ, Wofford JA, Zhao Y, Coloff JL, Ferguson EC, Wieman HL et al. (2009). Autophagy provides nutrients but can lead to Chop-dependent induction of Bim to sensitize growth factor-deprived cells to apoptosis. Mol Biol Cell 20: 1180–1191.
Aoki H, Takada Y, Kondo S, Sawaya R, Aggarwal BB, Kondo Y . (2007). Evidence that curcumin suppresses the growth of malignant gliomas in vitro and in vivo through induction of autophagy: role of Akt and extracellular signal-regulated kinase signaling pathways. Mol Pharmacol 72: 29–39.
Arico S, Petiot A, Bauvy C, Dubbelhuis PF, Meijer AJ, Codogno P et al. (2001). The tumor suppressor PTEN positively regulates macroautophagy by inhibiting the phosphatidylinositol 3-kinase/protein kinase B pathway. J Biol Chem 276: 35243–35246.
Barre B, Perkins ND . (2010). The Skp2 promoter integrates signaling through the NF-kappaB, p53, and Akt/GSK3beta pathways to regulate autophagy and apoptosis. Mol Cell 38: 524–538.
Bhoopathi P, Chetty C, Gujrati M, Dinh DH, Rao JS, Lakka S . (2010). Cathepsin B facilitates autophagy-mediated apoptosis in SPARC overexpressed primitive neuroectodermal tumor cells. Cell Death Differ 17: 1529–1539.
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A et al. (2005). p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171: 603–614.
Bren GD, Solan NJ, Miyoshi H, Pennington KN, Pobst LJ, Paya CV . (2001). Transcription of the RelB gene is regulated by NF-kappaB. Oncogene 20: 7722–7733.
Cao C, Subhawong T, Albert JM, Kim KW, Geng L, Sekhar KR et al. (2006). Inhibition of mammalian target of rapamycin or apoptotic pathway induces autophagy and radiosensitizes PTEN null prostate cancer cells. Cancer Res 66: 10040–10047.
Castino R, Thepparit C, Bellio N, Murphy D, Isidoro C . (2008). Akt induces apoptosis in neuroblastoma cells expressing a C98X vasopressin mutant following autophagy suppression. J Neuroendocrinol 20: 1165–1175.
Chang CP, Yang MC, Liu HS, Lin YS, Lei HY . (2007). Concanavalin A induces autophagy in hepatoma cells and has a therapeutic effect in a murine in situ hepatoma model. Hepatology 45: 286–296.
Chang YT, Tseng HC, Huang CC, Chen YP, Chiang HC, Chou FP . (2011). Relative down-regulation of apoptosis and autophagy genes in colorectal cancer. Eur J Clin Invest 41: 84–92.
Chen F, Castranova V . (2007). Nuclear factor-kappaB, an unappreciated tumor suppressor. Cancer Res 67: 11093–11098.
Chen Q, Xie W, Kuhn DJ, Voorhees PM, Lopez-Girona A, Mendy D et al. (2008). Targeting the p27 E3 ligase SCF(Skp2) results in p27- and Skp2-mediated cell-cycle arrest and activation of autophagy. Blood 111: 4690–4699.
Cliby W, Ritland S, Hartmann L, Dodson M, Halling KC, Keeney G et al. (1993). Human epithelial ovarian cancer allelotype. Cancer Res 53: 2393–2398.
Copetti T, Bertoli C, Dalla E, Demarchi F, Schneider C . (2009). p65/RelA modulates BECN1 transcription and autophagy. Mol Cell Biol 29: 2594–2608.
Crighton D, Wilkinson S, O'Prey J, Syed N, Smith P, Harrison PR et al. (2006). DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell 126: 121–134.
Criollo A, Senovilla L, Authier H, Maiuri MC, Morselli E, Vitale I et al. (2010). The IKK complex contributes to the induction of autophagy. EMBO J 29: 619–631.
de Haan CA, Molinari M, Reggiori F . (2010). Autophagy-independent LC3 function in vesicular traffic. Autophagy 6: 994–996.
Degenhardt K, Mathew R, Beaudoin B, Bray K, Anderson D, Chen G et al. (2006). Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 10: 51–64.
Ding WX, Ni HM, Gao W, Hou YF, Melan MA, Chen X et al. (2007). Differential effects of endoplasmic reticulum stress-induced autophagy on cell survival. J Biol Chem 282: 4702–4710.
Ding Y, Kim JK, Kim SI, Na HJ, Jun SY, Lee SJ et al. (2010). TGF-beta1 protects against mesangial cell apoptosis via induction of autophagy. J Biol Chem 285: 37909–37919.
Ding ZB, Shi YH, Zhou J, Qiu SJ, Xu Y, Dai Z et al. (2008). Association of autophagy defect with a malignant phenotype and poor prognosis of hepatocellular carcinoma. Cancer Res 68: 9167–9175.
Djavaheri-Mergny M, Amelotti M, Mathieu J, Besancon F, Bauvy C, Souquere S et al. (2006). NF-kappaB activation represses tumor necrosis factor-alpha-induced autophagy. J Biol Chem 281: 30373–30382.
