Abstract
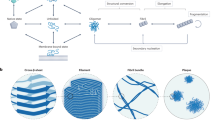
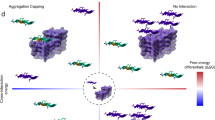
Although most proteins can assemble into amyloid-like fibrils in vitro under extreme conditions, how proteins form amyloid fibrils in vivo remains unresolved. Identifying rare aggregation-prone species under physiologically relevant conditions and defining their structural properties is therefore an important challenge. By solving the folding mechanism of the naturally amyloidogenic protein β-2-microglobulin at pH 7.0 and 37 °C and correlating the concentrations of different species with the rate of fibril elongation, we identify a specific folding intermediate, containing a non-native trans-proline isomer, as the direct precursor of fibril elongation. Structural analysis using NMR shows that this species is highly native-like but contains perturbation of the edge strands that normally protect β-sandwich proteins from self-association. The results demonstrate that aggregation pathways can involve self-assembly of highly native-like folding intermediates, and have implications for the prevention of this, and other, amyloid disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Dobson, C.M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Uversky, V.N. & Fink, A.L. Conformational constraints for amyloid fibrillation: the importance of being unfolded. Biochim. Biophys. Acta 1698, 131–153 (2004).
Colon, W. & Kelly, J.W. Partial denaturation of transthyretin is sufficient for amyloid fibril formation in vitro. Biochemistry 31, 8654–8660 (1992).
Calamai, M., Chiti, F. & Dobson, C.M. Amyloid fibril formation can proceed from different conformations of a partially unfolded protein. Biophys. J. 89, 4201–4210 (2005).
Kelly, J.W. The alternative conformations of amyloidogenic proteins and their multi-step assembly pathways. Curr. Opin. Struct. Biol. 8, 101–106 (1998).
Vendruscolo, M. & Dobson, C.M. Towards complete descriptions of the free-energy landscapes of proteins. Philos. Transact. A Math. Phys. Eng. Sci. 363, 433–450 (2005).
Jahn, T.R. & Radford, S.E. The Yin and Yang of protein folding. FEBS J. 272, 5962–5970 (2005).
Chiti, F. et al. Kinetic partitioning of protein folding and aggregation. Nat. Struct. Biol. 9, 137–143 (2002).
Lashuel, H.A., Lai, Z. & Kelly, J.W. Characterization of the transthyretin acid denaturation pathways by analytical ultracentrifugation: implications for wild-type, V30M, and L55P amyloid fibril formation. Biochemistry 37, 17851–17864 (1998).
Booth, D.R. et al. Instability, unfolding and aggregation of human lysozyme variants underlying amyloid fibrillogenesis. Nature 385, 787–793 (1997).
Liu, K., Cho, H.S., Lashuel, H.A., Kelly, J.W. & Wemmer, D.E. A glimpse of a possible amyloidogenic intermediate of transthyretin. Nat. Struct. Biol. 7, 754–757 (2000).
Khurana, R. et al. Partially folded intermediates as critical precursors of light chain amyloid fibrils and amorphous aggregates. Biochemistry 40, 3525–3535 (2001).
Verdone, G. et al. The solution structure of human β2-microglobulin reveals the prodromes of its amyloid transition. Protein Sci. 11, 487–499 (2002).
Jahn, T.R. & Radford, S.E. β2-microglobulin. in Amyloid Proteins: The Beta Sheet Conformation and Diseases Vol. 2 (ed. Sipe, J.D.) 667–695 (Wiley-VCH, Weinheim, Germany, 2005).
Gejyo, F., Homma, N., Suzuki, Y. & Arakawa, M. Serum levels of β2-microglobulin as a new form of amyloid protein in patients undergoing long-term hemodialysis. N. Engl. J. Med. 314, 585–586 (1986).
McParland, V.J., Kalverda, A.P., Homans, S.W. & Radford, S.E. Structural properties of an amyloid precursor of β2-microglobulin. Nat. Struct. Biol. 9, 326–331 (2002).
Platt, G.W., McParland, V.J., Kalverda, A.P., Homans, S.W. & Radford, S.E. Dynamics in the unfolded state of β2-microglobulin studied by NMR. J. Mol. Biol. 346, 279–294 (2005).
