Abstract
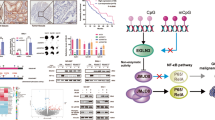
The IRX1 tumor suppressor gene is located on 5p15.33, a cancer susceptibility locus. Loss of heterozygosity of 5p15.33 in gastric cancer was identified in our previous work. In this study, we analyzed the molecular features and function of IRX1. We found that IRX1 expression was lost or reduced in gastric cancer. However, no mutations were identified in IRX1-encoding regions. IRX1 transcription was suppressed by hypermethylation, and the expression of IRX1 mRNA was partially restored in gastric cancer cells after 5-Aza-dC treatment. Restoring IRX1 expression in SGC-7901 and NCI-N87 gastric cancer cells inhibited growth, invasion and tumorigenesis in vitro and in vivo. We identified a number of target genes by global microarray analysis after IRX1 transfection combined with real-time PCR and chromatin immunoprecipitation assay. BDKRB2, an angiogenesis-related gene, HIST2H2BE and FGF7, cell proliferation and invasion-related genes, were identified as direct IRX1 target genes. The hypermethylation of IRX1 was not only detected in primary gastric cancer tissues but also in the peripheral blood of gastric cancer patients, suggesting IRX1 could potentially serve as a biomarker for gastric cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Alarcon P, Rodriguez-Seguel E, Fernandez-Gonzalez A, Rubio R, Gomez-Skarmeta JL . (2008). A dual requirement for Iroquois genes during Xenopus kidney development. Development 135: 3197–3207.
Asaka S, Fujimoto T, Akaishi J, Ogawa K, Onda M . (2006). Genetic prognostic index influences patient outcome for node-positive breast cancer. Surg Today 36: 793–801.
Becker MB, Zulch A, Bosse A, Gruss P . (2001). Irx1 and Irx2 expression in early lung development. Mech Dev 106: 155–158.
Bennett KL, Karpenko M, Lin MT, Claus R, Arab K, Dyckhoff G et al. (2008). Frequently methylated tumor suppressor genes in head and neck squamous cell carcinoma. Cancer Res 68: 4494–4499.
Bergman Y, Mostoslavsky R . (1998). DNA demethylation: turning genes on. Biol Chem 379: 401–407.
Cairns P . (2009). 5′-azacytidine expression arrays. Methods Mol Biol 507: 165–174.
Choi IS, Wu TT . (2005). Epigenetic alterations in gastric carcinogenesis. Cell Res 15: 247–254.
Duffy MJ, Napieralski R, Martens JW, Span PN, Spyratos F, Sweep FC et al. (2009). Methylated genes as new cancer biomarkers. Eur J Cancer 45: 335–346.
Duggan DJ, Bittner M, Chen Y, Meltzer P, Trent JM . (1999). Expression profiling using cDNA microarrays. Nat Genet 21: 10–14.
Esteller M . (2007). Epigenetic gene silencing in cancer: the DNA hypermethylome. Hum Mol Genet 16 (Spec No 1): R50–R59.
Ferguson CA, Tucker AS, Christensen L, Lau AL, Matzuk MM, Sharpe PT . (1998). Activin is an essential early mesenchymal signal in tooth development that is required for patterning of the murine dentition. Genes Dev 12: 2636–2649.
Garinis GA, Patrinos GP, Spanakis NE, Menounos PG . (2002). DNA hypermethylation: when tumour suppressor genes go silent. Hum Genet 111: 115–127.
Hao Y, Yu Y, Wang L, Yan M, Ji J, Qu Y et al. (2008). IPO-38 is identified as a novel serum biomarker of gastric cancer based on clinical proteomics technology. J Proteome Res 7: 3668–3677.
Hellebrekers DM, Melotte V, Vire E, Langenkamp E, Molema G, Fuks F et al. (2007). Identification of epigenetically silenced genes in tumor endothelial cells. Cancer Res 67: 4138–4148.
Huang Q, Chen W, Wang L, Lin W, Lin J, Lin X . (2008). Identification of transgelin as a potential novel biomarker for gastric adenocarcinoma based on proteomics technology. J Cancer Res Clin Oncol 134: 1219–1227.
Hurtubise A, Bernstein ML, Momparler RL . (2008). Preclinical evaluation of the antineoplastic action of 5-aza-2′-deoxycytidine and different histone deacetylase inhibitors on human Ewing’s sarcoma cells. Cancer Cell Int 8: 16.
Ikeda Y, Hayashi I, Kamoshita E, Yamazaki A, Endo H, Ishihara K et al. (2004). Host stromal bradykinin B2 receptor signaling facilitates tumor-associated angiogenesis and tumor growth. Cancer Res 64: 5178–5185.