Donohue E, Tovey A, Vogl AW, Arns S, Sternberg E, Young RN et al. (2011). Inhibition of autophagosome formation by the benzoporphyrin derivative verteporfin. J Biol Chem 286: 7290–7300.
Eby KG, Rosenbluth JM, Mays DJ, Marshall CB, Barton CE, Sinha S et al. (2010). ISG20L1 is a p53 family target gene that modulates genotoxic stress-induced autophagy. Mol Cancer 9: 95.
Eccles DM, Russell SE, Haites NE, Atkinson R, Bell DW, Gruber L et al. (1992). Early loss of heterozygosity on 17q in ovarian cancer. The Abe Ovarian Cancer Genetics Group. Oncogene 7: 2069–2072.
Edinger AL, Cinalli RM, Thompson CB . (2003). Rab7 prevents growth factor-independent survival by inhibiting cell-autonomous nutrient transporter expression. Dev Cell 5: 571–582.
Egan DF, Shackelford DB, Mihaylova MM, Gelino S, Kohnz RA, Mair W et al. (2011). Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331: 456–461.
Elgendy M, Sheridan C, Brumatti G, Martin SJ . (2011). Oncogenic ras-induced expression of noxa and beclin-1 promotes autophagic cell death and limits clonogenic survival. Mol Cell 42: 23–35.
Fan QW, Cheng C, Hackett C, Feldman M, Houseman BT, Nicolaides T et al. (2010). Akt and autophagy cooperate to promote survival of drug-resistant glioma. Sci Signal 3: ra81.
Feng Z, Levine AJ . (2010). The regulation of energy metabolism and the IGF-1/mTOR pathways by the p53 protein. Trends Cell Biol 20: 427–434.
Fu L, Kim YA, Wang X, Wu X, Yue P, Lonial S et al. (2009). Perifosine inhibits mammalian target of rapamycin signaling through facilitating degradation of major components in the mTOR axis and induces autophagy. Cancer Res 69: 8967–8976.
Fujii S, Mitsunaga S, Yamazaki M, Hasebe T, Ishii G, Kojima M et al. (2008). Autophagy is activated in pancreatic cancer cells and correlates with poor patient outcome. Cancer Sci 99: 1813–1819.
Fujiwara K, Daido S, Yamamoto A, Kobayashi R, Yokoyama T, Aoki H et al. (2008). Pivotal role of the cyclin-dependent kinase inhibitor p21WAF1/CIP1 in apoptosis and autophagy. J Biol Chem 283: 388–397.
Fung C, Lock R, Gao S, Salas E, Debnath J . (2008). Induction of autophagy during extracellular matrix detachment promotes cell survival. Mol Biol Cell 19: 797–806.
Futreal PA, Soderkvist P, Marks JR, Iglehart JD, Cochran C, Barrett JC et al. (1992). Detection of frequent allelic loss on proximal chromosome 17q in sporadic breast carcinoma using microsatellite length polymorphisms. Cancer Res 52: 2624–2627.
Gajewska M, Gajkowska B, Motyl T . (2005). Apoptosis and autophagy induced by TGF-B1 in bovine mammary epithelial BME-UV1 cells. J Physiol Pharmacol 56 (Suppl 3): 143–157.
Ganley IG, Lam du H, Wang J, Ding X, Chen S, Jiang X . (2009). ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J Biol Chem 284: 12297–12305.
Gao P, Bauvy C, Souquere S, Tonelli G, Liu L, Zhu Y et al. (2010). The Bcl-2 homology domain 3 mimetic gossypol induces both Beclin 1-dependent and Beclin 1-independent cytoprotective autophagy in cancer cells. J Biol Chem 285: 25570–25781.
Gao W, Shen Z, Shang L, Wang X . (2011). Upregulation of human autophagy-initiation kinase ULK1 by tumor suppressor p53 contributes to DNA-damage-induced cell death. Cell Death Differ (doi:10.1038/cdd.2011.33).
Gao X, Zacharek A, Salkowski A, Grignon DJ, Sakr W, Porter AT et al. (1995). Loss of heterozygosity of the BRCA1 and other loci on chromosome 17q in human prostate cancer. Cancer Res 55: 1002–1005.
Ghavami S, Asoodeh A, Klonisch T, Halayko AJ, Kadkhoda K, Kroczak TJ et al. (2008). Brevinin-2R(1) semi-selectively kills cancer cells by a distinct mechanism, which involves the lysosomal-mitochondrial death pathway. J Cell Mol Med 12: 1005–1022.
Gu Y, Wang C, Cohen A . (2004). Effect of IGF-1 on the balance between autophagy of dysfunctional mitochondria and apoptosis. FEBS Lett 577: 357–360.
Guo JY, Chen HY, Mathew R, Fan J, Strohecker AM, Karsli-Uzunbas G et al. (2011). Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev 25: 460–470.
Gupta A, Roy S, Lazar AJ, Wang WL, McAuliffe JC, Reynoso D et al. (2010). Autophagy inhibition and antimalarials promote cell death in gastrointestinal stromal tumor (GIST). Proc Natl Acad Sci USA 107: 14333–14338.
