Abstract
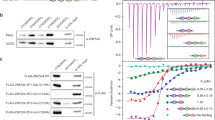
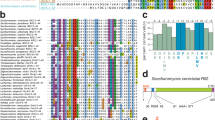
Cdc13, Stn1 and Ten1 are essential yeast proteins that both protect chromosome termini from unregulated resection and regulate telomere length. Cdc13, which localizes to telomeres through high-affinity binding to telomeric single-stranded DNA, has been extensively characterized, whereas the contribution(s) of the Cdc13-associated Stn1 and Ten1 proteins to telomere function have remained unclear. We show here that Stn1 and Ten1 are DNA-binding proteins with specificity for telomeric DNA substrates. Furthermore, Stn1 and Ten1 show similarities to Rpa2 and Rpa3, subunits of the heterotrimeric replication protein A (RPA) complex, which is the major single-stranded DNA–binding activity in eukaryotic cells. We propose that Cdc13, Stn1 and Ten1 function as a telomere-specific RPA-like complex. Identification of an RPA-like complex that is targeted to a specific region of the genome suggests that multiple RPA-like complexes have evolved, each making individual contributions to genomic stability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Shay, J.W. & Wright, W.E. in Telomeres 2nd edn. (eds. de Lange, T., Lundblad, V. & Blackburn, E.) 81–108 (Cold Spring Harbor Laboratory, New York, 2005).
Stewart, S.A. & Weinberg, R.A. Telomeres: cancer to human aging. Annu. Rev. Cell Dev. Biol. 22, 531–557 (2006).
Greider, C.W. & Blackburn, E.H. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43, 405–413 (1985).
Lundblad, V. & Szostak, J.W. A mutant with a defect in telomere elongation leads to senescence in yeast. Cell 57, 633–643 (1989).
Bodnar, A.G. et al. Extension of life-span by introduction of telomerase into normal human cells. Science 279, 349–352 (1998).
Smogorzewska, A. & de Lange, T. Regulation of telomerase by telomeric proteins. Annu. Rev. Biochem. 73, 177–208 (2004).
Hug, N. & Lingner, J. Telomere length homeostasis. Chromosoma 115, 413–425 (2006).
Blackburn, E.H. Telomere states and cell fates. Nature 408, 53–56 (2000).
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Nugent, C.I., Hughes, T.R., Lue, N.F. & Lundblad, V. Cdc13p: a single-strand telomeric DNA-binding protein with a dual role in yeast telomere maintenance. Science 274, 249–252 (1996).
Taggart, A.K., Teng, S.C. & Zakian, V.A. Est1p as a cell cycle-regulated activator of telomere-bound telomerase. Science 297, 1023–1026 (2002).
Weinert, T.A. & Hartwell, L.H. Cell cycle arrest of cdc mutants and specificity of the RAD9 checkpoint. Genetics 134, 63–80 (1993).
Garvik, B., Carson, M. & Hartwell, L. Single-stranded DNA arising at telomeres in cdc13 mutants may constitute a specific signal for the RAD9 checkpoint. Mol. Cell. Biol. 15, 6128–6138 (1995).
Evans, S.K. & Lundblad, V. Est1 and Cdc13 as comediators of telomerase access. Science 286, 117–120 (1999).
Pennock, E., Buckley, K. & Lundblad, V. Cdc13 delivers separate complexes to the telomere for end protection and replication. Cell 104, 387–396 (2001).
Bianchi, A., Negrini, S. & Shore, D. Delivery of yeast telomerase to a DNA break depends on the recruitment functions of Cdc13 and Est1. Mol. Cell 16, 139–146 (2004).
Grandin, N., Reed, S.I. & Charbonneau, M. Stn1, a new Saccharomyces cerevisiae protein, is implicated in telomere size regulation in association with Cdc13. Genes Dev. 11, 512–527 (1997).
Grandin, N., Damon, C. & Charbonneau, M. Ten1 functions in telomere end protection and length regulation in association with Stn1 and Cdc13. EMBO J. 20, 1173–1183 (2001).
Vodenicharov, M.D. & Wellinger, R.J. DNA degradation at unprotected telomeres in yeast is regulated by the CDK1 (CDC28/Clb) cell cycle kinase. Mol. Cell 24, 127–137 (2006).