Iorns E, Lord CJ, Grigoriadis A, McDonald S, Fenwick K, Mackay A et al. (2009). Integrated functional, gene expression and genomic analysis for the identification of cancer targets. PLoS One 4: e5120.
Ishihara K, Hayash I, Yamashina S, Majima M . (2001). A potential role of bradykinin in angiogenesis and growth of S-180 mouse tumors. Jpn J Pharmacol 87: 318–326.
Ishihara K, Kamata M, Hayashi I, Yamashina S, Majima M . (2002). Roles of bradykinin in vascular permeability and angiogenesis in solid tumor. Int Immunopharmacol 2: 499–509.
Jee CD, Kim MA, Jung EJ, Kim J, Kim WH . (2009). Identification of genes epigenetically silenced by CpG methylation in human gastric carcinoma. Eur J Cancer 45: 1282–1293.
Jin G, Xu L, Shu Y, Tian T, Liang J, Xu Y et al. (2009). Common genetic variants on 5p15.33 contribute to risk of lung adenocarcinoma in a Chinese population. Carcinogenesis 30: 987–990.
Kamei M, Kato M, Mochizuki K, Kuroda K, Sato S, Hashizume S et al. (1992). Serodiagnosis of cancers by ELISA of anti-histone H2B antibody. Biotherapy 4: 17–22.
Kawakami K, Brabender J, Lord RV, Groshen S, Greenwald BD, Krasna MJ et al. (2000). Hypermethylated APC DNA in plasma and prognosis of patients with esophageal adenocarcinoma. J Natl Cancer Inst 92: 1805–1811.
Knudson AG . (1996). Hereditary cancer: two hits revisited. J Cancer Res Clin Oncol 122: 135–140.
Knudson AG . (2001). Two genetic hits (more or less) to cancer. Nat Rev Cancer 1: 157–162.
Kuga D, Mizoguchi M, Guan Y, Hata N, Yoshimoto K, Shono T et al. (2008). Prevalence of copy-number neutral LOH in glioblastomas revealed by genomewide analysis of laser-microdissected tissues. Neuro Oncol 10: 995–1003.
Lai IR, Lee WJ, Huang MT, Lin HH . (2002). Comparison of serum CA72-4, CEA, TPA, CA19-9 and CA125 levels in gastric cancer patients and correlation with recurrence. Hepatogastroenterology 49: 1157–1160.
Lemaire M, Chabot GG, Raynal NJ, Momparler LF, Hurtubise A, Bernstein ML et al. (2008). Importance of dose-schedule of 5-aza-2′-deoxycytidine for epigenetic therapy of cancer. BMC Cancer 8: 128.
Lewis MT, Ross S, Strickland PA, Snyder CJ, Daniel CW . (1999). Regulated expression patterns of IRX-2, an Iroquois-class homeobox gene, in the human breast. Cell Tissue Res 296: 549–554.
Lu Y, Yu Y, Zhu Z, Xu H, Ji J, Bu L et al. (2005). Identification of a new target region by loss of heterozygosity at 5p15.33 in sporadic gastric carcinomas: genotype and phenotype related. Cancer Lett 224: 329–337.
McKay JD, Hung RJ, Gaborieau V, Boffetta P, Chabrier A, Byrnes G et al. (2008). Lung cancer susceptibility locus at 5p15.33. Nat Genet 40: 1404–1406.
Momparler RL . (2003). Cancer epigenetics. Oncogene 22: 6479–6483.
Perez-Diez A, Morgun A, Shulzhenko N . (2007). Microarrays for cancer diagnosis and classification. Adv Exp Med Biol 593: 74–85.
Plendl J, Snyman C, Naidoo S, Sawant S, Mahabeer R, Bhoola KD . (2000). Expression of tissue kallikrein and kinin receptors in angiogenic microvascular endothelial cells. Biol Chem 381: 1103–1115.
Quackenbush J . (2006). Microarray analysis and tumor classification. N Engl J Med 354: 2463–2472.
Robertson KD, Jones PA . (2000). DNA methylation: past, present and future directions. Carcinogenesis 21: 461–467.
Ropiquet F, Huguenin S, Villette JM, Ronfle V, Le Brun G, Maitland NJ et al. (1999). FGF7/KGF triggers cell transformation and invasion on immortalised human prostatic epithelial PNT1A cells. Int J Cancer 82: 237–243.
Schneider-Stock R, Ocker M . (2007). Epigenetic therapy in cancer: molecular background and clinical development of histone deacetylase and DNA methyltransferase inhibitors. IDrugs 10: 557–561.