Gutierrez MG, Munafo DB, Beron W, Colombo MI . (2004). Rab7 is required for the normal progression of the autophagic pathway in mammalian cells. J Cell Sci 117: 2687–2697.
Han J, Hou W, Goldstein LA, Lu C, Stolz DB, Yin XM et al. (2008). Involvement of protective autophagy in TRAIL resistance of apoptosis-defective tumor cells. J Biol Chem 283: 19665–19677.
Han W, Li L, Qiu S, Lu Q, Pan Q, Gu Y et al. (2007). Shikonin circumvents cancer drug resistance by induction of a necroptotic death. Mol Cancer Ther 6: 1641–1649.
Herrero-Martin G, Hoyer-Hansen M, Garcia-Garcia C, Fumarola C, Farkas T, Lopez-Rivas A et al. (2009). TAK1 activates AMPK-dependent cytoprotective autophagy in TRAIL-treated epithelial cells. EMBO J 28: 677–685.
Hoyer-Hansen M, Jaattela M . (2007). AMP-activated protein kinase: a universal regulator of autophagy? Autophagy 3: 381–383.
Hsu KF, Wu CL, Huang SC, Wu CM, Hsiao JR, Yo YT et al. (2009). Cathepsin L mediates resveratrol-induced autophagy and apoptotic cell death in cervical cancer cells. Autophagy 5: 451–460.
Huang S, Sinicrope FA . (2010). Celecoxib-induced apoptosis is enhanced by ABT-737 and by inhibition of autophagy in human colorectal cancer cells. Autophagy 6: 256–269.
Huang X, Bai HM, Chen L, Li B, Lu YC . (2010). Reduced expression of LC3B-II and Beclin 1 in glioblastoma multiforme indicates a down-regulated autophagic capacity that relates to the progression of astrocytic tumors. J Clin Neurosci 17: 1515–1519.
Hwang J, Lee HJ, Lee WH, Suk K . (2010). NF-kappaB as a common signaling pathway in ganglioside-induced autophagic cell death and activation of astrocytes. J Neuroimmunol 226: 66–72.
Indelicato M, Pucci B, Schito L, Reali V, Aventaggiato M, Mazzarino MC et al. (2010). Role of hypoxia and autophagy in MDA-MB-231 invasiveness. J Cell Physiol 223: 359–368.
Iwamaru A, Kondo Y, Iwado E, Aoki H, Fujiwara K, Yokoyama T et al. (2007). Silencing mammalian target of rapamycin signaling by small interfering RNA enhances rapamycin-induced autophagy in malignant glioma cells. Oncogene 26: 1840–1851.
Jager S, Bucci C, Tanida I, Ueno T, Kominami E, Saftig P et al. (2004). Role for Rab7 in maturation of late autophagic vacuoles. J Cell Sci 117: 4837–4848.
Jiang H, Martin V, Alonso M, Gomez-Manzano C, Fueyo J . (2010b). RB-E2F1: molecular rheostat for autophagy and apoptosis. Autophagy 6: 1216–1217.
Jiang H, Martin V, Gomez-Manzano C, Johnson DG, Alonso M, White E et al. (2010a). The RB-E2F1 pathway regulates autophagy. Cancer Res 70: 7882–7893.
Jung CH, Jun CB, Ro SH, Kim YM, Otto NM, Cao J et al. (2009). ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell 20: 1992–2003.
Kamitsuji Y, Kuroda J, Kimura S, Toyokuni S, Watanabe K, Ashihara E et al. (2008). The Bcr-Abl kinase inhibitor INNO-406 induces autophagy and different modes of cell death execution in Bcr-Abl-positive leukemias. Cell Death Differ 15: 1712–1722.
Kandouz M, Haidara K, Zhao J, Brisson ML, Batist G . (2010). The EphB2 tumor suppressor induces autophagic cell death via concomitant activation of the ERK1/2 and PI3K pathways. Cell Cycle 9: 398–407.
Kang C, You YJ, Avery L . (2007). Dual roles of autophagy in the survival of Caenorhabditis elegans during starvation. Genes Dev 21: 2161–2171.
Kang MR, Kim MS, Oh JE, Kim YR, Song SY, Kim SS et al. (2009). Frameshift mutations of autophagy-related genes ATG2B, ATG5, ATG9B and ATG12 in gastric and colorectal cancers with microsatellite instability. J Pathol 217: 702–706.
Karpathiou G, Sivridis E, Koukourakis M, Mikroulis D, Bouros D, Froudarakis M et al. (2010). LC3A autophagic activity and prognostic significance in non-small cell lung carcinomas. Chest (doi:10.1378/chest.10-1831).
Kashuba VI, Gizatullin RZ, Protopopov AI, Allikmets R, Korolev S, Li J et al. (1997). NotI linking/jumping clones of human chromosome 3: mapping of the TFRC, RAB7 and HAUSP genes to regions rearranged in leukemia and deleted in solid tumors. FEBS Lett 419: 181–185.
Kim EH, Sohn S, Kwon HJ, Kim SU, Kim MJ, Lee SJ et al. (2007). Sodium selenite induces superoxide-mediated mitochondrial damage and subsequent autophagic cell death in malignant glioma cells. Cancer Res 67: 6314–6324.
