Abstract
Assembly of fully functional ribosomes is a prerequisite for failsafe translation. This explains why maturing preribosomal subunits have to pass through an array of quality-control checkpoints, including nuclear export, to ensure that only properly assembled ribosomes engage in translation. Despite these safeguards, we found that nuclear pre-60S particles unable to remove a transient structure composed of ITS2 pre-rRNA and associated assembly factors, termed the 'foot', escape to the cytoplasm, where they can join with mature 40S subunits to catalyze protein synthesis. However, cells harboring these abnormal ribosomes show translation defects indicated by the formation of 80S ribosomes poised with pre-60S subunits carrying tRNAs in trapped hybrid states. To overcome this translational stress, the cytoplasmic surveillance machineries RQC and Ski-exosome target these malfunctioning ribosomes. Thus, pre-60S subunits that escape nuclear quality control can enter translation, but are caught by cytoplasmic surveillance mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Woolford, J.L. Jr. & Baserga, S.J. Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195, 643–681 (2013).
de la Cruz, J., Karbstein, K. & Woolford, J.L. Jr. Functions of ribosomal proteins in assembly of eukaryotic ribosomes in vivo. Annu. Rev. Biochem. 84, 93–129 (2015).
Kressler, D., Hurt, E. & Baßler, J. A puzzle of life: crafting ribosomal subunits. Trends Biochem. Sci. 42, 640–654 (2017).
Bradatsch, B. et al. Structure of the pre-60S ribosomal subunit with nuclear export factor Arx1 bound at the exit tunnel. Nat. Mol. Biol. 19, 1234–1241 (2012).
Leidig, C. et al. 60S ribosome biogenesis requires rotation of the 5S ribonucleoprotein particle. Nat. Commun. 5, 3491 (2014).
Wu, S. et al. Diverse roles of assembly factors revealed by structures of late nuclear pre-60S ribosomes. Nature 534, 133–137 (2016).
Sahasranaman, A. et al. Assembly of Saccharomyces cerevisiae 60S ribosomal subunits: role of factors required for 27S pre-rRNA processing. EMBO J. 30, 4020–4032 (2011).
Granneman, S., Petfalski, E. & Tollervey, D. A cluster of ribosome synthesis factors regulate pre-rRNA folding and 5.8S rRNA maturation by the Rat1 exonuclease. EMBO J. 30, 4006–4019 (2011).
Thoms, M. et al. The exosome is recruited to RNA substrates through specific adaptor proteins. Cell 162, 1029–1038 (2015).
Wu, S., Tan, D., Woolford, J.L. Jr., Dong, M.Q. & Gao, N. Atomic modeling of the ITS2 ribosome assembly subcomplex from cryo-EM together with mass spectrometry-identified protein-protein crosslinks 26, 103–112 (2017).
Gasse, L., Flemming, D. & Hurt, E. Coordinated ribosomal ITS2 RNA processing by the Las1 complex integrating endonuclease, polynucleotide kinase, and exonuclease activities. Mol. Cell 60, 808–815 (2015).
Allmang, C. et al. Functions of the exosome in rRNA, snoRNA and snRNA synthesis. EMBO J. 18, 5399–5410 (1999).
Fang, F., Phillips, S. & Butler, J.S. Rat1p and Rai1p function with the nuclear exosome in the processing and degradation of rRNA precursors. RNA 11, 1571–1578 (2005).
Briggs, M.W., Burkard, K.T.D. & Butler, J.S. Rrp6p, the yeast homologue of the human PM-Scl 100-kDa autoantigen, is essential for efficient 5.8 S rRNA 3′ end formation. J. Biol. Chem. 273, 13255–13263 (1998).
Pillon, M.C., Sobhany, M., Borgnia, M.J., Williams, J.G. & Stanley, R.E. Grc3 programs the essential endoribonuclease Las1 for specific RNA cleavage. Proc. Natl. Acad. Sci. USA 114, E5530–E5538 (2017).