Chandra, A., Hughes, T.R., Nugent, C.I. & Lundblad, V. Cdc13 both positively and negatively regulates telomere replication. Genes Dev. 15, 404–414 (2001).
Wold, M.S. Replication Protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 66, 61–92 (1997).
Iftode, C., Daniely, Y. & Borowiec, J.A. Replication Protein A (RPA): the eukaryotic SSB. Crit. Rev. Biochem. Mol. Biol. 34, 141–180 (1999).
Bochkarev, A. & Bochkareva, E. From RPA to BRCA2: lessons from single-stranded DNA binding by the OB-fold. Curr. Opin. Struct. Biol. 14, 36–42 (2004).
Soding, J., Biegert, A. & Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Bochkarev, A., Bochkareva, E., Frappier, L. & Edwards, A.M. The crystal structure of the complex of Replication Protein A subunits RPA32 and RPA14 reveals a mechanism for single-stranded DNA binding. EMBO J. 18, 4498–4504 (1999).
Ginalski, K., Elofsson, A., Fischer, D. & Rychlewski, L. 3D-Jury: a simple approach to improve protein structure predictions. Bioinformatics 19, 1015–1018 (2003).
Arcus, V. OB-fold domains: a snapshot of the evolution of sequence, structure and function. Curr. Opin. Struct. Biol. 12, 794–801 (2002).
Theobald, D.L., Mitton-Fry, R.M. & Wuttke, D.S. Nucleic acid recognition by OB-fold proteins. Annu. Rev. Biophys. Biomol. Struct. 32, 115–133 (2003).
Theobald, D.L. & Wuttke, D.S. Divergent evolution within protein superfolds inferred from profile-based phylogenetics. J. Mol. Biol. 354, 722–737 (2005).
Santocanale, C., Neecke, H., Longhese, M.P., Lucchini, G. & Plevani, P. Mutations in the gene encoding the 34 kDa subunit of yeast Replication Protein A cause defective S phase progression. J. Mol. Biol. 254, 595–607 (1995).
Philipova, D. et al. A hierarchy of SSB protomers in Replication Protein A. Genes Dev. 10, 2222–2233 (1996).
Maniar, H.S., Wilson, R. & Brill, S.J. Roles of replication protein-A subunits 2 and 3 in DNA replication fork movement in Saccharomyces cerevisiae. Genetics 145, 891–902 (1997).
Mitton-Fry, R.M., Anderson, E.M., Hughes, T.R., Lundblad, V. & Wuttke, D.S. Conserved structure for single-stranded telomeric DNA recognition. Science 296, 145–147 (2002).
Sibenaller, Z.A., Sorensen, B.R. & Wold, M.S. The 32- and 14-kilodalton subunits of Replication Protein A are responsible for species-specific interactions with single-stranded DNA. Biochemistry 37, 12496–12506 (1998).
Bochkareva, E., Frappier, L., Edwards, A.M. & Bochkarev, A. The RPA32 subunit of human Replication Protein A contains a single-stranded DNA-binding domain. J. Biol. Chem. 273, 3932–3936 (1998).
Lin, Y.L., Chen, C., Keshav, K.F., Winchester, E. & Dutta, A. Dissection of functional domains of the human DNA replication protein complex Replication Protein A. J. Biol. Chem. 271, 17190–17198 (1996).
Venclovas, C. & Thelen, M.P. Structure-based predictions of Rad1, Rad9, Hus1 and Rad17 participation in sliding clamp and clamp-loading complexes. Nucleic Acids Res. 28, 2481–2493 (2000).
Green, C.M., Erdjument-Bromage, H., Tempst, P. & Lowndes, N.F. A novel Rad24 checkpoint protein complex closely related to replication factor C. Curr. Biol. 10, 39–42 (2000).
Parrilla-Castellar, E.R., Arlander, S.J. & Karnitz, L. Dial 9–1-1 for DNA damage: the Rad9-Hus1-Rad1 (9–1-1) clamp complex. DNA Repair (Amst.) 3, 1009–1014 (2004).
Aroya, S.B. & Kupiec, M. The Elg1 replication factor C-like complex: a novel guardian of genome stability. DNA Repair (Amst.) 4, 409–417 (2005).