Kim J, Kundu M, Viollet B, Guan KL . (2011). AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol 13: 132–141.
Kim KW, Hwang M, Moretti L, Jaboin JJ, Cha YI, Lu B . (2008b). Autophagy upregulation by inhibitors of caspase-3 and mTOR enhances radiotherapy in a mouse model of lung cancer. Autophagy 4: 659–668.
Kim KW, Moretti L, Mitchell LR, Jung DK, Lu B . (2009b). Combined Bcl-2/mammalian target of rapamycin inhibition leads to enhanced radiosensitization via induction of apoptosis and autophagy in non-small cell lung tumor xenograft model. Clin Cancer Res 15: 6096–6105.
Kim MS, Jeong EG, Ahn CH, Kim SS, Lee SH, Yoo NJ . (2008a). Frameshift mutation of UVRAG, an autophagy-related gene, in gastric carcinomas with microsatellite instability. Hum Pathol 39: 1059–1063.
Kim RH, Coates JM, Bowles TL, McNerney GP, Sutcliffe J, Jung JU et al. (2009a). Arginine deiminase as a novel therapy for prostate cancer induces autophagy and caspase-independent apoptosis. Cancer Res 69: 700–708.
Kiyono K, Suzuki HI, Matsuyama H, Morishita Y, Komuro A, Kano MR et al. (2009). Autophagy is activated by TGF-beta and potentiates TGF-beta-mediated growth inhibition in human hepatocellular carcinoma cells. Cancer Res 69: 8844–8852.
Koesters R, Kaissling B, Lehir M, Picard N, Theilig F, Gebhardt R et al. (2010). Tubular overexpression of transforming growth factor-beta1 induces autophagy and fibrosis but not mesenchymal transition of renal epithelial cells. Am J Pathol 177: 632–643.
Komata T, Kanzawa T, Takeuchi H, Germano IM, Schreiber M, Kondo Y et al. (2003). Antitumour effect of cyclin-dependent kinase inhibitors (p16(INK4A), p18(INK4C), p19(INK4D), p21(WAF1/CIP1) and p27(KIP1)) on malignant glioma cells. Br J Cancer 88: 1277–1280.
Koukourakis MI, Giatromanolaki A, Sivridis E, Pitiakoudis M, Gatter KC, Harris AL . (2010). Beclin 1 over- and underexpression in colorectal cancer: distinct patterns relate to prognosis and tumour hypoxia. Br J Cancer 103: 1209–1214.
Kuo HP, Lee DF, Chen CT, Liu M, Chou CK, Lee HJ et al. (2010). ARD1 stabilization of TSC2 suppresses tumorigenesis through the mTOR signaling pathway. Sci Signal 3: ra9.
Laane E, Tamm KP, Buentke E, Ito K, Kharaziha P, Oscarsson J et al. (2009). Cell death induced by dexamethasone in lymphoid leukemia is mediated through initiation of autophagy. Cell Death Differ 16: 1018–1029.
Lazova R, Klump V, Pawelek J . (2010). Autophagy in cutaneous malignant melanoma. J Cutan Pathol 37: 256–268.
Lazova R, Pawelek JM . (2009). Why do melanomas get so dark? Exp Dermatol 18: 934–938.
Levine B, Kroemer G . (2008). Autophagy in the pathogenesis of disease. Cell 132: 27–42.
Levine B, Mizushima N, Virgin HW . (2011). Autophagy in immunity and inflammation. Nature 469: 323–335.
Li BX, Li CY, Peng RQ, Wu XJ, Wang HY, Wan DS et al. (2009). The expression of beclin 1 is associated with favorable prognosis in stage IIIB colon cancers. Autophagy 5: 303–306.
Li X, Fan Z . (2010). The epidermal growth factor receptor antibody cetuximab induces autophagy in cancer cells by downregulating HIF-1alpha and Bcl-2 and activating the beclin 1/hVps34 complex. Cancer Res 70: 5942–5952.
Li Z, Chen B, Wu Y, Jin F, Xia Y, Liu X . (2010). Genetic and epigenetic silencing of the beclin 1 gene in sporadic breast tumors. BMC Cancer 10: 98.
Lian J, Wu X, He F, Karnak D, Tang W, Meng Y et al. (2011). A natural BH3 mimetic induces autophagy in apoptosis-resistant prostate cancer via modulating Bcl-2-Beclin1 interaction at endoplasmic reticulum. Cell Death Differ 18: 60–71.
Liang J, Shao SH, Xu ZX, Hennessy B, Ding Z, Larrea M et al. (2007). The energy sensing LKB1-AMPK pathway regulates p27(kip1) phosphorylation mediating the decision to enter autophagy or apoptosis. Nat Cell Biol 9: 218–224.
Liu D, Huang Y, Chen B, Zeng J, Guo N, Zhang S et al. (2011). Activation of mammalian target of rapamycin pathway confers adverse outcome in nonsmall cell lung carcinoma. Cancer (doi:10.1002/cncr.25959).
Lock R, Roy S, Kenific CM, Su JS, Salas E, Ronen SM et al. (2011). Autophagy facilitates glycolysis during Ras-mediated oncogenic transformation. Mol Biol Cell 22: 165–178.