Shaker S, Bernstein M, Momparler RL . (2004). Antineoplastic action of 5-aza-2′-deoxycytidine. (Dacogen) and depsipeptide on Raji lymphoma cells. Oncol Rep 11: 1253–1256.
Shaoul R, Eliahu L, Sher I, Hamlet Y, Miselevich I, Goldshmidt O et al. (2006). Elevated expression of FGF7 protein is common in human gastric diseases. Biochem Biophys Res Commun 350: 825–833.
Tamura M, Gu J, Matsumoto K, Aota S, Parsons R, Yamada KM . (1998). Inhibition of cell migration, spreading, and focal adhesions by tumor suppressor PTEN. Science 280: 1614–1617.
Tamura M, Gu J, Takino T, Yamada KM . (1999). Tumor suppressor PTEN inhibition of cell invasion, migration, and growth: differential involvement of focal adhesion kinase and p130Cas. Cancer Res 59: 442–449.
Telford JL, Covacci A, Ghiara P, Montecucco C, Rappuoli R . (1994). Unravelling the pathogenic role of Helicobacter pylori in peptic ulcer: potential new therapies and vaccines. Trends Biotechnol 12: 420–426.
Tocchi A, Costa G, Lepre L, Liotta G, Mazzoni G, Cianetti A et al. (1998). The role of serum and gastric juice levels of carcinoembryonic antigen, CA19.9 and CA72.4 in patients with gastric cancer. J Cancer Res Clin Oncol 124: 450–455.
Tommasi S, Karm DL, Wu X, Yen Y, Pfeifer GP . (2009). Methylation of homeobox genes is a frequent and early epigenetic event in breast cancer. Breast Cancer Res 11: R14.
Toyota M, Issa JP . (2005). Epigenetic changes in solid and hematopoietic tumors. Semin Oncol 32: 521–530.
Tycko B . (2000). Epigenetic gene silencing in cancer. J Clin Invest 105: 401–407.
Ushijima T, Nakajima T, Maekita T . (2006). DNA methylation as a marker for the past and future. J Gastroenterol 41: 401–407.
van Tuyl M, Liu J, Groenman F, Ridsdale R, Han RN, Venkatesh V et al. (2006). Iroquois genes influence proximo-distal morphogenesis during rat lung development. Am J Physiol Lung Cell Mol Physiol 290: L777–L789.
Yamashita K, Upadhyay S, Osada M, Hoque MO, Xiao Y, Mori M et al. (2002). Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell 2: 485–495.
Ychou M, Duffour J, Karma A, Gourgou S, Grenier J . (2000). Clinical significance and prognostic value of CA72-4 compared with CEA and CA19-9 in patients with gastric cancer. Dis Markers 16: 105–110.
Yu YY, Ji J, Lu Y, Bu L, Liu BY, Zhu ZG et al. (2006). [High-resolution analysis of chromosome 5 and identification of candidate genes in gastric cancer]. Zhonghua Zhong Liu Za Zhi 28: 84–87.
Zhao H, Kim Y, Wang P, Lapointe J, Tibshirani R, Pollack JR et al. (2005). Genome-wide characterization of gene expression variations and DNA copy number changes in prostate cancer cell lines. Prostate 63: 187–197.
Acknowledgements
This work was supported, in part by grants from the National Natural Science Foundation of China (30572127, 30770961 and 30973486), the Chinese National High Tech Program (863-2006AA02A402, 2006AA02A301), the Shanghai Pu Jiang Project (PJ200700367), Key Research Project from Shanghai Science and Technology Commission (09DZ1950101 and 09JC1409600), and Shanghai Charity Foundation for Cancer Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Guo, X., Liu, W., Pan, Y. et al. Homeobox gene IRX1 is a tumor suppressor gene in gastric carcinoma. Oncogene 29, 3908–3920 (2010). https://doi.org/10.1038/onc.2010.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.143
Keywords
This article is cited by
-
Analysis of different adipose depot gene expression in cachectic patients with gastric cancer
Nutrition & Metabolism (2022)
-
Prognostic value of Iroquois homeobox 1 methylation in non-small cell lung cancers
Genes & Genomics (2020)
-
Gastric cancer and gene copy number variation: emerging cancer drivers for targeted therapy
Oncogene (2016)
-
Vimentin Regulates Neuroplasticity in Transected Spinal Cord Rats Associated with micRNA138
Molecular Neurobiology (2015)
-
Hypermethylation of EBF3 and IRX1 Genes in Synovial Fibroblasts of Patients with Rheumatoid Arthritis
Molecules and Cells (2013)