Chiti, F. et al. A partially structured species of β2-microglobulin is significantly populated under physiological conditions and involved in fibrillogenesis. J. Biol. Chem. 276, 46714–46721 (2001).
Chiti, F. et al. Detection of two partially structured species in the folding process of the amyloidogenic protein β2-microglobulin. J. Mol. Biol. 307, 379–391 (2001).
Kameda, A. et al. Nuclear magnetic resonance characterization of the refolding intermediate of β2-microglobulin trapped by non-native prolyl peptide bond. J. Mol. Biol. 348, 383–397 (2005).
Naiki, H. et al. Establishment of a kinetic model of dialysis-related amyloid fibril extension in vitro. Amyloid 4, 223–232 (1997).
Yamamoto, S. et al. Glycosaminoglycans enhance the trifluoroethanol-induced extension of β2-microglobulin-related amyloid fibrils at a neutral pH. J. Am. Soc. Nephrol. 15, 126–133 (2004).
Balbach, J. & Schmid, F.X. Proline isomerisation and its catalysis in protein folding. in Mechanisms of Protein Folding 2nd edn (ed. Pain, R.H.) 212–249 (Oxford University Press, Oxford, 2000).
Benyamini, H., Gunasekaran, K., Wolfson, H. & Nussinov, R. β2-microglobulin amyloidosis: insights from conservation analysis and fibril modelling by protein docking techniques. J. Mol. Biol. 330, 159–174 (2003).
Parker, M.J., Dempsey, C.E., Lorch, M. & Clarke, A.R. Acquisition of native β-strand topology during the rapid collapse phase of protein folding. Biochemistry 36, 13396–13405 (1997).
Sambashivan, S., Liu, Y., Sawaya, M.R., Gingery, M. & Eisenberg, D. Amyloid-like fibrils of ribonuclease A with three-dimensional domain-swapped and native-like structure. Nature 437, 266–269 (2005).
Richardson, J.S. & Richardson, D.C. Natural β-sheet proteins use negative design to avoid edge-to-edge aggregation. Proc. Natl. Acad. Sci. USA 99, 2754–2759 (2002).
Elam, J.S. et al. Amyloid-like filaments and water-filled nanotubes formed by SOD1 mutant proteins linked to familial ALS. Nat. Struct. Biol. 10, 461–467 (2003).
Serag, A.A., Altenbach, C., Gingery, M., Hubbell, W.L. & Yeates, T.O. Arrangement of subunits and ordering of β-strands in an amyloid sheet. Nat. Struct. Biol. 9, 734–739 (2002).
Trinh, C.H., Smith, D.P., Kalverda, A.P., Phillips, S.E. & Radford, S.E. Crystal structure of monomeric human β2-microglobulin reveals clues to its amyloidogenic properties. Proc. Natl. Acad. Sci. USA 99, 9771–9776 (2002).
Jones, S., Smith, D.P. & Radford, S.E. Role of the N and C-terminal strands of β2-microglobulin in amyloid formation at neutral pH. J. Mol. Biol. 330, 935–941 (2003).
Esposito, G. et al. Removal of the N-terminal hexapeptide from human β2-microglobulin facilitates protein aggregation and fibril formation. Protein Sci. 9, 831–845 (2000).
Chien, P., Weissman, J.S. & DePace, A.H. Emerging principles of conformation-based prion inheritance. Annu. Rev. Biochem. 73, 617–656 (2004).
Jimenez, J.L. et al. The protofilament structure of insulin amyloid fibrils. Proc. Natl. Acad. Sci. USA 99, 9196–9201 (2002).
Barral, J.M., Broadley, S.A., Schaffar, G. & Hartl, F.U. Roles of molecular chaperones in protein misfolding diseases. Semin. Cell Dev. Biol. 15, 17–29 (2004).
Mallis, R.J., Brazin, K.N., Fulton, D.B. & Andreotti, A.H. Structural characterization of a proline-driven conformational switch within the Itk SH2 domain. Nat. Struct. Biol. 9, 900–905 (2002).
Eckert, B., Martin, A., Balbach, J. & Schmid, F.X. Prolyl isomerization as a molecular timer in phage infection. Nat. Struct. Mol. Biol. 12, 619–623 (2005).