Lorin S, Borges A, Ribeiro Dos Santos L, Souquere S, Pierron G, Ryan KM et al. (2009). c-Jun NH2-terminal kinase activation is essential for DRAM-dependent induction of autophagy and apoptosis in 2-methoxyestradiol-treated Ewing sarcoma cells. Cancer Res 69: 6924–6931.
Lorin S, Pierron G, Ryan KM, Codogno P, Djavaheri-Mergny M . (2010). Evidence for the interplay between JNK and p53-DRAM signalling pathways in the regulation of autophagy. Autophagy 6: 153–154.
Lovas A, Radke D, Albrecht D, Yilmaz ZB, Moller U, Habenicht AJ et al. (2008). Differential RelA- and RelB-dependent gene transcription in LTbetaR-stimulated mouse embryonic fibroblasts. BMC Genomics 9: 606.
Lu Z, Luo RZ, Lu Y, Zhang X, Yu Q, Khare S et al. (2008). The tumor suppressor gene ARHI regulates autophagy and tumor dormancy in human ovarian cancer cells. J Clin Invest 118: 3917–3929.
Lum JJ, Bauer DE, Kong M, Harris MH, Li C, Lindsten T et al. (2005). Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 120: 237–248.
Ma X, Piao S, Wang DW, McAfee QW, Nathanson KL, Lum JJ et al. (2011). Measurements of tumor cell autophagy predict invasiveness, resistance to chemotherapy, and survival in melanoma. Clin Cancer Res 17: 1–12.
Maddodi N, Huang W, Havighurst T, Kim K, Longley BJ, Setaluri V . (2010). Induction of autophagy and inhibition of melanoma growth in vitro and in vivo by hyperactivation of oncogenic BRAF. J Invest Dermatol 130: 1657–1667.
Maiuri MC, Criollo A, Tasdemir E, Vicencio JM, Tajeddine N, Hickman JA et al. (2007a). BH3-only proteins and BH3 mimetics induce autophagy by competitively disrupting the interaction between Beclin 1 and Bcl-2/Bcl-X(L). Autophagy 3: 374–346.
Maiuri MC, Galluzzi L, Morselli E, Kepp O, Malik SA, Kroemer G . (2010). Autophagy regulation by p53. Curr Opin Cell Biol 22: 181–185.
Maiuri MC, Le Toumelin G, Criollo A, Rain JC, Gautier F, Juin P et al. (2007b). Functional and physical interaction between Bcl-X(L) and a BH3-like domain in Beclin-1. EMBO J 26: 2527–2539.
Maiuri MC, Malik SA, Morselli E, Kepp O, Criollo A, Mouchel PL et al. (2009). Stimulation of autophagy by the p53 target gene Sestrin2. Cell Cycle 8: 1571–1576.
Marino G, Salvador-Montoliu N, Fueyo A, Knecht E, Mizushima N, Lopez-Otin C . (2007). Tissue-specific autophagy alterations and increased tumorigenesis in mice deficient in Atg4C/autophagin-3. J Biol Chem 282: 18573–18583.
Mathew R, Kongara S, Beaudoin B, Karp CM, Bray K, Degenhardt K et al. (2007). Autophagy suppresses tumor progression by limiting chromosomal instability. Genes Dev 21: 1367–1381.
Mathieu V, Le Mercier M, De Neve N, Sauvage S, Gras T, Roland I et al. (2007). Galectin-1 knockdown increases sensitivity to temozolomide in a B16F10 mouse metastatic melanoma model. J Invest Dermatol 127: 2399–2410.
McPhee CK, Logan MA, Freeman MR, Baehrecke EH . (2010). Activation of autophagy during cell death requires the engulfment receptor Draper. Nature 465: 1093–1096.
Meschini S, Condello M, Calcabrini A, Marra M, Formisano G, Lista P et al. (2008). The plant alkaloid voacamine induces apoptosis-independent autophagic cell death on both sensitive and multidrug resistant human osteosarcoma cells. Autophagy 4: 1020–1033.
Miao Y, Zhang Y, Chen Y, Chen L, Wang F . (2010). GABARAP is overexpressed in colorectal carcinoma and correlates with shortened patient survival. Hepatogastroenterology 57: 257–261.
Miracco C, Cevenini G, Franchi A, Luzi P, Cosci E, Mourmouras V et al. (2010). Beclin 1 and LC3 autophagic gene expression in cutaneous melanocytic lesions. Hum Pathol 41: 503–512.
Miracco C, Cosci E, Oliveri G, Luzi P, Pacenti L, Monciatti I et al. (2007). Protein and mRNA expression of autophagy gene Beclin 1 in human brain tumours. Int J Oncol 30: 429–436.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . (2008). Autophagy fights disease through cellular self-digestion. Nature 451: 1069–1075.
Morselli E, Tasdemir E, Maiuri MC, Galluzzi L, Kepp O, Criollo A et al. (2008). Mutant p53 protein localized in the cytoplasm inhibits autophagy. Cell Cycle 7: 3056–3061.
Narita M, Young AR . (2009). Autophagy facilitates oncogene-induced senescence. Autophagy 5: 1046–1047.