Mitchell, P., Petfalski, E. & Tollervey, D. The 3′ end of yeast 5.8S rRNA is generated by an exonuclease processing mechanism. Genes Dev. 10, 502–513 (1996).
de la Cruz, J., Kressler, D., Tollervey, D. & Linder, P. Dob1p (Mtr4p) is a putative ATP-dependent RNA helicase required for the 3′ end formation of 5.8S rRNA in Saccharomyces cerevisiae. EMBO J. 17, 1128–1140 (1998).
Faber, A.W., Vos, H.R., Vos, J.C. & Raué, H.A. 5′-end formation of yeast 5.8SL rRNA is an endonucleolytic event. Biochem. Biophys. Res. Commun. 345, 796–802 (2006).
van Hoof, A., Lennertz, P. & Parker, R. Three conserved members of the RNase D family have unique and overlapping functions in the processing of 5S, 5.8S, U4, U5, RNase MRP and RNase P RNAs in yeast. EMBO J. 19, 1357–1365 (2000).
Thomson, E. & Tollervey, D. The final step in 5.8S rRNA processing is cytoplasmic in Saccharomyces cerevisiae. Mol. Cell. Biol. 30, 976–984 (2010).
Dez, C., Houseley, J. & Tollervey, D. Surveillance of nuclear-restricted pre-ribosomes within a subnucleolar region of Saccharomyces cerevisiae. EMBO J. 25, 1534–1546 (2006).
Houseley, J. & Tollervey, D. Yeast Trf5p is a nuclear poly(A) polymerase. EMBO Rep. 7, 205–211 (2006).
Matsuo, Y. et al. Coupled GTPase and remodelling ATPase activities form a checkpoint for ribosome export. Nature 505, 112–116 (2014).
Sarkar, A., Pech, M., Thoms, M., Beckmann, R. & Hurt, E. Ribosome-stalk biogenesis is coupled with recruitment of nuclear-export factor to the nascent 60S subunit. Nat. Struct. Mol. Biol. 23, 1074–1082 (2016).
Espinar-Marchena, F.J., Babiano, R. & Cruz, J. Placeholder factors in ribosome biogenesis: please, pave my way. Microb. Cell 4, 144–168 (2017).
Strunk, B.S. et al. Ribosome assembly factors prevent premature translation initiation by 40S assembly intermediates. Science 333, 1449–1453 (2011).
Karbstein, K. Quality control mechanisms during ribosome maturation. Trends Cell Biol. 23, 242–250 (2013).
Weis, F. et al. Mechanism of eIF6 release from the nascent 60S ribosomal subunit. Nat. Struct. Mol. Biol. 22, 914–919 (2015).
Ma, C. et al. Structural snapshot of cytoplasmic pre-60S ribosomal particles bound by Nmd3, Lsg1, Tif6 and Reh1. Nat. Struct. Mol. Biol. 24, 214–220 (2017).
Lo, K.Y. et al. Defining the pathway of cytoplasmic maturation of the 60S ribosomal subunit. Mol. Cell 39, 196–208 (2010).
Cole, S.E., LaRiviere, F.J., Merrikh, C.N. & Moore, M.J. A convergence of rRNA and mRNA quality control pathways revealed by mechanistic analysis of nonfunctional rRNA decay. Mol. Cell 34, 440–450 (2009).
LaRiviere, F.J., Cole, S.E., Ferullo, D.J. & Moore, M.J. A late-acting quality control process for mature eukaryotic rRNAs. Mol. Cell 24, 619–626 (2006).
Soudet, J., Gélugne, J.P., Belhabich-Baumas, K., Caizergues-Ferrer, M. & Mougin, A. Immature small ribosomal subunits can engage in translation initiation in Saccharomyces cerevisiae. EMBO J. 29, 80–92 (2010).
Rodríguez-Galán, O., García-Gómez, J.J., Kressler, D. & de la Cruz, J. Immature large ribosomal subunits containing the 7S pre-rRNA can engage in translation in Saccharomyces cerevisiae. RNA Biol. 12, 838–846 (2015).
