Abstract
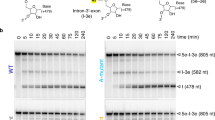
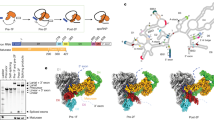
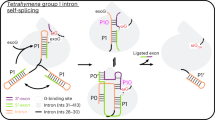
Despite the biological importance of self-splicing group II introns, little is known about their structural organization. Synthetic incorporation of site-specific photo-cross-linkers within catalytic domains resulted in functional distance constraints that, when combined with known tertiary interactions, provide a three-dimensional view of the active intron architecture. All functionalities important for both steps of splicing are proximal before the first step, suggestive of a single active-site region for group II intron catalysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lehmann, K. & Schmidt, U. Crit. Rev. Biochem. Mol. Biol. 38, 249–303 (2003).
Boeke, J. Genome Res. 13, 1975–1983 (2003).
Costa, M., Michel, F. & Westhof, E. EMBO J. 19, 5007–5018 (2000).
Fedorova, O., Mitros, T. & Pyle, A.M. J. Mol. Biol. 330, 197–209 (2003).
Mikheeva, S., Murray, H.L., Zhou, H., Turczyk, B.M. & Jarrell, K.A. RNA 6, 1509–1515 (2000).
Podar, M. & Perlman, P.S. RNA 5, 318–329 (1999).
Jacquier, A. & Jacquesson-Breuleux, N. J. Mol. Biol. 219, 415–428 (1991).
Chanfreau, G. & Jacquier, A. Science 266, 1383–1387 (1994).
Chanfreau, G. & Jacquier, A. EMBO J. 15, 3466–3476 (1996).
Adams, P.L., Stahley, M.R., Kosek, A.B., Wang, J. & Strobel, S.A. Nature 430, 45–50 (2004).
Acknowledgements
We thank O. Fedorova for help with photo-cross-linking and RNA synthesis, L. Wadley, M. Schmeing and J. Wang for help with modeling and L. Shapiro for helpful discussions. This research was supported by fellowships from Praxis XXI, Gulbenkian 43058 (A.D.L.) and by US National Institutes of Health grants GM0822414, and GM0822415 (S.H.) and GM50313 (A.M.P.). A.M.P. is an Investigator with the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Schematic group II intron secondary structure. (PDF 412 kb)
Supplementary Fig. 2
The products of splicing from a D56 molecule containing a single-nucleotide 3′-exon. (PDF 410 kb)
Supplementary Fig. 3
Secondary structure of the intron with color coding of corresponding domains in the tertiary structure that were modeled (Supplementary Fig. 4). (PDF 957 kb)
Supplementary Fig. 4
Three-dimensional model, color-coded as in Supplementary Fig. 3. (PDF 198 kb)
Supplementary Table 1
Crosslinks identified in the present study. (PDF 67 kb)
Supplementary Table 2
Additional known constraints on group II intron architecture. (PDF 101 kb)
Rights and permissions
About this article
Cite this article
de Lencastre, A., Hamill, S. & Pyle, A. A single active-site region for a group II intron. Nat Struct Mol Biol 12, 626–627 (2005). https://doi.org/10.1038/nsmb957
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb957
This article is cited by
-
Visualizing group II intron dynamics between the first and second steps of splicing
Nature Communications (2020)
-
Inactivation of group II intron RmInt1 in the Sinorhizobium meliloti genome
Scientific Reports (2015)
-
Now on display: a gallery of group II intron structures at different stages of catalysis
Mobile DNA (2013)
-
Crystal structure of a group II intron in the pre-catalytic state
Nature Structural & Molecular Biology (2012)
-
Exon sequence requirements for excision in vivo of the bacterial group II intron RmInt1
BMC Molecular Biology (2011)