Negri T, Tarantino E, Orsenigo M, Reid JF, Gariboldi M, Zambetti M et al. (2010). Chromosome band 17q21 in breast cancer: significant association between beclin 1 loss and HER2/NEU amplification. Genes Chromosomes Cancer 49: 901–909.
Nishida Y, Arakawa S, Fujitani K, Yamaguchi H, Mizuta T, Kanaseki T et al. (2009). Discovery of Atg5/Atg7-independent alternative macroautophagy. Nature 461: 654–658.
Nivon M, Richet E, Codogno P, Arrigo AP, Kretz-Remy C . (2009). Autophagy activation by NFkappaB is essential for cell survival after heat shock. Autophagy 5: 766–783.
Noble CG, Dong JM, Manser E, Song H . (2008). Bcl-xL and UVRAG cause a monomer-dimer switch in Beclin1. J Biol Chem 283: 26274–26282.
Nomura H, Uzawa K, Yamano Y, Fushimi K, Ishigami T, Kouzu Y et al. (2009). Overexpression and altered subcellular localization of autophagy-related 16-like 1 in human oral squamous-cell carcinoma: correlation with lymphovascular invasion and lymph-node metastasis. Hum Pathol 40: 83–91.
O'Donovan TR, O'Sullivan GC, McKenna S . (2011). Induction of autophagy by drug-resistant esophageal cancer cells promotes their survival and recovery following treatment with chemotherapeutics. Autophagy 7: 509–524.
Ostenfeld MS, Hoyer-Hansen M, Bastholm L, Fehrenbacher N, Olsen OD, Groth-Pedersen L et al. (2008). Anti-cancer agent siramesine is a lysosomotropic detergent that induces cytoprotective autophagosome accumulation. Autophagy 4: 487–499.
Othman EQ, Kaur G, Mutee AF, Muhammad TS, Tan ML . (2009). Immunohistochemical expression of MAP1LC3A and MAP1LC3B protein in breast carcinoma tissues. J Clin Lab Anal 23: 249–258.
Paglin S, Lee NY, Nakar C, Fitzgerald M, Plotkin J, Deuel B et al. (2005). Rapamycin-sensitive pathway regulates mitochondrial membrane potential, autophagy, and survival in irradiated MCF-7 cells. Cancer Res 65: 11061–11070.
Pankiv S, Alemu EA, Brech A, Bruun JA, Lamark T, Overvatn A et al. (2010). FYCO1 is a Rab7 effector that binds to LC3 and PI3P to mediate microtubule plus end-directed vesicle transport. J Cell Biol 188: 253–269.
Pankiv S, Johansen T . (2010). FYCO1: linking autophagosomes to microtubule plus end-directing molecular motors. Autophagy 6: 550–552.
Park MA, Yacoub A, Rahmani M, Zhang G, Hart L, Hagan MP et al. (2008). OSU-03012 stimulates PKR-like endoplasmic reticulum-dependent increases in 70-kDa heat shock protein expression, attenuating its lethal actions in transformed cells. Mol Pharmacol 73: 1168–1184.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. (2005). Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122: 927–939.
Pirtoli L, Cevenini G, Tini P, Vannini M, Oliveri G, Marsili S et al. (2009). The prognostic role of Beclin 1 protein expression in high-grade gliomas. Autophagy 5: 930–936.
Platt FM, Hurst CD, Taylor CF, Gregory WM, Harnden P, Knowles MA . (2009). Spectrum of phosphatidylinositol 3-kinase pathway gene alterations in bladder cancer. Clin Cancer Res 15: 6008–6017.
Polager S, Ofir M, Ginsberg D . (2008). E2F1 regulates autophagy and the transcription of autophagy genes. Oncogene 27: 4860–4864.
Portillo JA, Okenka G, Reed E, Subauste A, Van Grol J, Gentil K et al. (2010). The CD40-autophagy pathway is needed for host protection despite IFN-Gamma-dependent immunity and CD40 induces autophagy via control of P21 levels. PLoS One 5: e14472.
Qadir MA, Kwok B, Dragowska WH, To KH, Le D, Bally MB et al. (2008). Macroautophagy inhibition sensitizes tamoxifen-resistant breast cancer cells and enhances mitochondrial depolarization. Breast Cancer Res Treat 112: 389–403.
Reef S, Kimchi A . (2008). Nucleolar p19ARF, unlike mitochondrial smARF, is incapable of inducing p53-independent autophagy. Autophagy 4: 866–869.
Reef S, Shifman O, Oren M, Kimchi A . (2007). The autophagic inducer smARF interacts with and is stabilized by the mitochondrial p32 protein. Oncogene 26: 6677–6683.
Reef S, Zalckvar E, Shifman O, Bialik S, Sabanay H, Oren M et al. (2006). A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol Cell 22: 463–475.
Roca H, Varsos Z, Pienta KJ . (2008). CCL2 protects prostate cancer PC3 cells from autophagic death via phosphatidylinositol 3-kinase/AKT-dependent survivin up-regulation. J Biol Chem 283: 25057–25073.