Brandman, O. et al. A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 151, 1042–1054 (2012).
Anderson, J.S. & Parker, R.P. The 3′ to 5′ degradation of yeast mRNAs is a general mechanism for mRNA turnover that requires the SKI2 DEVH box protein and 3′ to 5′ exonucleases of the exosome complex. EMBO J. 17, 1497–1506 (1998).
van Hoof, A., Frischmeyer, P.A., Dietz, H.C. & Parker, R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295, 2262–2264 (2002).
Hung, N.J. & Johnson, A.W. Nuclear recycling of the pre-60S ribosomal subunit-associated factor Arx1 depends on Rei1 in Saccharomyces cerevisiae. Mol. Cell. Biol. 26, 3718–3727 (2006).
Lebreton, A. et al. A functional network involved in the recycling of nucleocytoplasmic pre-60S factors. J. Cell Biol. 173, 349–360 (2006).
Saveanu, C. et al. Sequential protein association with nascent 60S ribosomal particles. Mol. Cell. Biol. 23, 4449–4460 (2003).
Ban, N. et al. A new system for naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169 (2014).
Shen, P.S. et al. Protein synthesis. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains. Science 347, 75–78 (2015).
Lyumkis, D. et al. Structural basis for translational surveillance by the large ribosomal subunit-associated protein quality control complex. Proc. Natl. Acad. Sci. USA 111, 15981–15986 (2014).
Bengtson, M.H. & Joazeiro, C.A. Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 467, 470–473 (2010).
Defenouillère, Q. et al. Cdc48-associated complex bound to 60S particles is required for the clearance of aberrant translation products. Proc. Natl. Acad. Sci. USA 110, 5046–5051 (2013).
Osuna, B.A., Howard, C.J., Kc, S., Frost, A. & Weinberg, D.E. In vitro analysis of RQC activities provides insights into the mechanism and function of CAT tailing. eLife 6, e27949 (2017).
Remacha, M. et al. Proteins P1, P2, and P0, components of the eukaryotic ribosome stalk. New structural and functional aspects. Biochem. Cell Biol. 73, 959–968 (1995).
Behrmann, E. et al. Structural snapshots of actively translating human ribosomes. Cell 161, 845–857 (2015).
Brandman, O. & Hegde, R.S. Ribosome-associated protein quality control. Nat. Struct. Mol. Biol. 23, 7–15 (2016).
Schmidt, C. et al. The cryo-EM structure of a ribosome-Ski2-Ski3-Ski8 helicase complex. Science 354, 1431–1433 (2016).
Inada, T. The ribosome as a platform for mRNA and nascent polypeptide quality control. Trends Biochem. Sci. 42, 5–15 (2017).
Pestov, D.G. & Shcherbik, N. Rapid cytoplasmic turnover of yeast ribosomes in response to rapamycin inhibition of TOR. Mol. Cell. Biol. 32, 2135–2144 (2012).
Medina, D.A. et al. Cytoplasmic 5′-3′ exonuclease Xrn1p is also a genome-wide transcription factor in yeast. Front. Genet. 5, 1 (2014).
Mitchell, P. & Tollervey, D. mRNA stability in eukaryotes. Curr. Opin. Genet. Dev. 10, 193–198 (2000).
Kobayashi, K. et al. Structural basis for mRNA surveillance by archaeal Pelota and GTP-bound EF1α complex. Proc. Natl. Acad. Sci. USA 107, 17575–17579 (2010).
Shoemaker, C.J., Eyler, D.E. & Green, R. Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330, 369–372 (2010).
Matsuo, Y. et al. Ubiquitination of stalled ribosome triggers ribosome-associated quality control. Nat. Commun. 8, 159 (2017).
Sitron, C.S., Park, J.H. & Brandman, O. Asc1, Hel2, and Slh1 couple translation arrest to nascent chain degradation. RNA 23, 798–810 (2017).