Lummis, S.C. et al. Cis-trans isomerization at a proline opens the pore of a neurotransmitter-gated ion channel. Nature 438, 248–252 (2005).
Wigley, W.C. et al. A protein sequence that can encode native structure by disfavoring alternate conformations. Nat. Struct. Biol. 9, 381–388 (2002).
Liou, Y.C. et al. Role of the prolyl isomerase Pin1 in protecting against age-dependent neurodegeneration. Nature 424, 556–561 (2003).
Anfinsen, C.B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973).
Kad, N.M., Thomson, N.H., Smith, D.P., Smith, D.A. & Radford, S.E. β2-microglobulin and its deamidated variant, N17D form amyloid fibrils with a range of morphologies in vitro. J. Mol. Biol. 313, 559–571 (2001).
Nilsson, M.R. Techniques to study amyloid fibril formation in vitro. Methods 34, 151–160 (2004).
Myers, S.L. et al. A systematic study of the effect of physiological factors on β2-microglobulin amyloid formation at neutral pH. Biochemistry (in the press).
O'Nuallain, B. & Wetzel, R. Conformational antibodies recognizing a generic amyloid fibril epitope. Proc. Natl. Acad. Sci. USA 99, 1485–1490 (2002).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B.A. & Blevins, R.A. NMRview: a computer program for the visualisation and analysis for NMR data. J. Biol. NMR 4, 603–614 (1994).
Lapidus, L.J., Eaton, W.A. & Hofrichter, J. Measuring the rate of intramolecular contact formation in polypeptides. Proc. Natl. Acad. Sci. USA 97, 7220–7225 (2000).
DeLano, W. The PyMOL Molecular Graphics System. (DeLano Scientific, San Carlos, California, USA, 2002).
Gruebele, M. Protein folding: the free energy surface. Curr. Opin. Struct. Biol. 12, 161–168 (2002).
Acknowledgements
We thank A.P. Kalverda, S.L. Myers, S. Jones and I.J. Morten for their help and support throughout this work, M.B. Pepys, G.A. Tennent, R. Wetzel and J.D. Fryer for kindly providing materials used and C.M. Dobson, F. Chiti and members of S.E.R.'s and S.W.H.'s group for helpful discussions. T.R.J. was supported by the Wellcome Trust, M.J.P. is a Biotechnology and Biological Sciences Research Council David Phillips Fellow and University of Leeds Research Fellow and S.E.R. is a Biotechnology and Biological Sciences Research Council Professorial Fellow.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Double-jump refolding experiment for wild-type β2m. (PDF 133 kb)
Supplementary Fig. 2
Double-jump unfolding kinetics for wild-type β2m. (PDF 333 kb)
Supplementary Fig. 3
Global analysis of the folding and unfolding kinetics of wild-type β2m. (PDF 183 kb)
Supplementary Fig. 4
Global analysis of the folding and unfolding kinetics of P32G β2m. (PDF 345 kb)
Supplementary Table 1
Parameters for wild-type β2m folding kinetics (PDF 65 kb)
Supplementary Table 2
Parameters for P32G β2m folding kinetics (PDF 58 kb)
Supplementary Methods
Methods used for double-jump experiments and kinetic modeling (PDF 97 kb)
Rights and permissions
About this article
Cite this article
Jahn, T., Parker, M., Homans, S. et al. Amyloid formation under physiological conditions proceeds via a native-like folding intermediate. Nat Struct Mol Biol 13, 195–201 (2006). https://doi.org/10.1038/nsmb1058
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1058
This article is cited by
-
Disease-relevant β2-microglobulin variants share a common amyloid fold
Nature Communications (2023)
-
Cryo-EM structure of ex vivo fibrils associated with extreme AA amyloidosis prevalence in a cat shelter
Nature Communications (2022)
-
Partially native intermediates mediate misfolding of SOD1 in single-molecule folding trajectories
Nature Communications (2017)
-
The native state of prion protein (PrP) directly inhibits formation of PrP-amyloid fibrils in vitro
Scientific Reports (2017)
-
Stepwise unfolding of human β2-microglobulin into a disordered amyloidogenic precursor at low pH
European Biophysics Journal (2017)