Russell SE, Hickey GI, Lowry WS, White P, Atkinson RJ . (1990). Allele loss from chromosome 17 in ovarian cancer. Oncogene 5: 1581–1583.
Saito H, Inazawa J, Saito S, Kasumi F, Koi S, Sagae S et al. (1993). Detailed deletion mapping of chromosome 17q in ovarian and breast cancers: 2-cM region on 17q21.3 often and commonly deleted in tumors. Cancer Res 53: 3382–3385.
Sanchez-Cespedes M . (2007). A role for LKB1 gene in human cancer beyond the Peutz-Jeghers syndrome. Oncogene 26: 7825–7832.
Scarlatti F, Maffei R, Beau I, Codogno P, Ghidoni R . (2008). Role of non-canonical Beclin 1-independent autophagy in cell death induced by resveratrol in human breast cancer cells. Cell Death Differ 15: 1318–1329.
Scherz-Shouval R, Weidberg H, Gonen C, Wilder S, Elazar Z, Oren M . (2010). p53-dependent regulation of autophagy protein LC3 supports cancer cell survival under prolonged starvation. Proc Natl Acad Sci USA 107: 18511–18516.
Serrano M, Lin AW, McCurrach ME, Beach D, Lowe SW . (1997). Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 88: 593–602.
Shingu T, Fujiwara K, Bogler O, Akiyama Y, Moritake K, Shinojima N et al. (2009). Inhibition of autophagy at a late stage enhances imatinib-induced cytotoxicity in human malignant glioma cells. Int J Cancer 124: 1060–1071.
Sivridis E, Koukourakis MI, Zois CE, Ledaki I, Ferguson DJ, Harris AL et al. (2010). LC3A-positive light microscopy detected patterns of autophagy and prognosis in operable breast carcinomas. Am J Pathol 176: 2477–2489.
Steffan JJ, Cardelli JA . (2010). Thiazolidinediones induce Rab7-RILP-MAPK-dependent juxtanuclear lysosome aggregation and reduce tumor cell invasion. Traffic 11: 274–286.
Steffan JJ, Williams BC, Welbourne T, Cardelli JA . (2010). HGF-induced invasion by prostate tumor cells requires anterograde lysosome trafficking and activity of Na+-H+ exchangers. J Cell Sci 123: 1151–1159.
Stenmark H . (2010). The Sir Hans Krebs Lecture. How a lipid mediates tumour suppression. Delivered on 29 June 2010 at the 35th FEBS Congress in Gothenburg, Sweden. FEBS J 277: 4837–4848.
Suzuki HI, Kiyono K, Miyazono K . (2010). Regulation of autophagy by transforming growth factor-beta (TGFbeta) signaling. Autophagy 6: 645–647.
Tangir J, Muto MG, Berkowitz RS, Welch WR, Bell DA, Mok SC . (1996). A 400 kb novel deletion unit centromeric to the BRCA1 gene in sporadic epithelial ovarian cancer. Oncogene 12: 735–740.
Tasdemir E, Maiuri MC, Galluzzi L, Vitale I, Djavaheri-Mergny M, D'Amelio M et al. (2008). Regulation of autophagy by cytoplasmic p53. Nat Cell Biol 10: 676–687.
Tormo D, Checinska A, Alonso-Curbelo D, Perez-Guijarro E, Canon E, Riveiro-Falkenbach E et al. (2009). Targeted activation of innate immunity for therapeutic induction of autophagy and apoptosis in melanoma cells. Cancer Cell 16: 103–114.
Totoki Y, Tatsuno K, Yamamoto S, Arai Y, Hosoda F, Ishikawa S et al. (2011). High-resolution characterization of a hepatocellular carcinoma genome. Nat Genet 43: 464–469.
Turcotte S, Chan DA, Sutphin PD, Hay MP, Denny WA, Giaccia AJ . (2008). A molecule targeting VHL-deficient renal cell carcinoma that induces autophagy. Cancer Cell 14: 90–102.
Ueno T, Sato W, Horie Y, Komatsu M, Tanida I, Yoshida M et al. (2008). Loss of Pten, a tumor suppressor, causes the strong inhibition of autophagy without affecting LC3 lipidation. Autophagy 4: 692–700.
Voss V, Senft C, Lang V, Ronellenfitsch MW, Steinbach JP, Seifert V et al. (2010). The pan-Bcl-2 inhibitor (-)-gossypol triggers autophagic cell death in malignant glioma. Mol Cancer Res 8: 1002–1016.
Wan XB, Fan XJ, Chen MY, Xiang J, Huang PY, Guo L et al. (2010). Elevated Beclin 1 expression is correlated with HIF-1alpha in predicting poor prognosis of nasopharyngeal carcinoma. Autophagy 6: 395–404.
Wang J, Whiteman MW, Lian H, Wang G, Singh A, Huang D et al. (2009). A non-canonical MEK/ERK signaling pathway regulates autophagy via regulating Beclin 1. J Biol Chem 284: 21412–21424.
Wild P, Dikic I . (2010). Mitochondria get a Parkin’ ticket. Nat Cell Biol 12: 104–106.
Wu WK, Cho CH, Lee CW, Wu K, Fan D, Yu J et al. (2010a). Proteasome inhibition: a new therapeutic strategy to cancer treatment. Cancer Lett 293: 15–22.