Kuroha, K. et al. Receptor for activated C kinase 1 stimulates nascent polypeptide-dependent translation arrest. EMBO Rep. 11, 956–961 (2010).
Tsai, A. et al. The impact of aminoglycosides on the dynamics of translation elongation. Cell Rep. 3, 497–508 (2013).
Garreau de Loubresse, N. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014).
Sundaramoorthy, E. et al. ZNF598 and RACK1 regulate mammalian ribosome-associated quality control function by mediating regulatory 40S ribosomal ubiquitylation. Mol. Cell 65, 751–760.e4 (2017).
Juszkiewicz, S. & Hegde, R.S. Initiation of quality control during poly(A) translation requires site-specific ribosome ubiquitination. Mol. Cell 65, 743–750.e4 (2017).
Garzia, A. et al. The E3 ubiquitin ligase and RNA-binding protein ZNF598 orchestrates ribosome quality control of premature polyadenylated mRNAs. Nat. Commun. 8, 16056 (2017).
Barrio-Garcia, C. et al. Architecture of the Rix1-Rea1 checkpoint machinery during pre-60S-ribosome remodeling. Nat. Struct. Mol. Biol. 23, 37–44 (2016).
Lafontaine, D.L.A. A 'garbage can' for ribosomes: how eukaryotes degrade their ribosomes. Trends Biochem. Sci. 35, 267–277 (2010).
Choe, Y.J. et al. Failure of RQC machinery causes protein aggregation and proteotoxic stress. Nature 531, 191–195 (2016).
Letzring, D.P., Wolf, A.S., Brule, C.E. & Grayhack, E.J. Translation of CGA codon repeats in yeast involves quality control components and ribosomal protein L1. RNA 19, 1208–1217 (2013).
Ikeuchi, K. & Inada, T. Ribosome-associated Asc1/RACK1 is required for endonucleolytic cleavage induced by stalled ribosome at the 3′ end of nonstop mRNA. Sci. Rep. 6, 28234 (2016).
Ossareh-Nazari, B. et al. Ubiquitylation by the Ltn1 E3 ligase protects 60S ribosomes from starvation-induced selective autophagy. J. Cell Biol. 204, 909–917 (2014).
Myasnikov, A.G. et al. The molecular structure of the left-handed supra-molecular helix of eukaryotic polyribosomes. Nat. Commun. 5, 5294 (2014).
Armache, J.P. et al. Cryo-EM structure and rRNA model of a translating eukaryotic 80S ribosome at 5.5-A resolution. Proc. Natl. Acad. Sci. USA 107, 19748–19753 (2010).
Beckmann, R. et al. Architecture of the protein-conducting channel associated with the translating 80S ribosome. Cell 107, 361–372 (2001).
Sasaki, M. et al. Regulation of the MDM2-P53 pathway and tumor growth by PICT1 via nucleolar RPL11. Nat. Med. 17, 944–951 (2011).
Butterfield, R.J. et al. Congenital lethal motor neuron disease with a novel defect in ribosome biogenesis. Neurology 82, 1322–1330 (2014).
Schmidt, C. et al. Structure of the hypusinylated eukaryotic translation factor eIF-5A bound to the ribosome. Nucleic Acids Res. 44, 1944–1951 (2016).
Longtine, M.S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Thomas, B.J. & Rothstein, R. Elevated recombination rates in transcriptionally active DNA. Cell 56, 619–630 (1989).
Bradatsch, B. et al. Arx1 functions as an unorthodox nuclear export receptor for the 60S preribosomal subunit. Mol. Cell 27, 767–779 (2007).
Simos, G. et al. The yeast protein Arc1p binds to tRNA and functions as a cofactor for the methionyl- and glutamyl-tRNA synthetases. EMBO J. 15, 5437–5448 (1996).
Vilardell, J. & Warner, J.R. Ribosomal protein L32 of Saccharomyces cerevisiae influences both the splicing of its own transcript and the processing of rRNA. Mol. Cell. Biol. 17, 1959–1965 (1997).