Wold, M.S., Weinberg, D.H., Virshup, D.M., Li, J.J. & Kelly, T.J. Identification of cellular proteins required for simian virus 40 DNA replication. J. Biol. Chem. 264, 2801–2809 (1989).
Bochkareva, E., Belegu, V., Korolev, S. & Bochkarev, A. Structure of the major single-stranded DNA-binding domain of Replication Protein A suggests a dynamic mechanism for DNA binding. EMBO J. 20, 612–618 (2001).
Theobald, D.L. & Wuttke, D.S. Prediction of multiple tandem OB-fold domains in telomere end-binding proteins Pot1 and Cdc13. Structure 12, 1877–1879 (2004).
Zou, L. & Elledge, S.J. Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science 300, 1542–1548 (2003).
Lisby, M., Barlow, J.H., Burgess, R.C. & Rothstein, R. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004).
Wellinger, R.J., Wolf, A.J. & Zakian, V.A. Saccharomyces telomeres acquire single-strand TG1–3 tails late in S phase. Cell 72, 51–60 (1993).
Wellinger, R.J., Ethier, K., Labrecque, P. & Zakian, V.A. Evidence for a new step in telomere maintenance. Cell 85, 423–433 (1996).
Takata, H., Kanoh, Y., Gunge, N., Shirahige, K. & Matsuura, A. Reciprocal association of the budding yeast ATM-related proteins Tel1 and Mec1 with telomeres in vivo. Mol. Cell 14, 515–522 (2004).
Smith, J., Zou, H. & Rothstein, R. Characterization of genetic interactions with RFA1: the role of RPA in DNA replication and telomere maintenance. Biochimie 82, 71–78 (2000).
Schramke, V. et al. RPA regulates telomerase action by providing Est1p access to chromosome ends. Nat. Genet. 36, 46–54 (2004).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Eddy, S.R. Profile hidden Markov models. Bioinformatics 14, 755–763 (1998).
Poirot, O., O'Toole, E. & Notredame, C. Tcoffee@igs: A web server for computing, evaluating and combining multiple sequence alignments. Nucleic Acids Res. 31, 3503–3506 (2003).
Poirot, O., Suhre, K., Abergel, C., O'Toole, E. & Notredame, C. 3DCoffee@igs: a web server for combining sequences and structures into a multiple sequence alignment. Nucleic Acids Res. 32, W37–W40 (2004).
Do, C.B., Mahabhashyam, M.S., Brudno, M. & Batzoglou, S. ProbCons: Probabilistic consistency-based multiple sequence alignment. Genome Res. 15, 330–340 (2005).
Acknowledgements
We thank the Brill (Rutgers University), Elledge (Harvard Medical School) and Wold (University of Iowa) laboratories for gifts of strains and plasmids, D. Wuttke and M. Wold for scientific conversations and advice, and E. Ford for technical assistance. This research was supported by Department of Defense postdoctoral fellowship DAMD 17-02-1-0276 (to R.B.C.), by grant GM55867 from the US National Institutes of Health and by the Lebensfeld Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Relative binding of Stn1, Ten1 and Stn164-199. (PDF 423 kb)
Supplementary Fig. 2
Yeast two-hybrid analysis with Cdc13, Stn1 and Ten1. (PDF 211 kb)
Supplementary Fig. 3
Comparison of the domain structure of subunits of the RPA and Cdc13-Stn1-Ten1 complexes. (PDF 384 kb)
Supplementary Table 1
List of plasmids used in this work. (PDF 63 kb)
Rights and permissions
About this article
Cite this article
Gao, H., Cervantes, R., Mandell, E. et al. RPA-like proteins mediate yeast telomere function. Nat Struct Mol Biol 14, 208–214 (2007). https://doi.org/10.1038/nsmb1205
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1205
This article is cited by
-
CST–polymerase α-primase solves a second telomere end-replication problem
Nature (2024)
-
Reconstitution of a telomeric replicon organized by CST
Nature (2022)
-
Shaping human telomeres: from shelterin and CST complexes to telomeric chromatin organization
Nature Reviews Molecular Cell Biology (2021)
-
Structural insights into telomere protection and homeostasis regulation by yeast CST complex
Nature Structural & Molecular Biology (2020)
-
Fine tuning the level of the Cdc13 telomere-capping protein for maximal chromosome stability performance
Current Genetics (2019)