Wu WK, Cho CH, Lee CW, Wu YC, Yu L, Li ZJ et al. (2010c). Macroautophagy and ERK phosphorylation counteract the antiproliferative effect of proteasome inhibitor in gastric cancer cells. Autophagy 6: 228–238.
Wu WK, Sakamoto KM, Milani M, Aldana-Masankgay G, Fan D, Wu K et al. (2010b). Macroautophagy modulates cellular response to proteasome inhibitors in cancer therapy. Drug Resist Updat 13: 87–92.
Xu DZ, Geng QR, Tian Y, Cai MY, Fang XJ, Zhan YQ et al. (2010). Activated mammalian target of rapamycin is a potential therapeutic target in gastric cancer. BMC Cancer 10: 536.
Yacoub A, Hamed HA, Allegood J, Mitchell C, Spiegel S, Lesniak MS et al. (2010). PERK-dependent regulation of ceramide synthase 6 and thioredoxin play a key role in mda-7/IL-24-induced killing of primary human glioblastoma multiforme cells. Cancer Res 70: 1120–1129.
Yang PM, Liu YL, Lin YC, Shun CT, Wu MS, Chen CC . (2010). Inhibition of autophagy enhances anticancer effects of atorvastatin in digestive malignancies. Cancer Res 70: 7699–7709.
Yang S, Wang X, Contino G, Liesa M, Sahin E, Ying H et al. (2011). Pancreatic cancers require autophagy for tumor growth. Genes Dev 25: 717–729.
Yang Z, Klionsky DJ . (2010). Mammalian autophagy: core molecular machinery and signaling regulation. Curr Opin Cell Biol 22: 124–131.
Yaswen P, Campisi J . (2007). Oncogene-induced senescence pathways weave an intricate tapestry. Cell 128: 233–234.
Yee KS, Wilkinson S, James J, Ryan KM, Vousden KH . (2009). PUMA- and Bax-induced autophagy contributes to apoptosis. Cell Death Differ 16: 1135–1145.
Yoshioka A, Miyata H, Doki Y, Yamasaki M, Sohma I, Gotoh K et al. (2008). LC3, an autophagosome marker, is highly expressed in gastrointestinal cancers. Int J Oncol 33: 461–468.
Young AR, Narita M, Ferreira M, Kirschner K, Sadaie M, Darot JF et al. (2009). Autophagy mediates the mitotic senescence transition. Genes Dev 23: 798–803.
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L et al. (2006). Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol 8: 1124–1132.
Yue Z, Jin S, Yang C, Levine AJ, Heintz N . (2003). Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA 100: 15077–15082.
Zeng X, Kinsella TJ . (2008). Mammalian target of rapamycin and S6 kinase 1 positively regulate 6-thioguanine-induced autophagy. Cancer Res 68: 2384–2390.
Zhang L, Yu J, Pan H, Hu P, Hao Y, Cai W et al. (2007). Small molecule regulators of autophagy identified by an image-based high-throughput screen. Proc Natl Acad Sci USA 104: 19023–19028.
Zhang YJ, Dai Q, Sun DF, Xiong H, Tian XQ, Gao FH et al. (2009b). mTOR signaling pathway is a target for the treatment of colorectal cancer. Ann Surg Oncol 16: 2617–2628.
Zhang Z, Shao Z, Xiong L, Che B, Deng C, Xu W . (2009a). Expression of Beclin1 in osteosarcoma and the effects of down-regulation of autophagy on the chemotherapeutic sensitivity. J Huazhong Univ Sci Technolog Med Sci 29: 737–740.
Zhao Y, Yang J, Liao W, Liu X, Zhang H, Wang S et al. (2010). Cytosolic FoxO1 is essential for the induction of autophagy and tumour suppressor activity. Nat Cell Biol 12: 665–675.
Zhou F, Yang Y, Xing D . (2011). Bcl-2 and Bcl-xL play important roles in the crosstalk between autophagy and apoptosis. FEBS J 278: 403–413.
Acknowledgements
This work was supported by research grant from the CUHK Group Research Scheme (3110043) and CUHK Focused Investments Scheme-Scheme C.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Wu, W., Coffelt, S., Cho, C. et al. The autophagic paradox in cancer therapy. Oncogene 31, 939–953 (2012). https://doi.org/10.1038/onc.2011.295
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.295
Keywords
This article is cited by
-
Genetic assessment of pathogenic germline alterations in lysosomal genes among Asian patients with pancreatic ductal adenocarcinoma
Journal of Translational Medicine (2023)
-
Autophagy in Cancer Therapy: Advances and Prospects
Current Molecular Biology Reports (2023)
-
Exosomal non-coding RNAs-mediated EGFR-TKIs resistance in NSCLC with EGFR mutation
Medical Oncology (2023)
-
FOXA1 prevents nutrients deprivation induced autophagic cell death through inducing loss of imprinting of IGF2 in lung adenocarcinoma
Cell Death & Disease (2022)
-
PNO1 regulates autophagy and apoptosis of hepatocellular carcinoma via the MAPK signaling pathway
Cell Death & Disease (2021)