Du, Y.C. & Stillman, B. Yph1p, an ORC-interacting protein: potential links between cell proliferation control, DNA replication, and ribosome biogenesis. Cell 109, 835–848 (2002).
Jäger, S., Strayle, J., Heinemeyer, W. & Wolf, D.H. Cic1, an adaptor protein specifically linking the 26S proteasome to its substrate, the SCF component Cdc4. EMBO J. 20, 4423–4431 (2001).
Vilella, M.D., Remacha, M., Ortiz, B.L., Mendez, E. & Ballesta, J.P. Characterization of the yeast acidic ribosomal phosphoproteins using monoclonal antibodies. Proteins L44/L45 and L44′ have different functional roles. Eur. J. Biochem. 196, 407–414 (1991).
Tron, T., Yang, M., Dick, F.A., Schmitt, M.E. & Trumpower, B.L. QSR1, an essential yeast gene with a genetic relationship to a subunit of the mitochondrial cytochrome bc1 complex, is homologous to a gene implicated in eukaryotic cell differentiation. J. Biol. Chem. 270, 9961–9970 (1995).
Frey, S., Pool, M. & Seedorf, M. Scp160p, an RNA-binding, polysome-associated protein, localizes to the endoplasmic reticulum of Saccharomyces cerevisiae in a microtubule-dependent manner. J. Biol. Chem. 276, 15905–15912 (2001).
Dieci, G., Bottarelli, L. & Ottonello, S. A general procedure for the production of antibody reagents against eukaryotic ribosomal proteins. Protein Pept. Lett. 12, 555–560 (2005).
Ho, J.H., Kallstrom, G. & Johnson, A.W. Nmd3p is a Crm1p-dependent adapter protein for nuclear export of the large ribosomal subunit. J. Cell Biol. 151, 1057–1066 (2000).
Kemmler, S., Occhipinti, L., Veisu, M. & Panse, V.G. Yvh1 is required for a late maturation step in the 60S biogenesis pathway. J. Cell Biol. 186, 863–880 (2009).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Tollervey, D. & Mattaj, I.W. Fungal small nuclear ribonucleoproteins share properties with plant and vertebrate U-snRNPs. EMBO J. 6, 469–476 (1987).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Liu, X. & Wang, H.W. Single particle electron microscopy reconstruction of the exosome complex using the random conical tilt method. J. Vis. Exp. 49, 2574 (2011).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Chen, J.Z. & Grigorieff, N. SIGNATURE: a single-particle selection system for molecular electron microscopy. J. Struct. Biol. 157, 168–173 (2007).
Scheres, S.H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Grigorieff, N. FREALIGN: high-resolution refinement of single particle structures. J. Struct. Biol. 157, 117–125 (2007).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Rosenthal, P.B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F.J. & Tagare, H.D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We would like to thank O. Berninghausen and C. Ungewickell for cryo-EM data collection and vitrifying samples, as well as P. Ihrig and J. Reichert (BZH) for performing all MS analyses. We are grateful to A.W. Johnson (University of Texas at Austin), B. Stillman (Cold Spring Harbor Laboratory), S. Ottonello (Università di Parma), J.R. Warner (Albert Einstein College of Medicine), J.P. Ballesta (Centro de Biologia Molecular Severo Ochoa), M. Fromont-Racine (Institut Pasteur), M. Seedorf (ZMBH University of Heidelberg), V. Panse (University of Zurich), B.L. Trumpower (Dartmouth Medical School) and E. Tosta for their gift of antibodies. We also thank C. Joazeiro (ZMBH University of Heidelberg) for critically reading the manuscript. E.H. is a recipient of grants from the Deutsche Forschungsgemeinschaft DFG (HU363/10-5, HU363/12-1).
Author information
Authors and Affiliations
Contributions
M.T., A.S., C.B.-G., R.B. and E.H. designed the study and analyzed the data. A.S. and M.T. performed all experiments except cryo-EM, which was done by C.B.-G. and R.B. E.T. performed the northern blot presented in Supplementary Figure 5c. D.F. performed the negative-stain EM. A.S., M.T., C.B.-G., E.T., R.B. and E.H. interpreted the results and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Impaired ITS2 processing results in mislocalization of foot-factors to the cytoplasm.
(a) Schematic representation of ITS2 processing. The endonuclease Las1 initiates ITS2 processing by cleaving at site C2 (of the pre-rRNA) that leads to the separation of the 7S and the 26S rRNA pathways. The 7S pathway proceeds by Nop53 mediated recruitment of Mtr4 and the exosome in the nucleus, which finally leads to mature 5.8S (in the cytoplasm) via different intermediates (indicated in the scheme). (b) Western blot analysis of whole cell lysates from the Las1-Aid strain (upper panel) and the Las1-Aid Nsa3-Flag Rqc2-TAP strain (lower panel), under conditions of Las1 expression (- Auxin) and Las1 depletion (+ Auxin). Samples were probed with the indicated antibodies. The results show that the levels of foot factors remain unperturbed upon Las1 depletion. The arrow points to the western blot signal of the HA tagged Las1-Aid and the asterisk indicates a background signal from the HA antibody (lower panel). (c) The subcellular localization of the GFP-tagged Nop7, Nsa3, Nug2 and Rpf2 from nop53Δ yeast cells under the expression of plasmid borne NOP53 wild-type, nop53 5×Ala and nop53 D64R mutant alleles. Cells were analyzed by fluorescence and DIC microscopy. Scale bar, 5 μm. (d) Whole cell lysates derived from nop53Δ yeast cells harboring the N-terminal TAP-Flag tagged NOP53 wild-type or the nop53 5×Ala allele, where analyzed by sucrose gradient centrifugation. The 40S, 60S, 80S and polysome fractions are indicated and halfmers are marked by arrowheads. The sucrose gradient fractions were analyzed by western blot analysis and probed with anti-ProtA (Nop53) and anti-L3 antibodies.
Supplementary Figure 2 Negative-stain EM of the Rqc2-TAP Nsa3-Flag particle under Las1 depletion conditions in comparison to mature 60S particles.
(a) Coomassie stained SDS-PAGE of the split purified Rqc2-TAP Nsa3-Flag particle obtained after depletion of Las1-Aid (+ Auxin, 2 h), which was used for subsequent negative stain EM analysis. (b) Selected 2D class averages of the Rqc2-TAP Nsa3-Flag particle purified from Las1 depleted cells. The class average shown in Fig. 3d is marked with a red box. Arrowheads point to a diffuse density, which might correspond to Ltn1. (c) Selected 2D class averages of an L24A-FTpA particle, which is a mixture of mature 60S subunits and 80S ribosomes. The population of the mature 60S subunit particle (marked with a red box) in the total dataset is ~3%.
Supplementary Figure 3 Cryo-EM sorting scheme of the TAP-Flag-Nop53 particle obtained after Las1 depletion.
The particles obtained from the TAP-Flag-Nop53 purification after Las1 depletion were classified using iterative multi-reference projection alignment. The data was sorted into 5 different classes, all of them displaying density for the foot structure. Class 2, 3 and 5 showed a strong density for the 60S subunit, but only very noisy density in the 40S region. Classes 1 and 4 represented 80S ribosomes in the absence or presence of tRNAs, respectively. These two classes were taken together for a new round of classification that gave the final classes containing empty 80S and 80S with tRNAs. Then, the final classes were refined using only the particles with higher cross correlation. The final structure is highlighted in green. Other classification rounds were performed with different approaches in order to prove that particles were fully classified (see Online Methods). In the final classes the density for the 40S subunit is displayed in yellow, the A/P tRNA in green, the foot structure in orange and eIF6 in red.
Supplementary Figure 4 Resolution of the foot containing 80S particle with comparison to peptidyl-tRNA-60S subunits bound to Rqc2 and Ltn1.
(a) Overall resolution measured at the 0.143 cut-off of the FSC from two independent datasets. The final subpopulation was refined to a resolution of 7.3 Å. (b) Nop53 Las1-depleted 80S map colored according to its local resolution. (c) Comparison between the cryo-EM structure of peptidyl-tRNA-60S ribosomes bound to Rqc2 and Ltn1 (Shen, P.S. et al., Science. 347, 75-8, 2015) (EMDB-6170; overall structure in light blue, Ltn1 and Rqc2 are respectively highlighted in tan and purple) and the reconstruction of the foot-containing 80S particle (gray, foot structure shown in orange). For easier visualization the 40S subunit of the latter reconstruction was omitted.
Supplementary Figure 5 The nop53 5×Ala mutant is genetically and functionally linked to cytoplasmic surveillance factors.
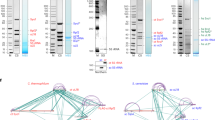
(a) Growth analysis comparing a wild-type strain and the indicated deletions strains under galactose overexpression of plasmid-based NOP53 wild-type or nop53 5×Ala mutant alleles (under control of the GAL1-10 promoter). An empty vector served as control. Cells were spotted in 10-fold serial dilutions on SDC-Leu (glucose) and SGC-Leu (galactose) medium and cell growth at 30°C was monitored after 2 and 3 days, respectively. (b) Polysome gradient analysis of whole cell lysates derived from a Nsa3-Flag and Nsa3-Flag ski2Δ strain after overexpression (for 8 h) of the dominant negative GAL::nop53 5×Ala mutant. The fractions containing 40S, 60S, 80S and polysomes are indicated. The sucrose gradient fractions were analyzed by western blot analysis and probed with the indicated antibodies. (c) Analysis of pre-rRNA and rRNAs extracted from the indicated yeast strains. Galactose induction was performed for 8h, following which RNAs were extracted and analyzed by Northern blotting. Oligonucleotide probes used for Northern analysis: precursors to 5.8S rRNA (top panel) were detected with the 020 oligo probe (TGAGAAGGAAATGACGCT) and the 5S rRNA was detected with the 041 oligo probe (CTACTCGGTCAGGCTC). Values shown indicate the relative abundance of 7S pre-rRNA compared to the wild-type lane (1), when normalized to 5S rRNA levels. Asterisk marks the abnormal processing intermediate between 7S and 5.8S+30.
Supplementary Figure 6 Analysis of the foot-containing 80S structure.
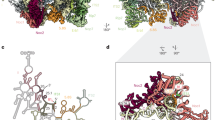
(a) Magnification of the foot structure (PDB ID: 3JCT) fitted into the density of the 80S ribosome presented in this study. Factors belonging to the foot structure as well as ITS2 are highlighted in different colors and labeled accordingly. The structure of the 60S subunit (PDB ID: 5TGM) is fitted in order to display the main interactions between the foot structure and the 60S subunit. R-proteins interacting with the foot L8 (eL8), L25 (uL23) and L27 (eL27) are highlighted in green, yellow and red respectively (Wu, S. et al., Nature. 534, 133-7, 2016). 25S and 5.8S rRNAs are shown in gray and the rest of the r-proteins in beige. (b) The fit of the foot containing 80S ribosome into the density of actively translating polysomes (EMDB-2790) demonstrates that the foot (orange) would clash with the small subunit of the next ribosome (light blue). (c) The transition between the two conformations of ES27 (displayed in green and blue, PDB ID: 3IZF and 3IZD) may be obstructed by the presence of the foot.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–2 (PDF 1703 kb)
Supplementary Data Set 1
Uncropped gels. (PDF 14677 kb)
Rights and permissions
About this article
Cite this article
Sarkar, A., Thoms, M., Barrio-Garcia, C. et al. Preribosomes escaping from the nucleus are caught during translation by cytoplasmic quality control. Nat Struct Mol Biol 24, 1107–1115 (2017). https://doi.org/10.1038/nsmb.3495
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3495