Abstract
Host and virus interactions occurring at the post-transcriptional level are critical for infection but remain poorly understood. Here, we performed comprehensive transcriptome-wide analyses revealing that human cytomegalovirus (HCMV) infection results in widespread alternative splicing (AS), shortening of 3′ untranslated regions (3′ UTRs) and lengthening of poly(A)-tails in host gene transcripts. We found that the host RNA-binding protein CPEB1 was highly induced after infection, and ectopic expression of CPEB1 in noninfected cells recapitulated infection-related post-transcriptional changes. CPEB1 was also required for poly(A)-tail lengthening of viral RNAs important for productive infection. Strikingly, depletion of CPEB1 reversed infection-related cytopathology and post-transcriptional changes, and decreased productive HCMV titers. Host RNA processing was also altered in herpes simplex virus-2 (HSV-2)-infected cells, thereby indicating that this phenomenon might be a common occurrence during herpesvirus infections. We anticipate that our work may serve as a starting point for therapeutic targeting of host RNA-binding proteins in herpesvirus infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
NCBI Reference Sequence
References
Staras, S.A. et al. Seroprevalence of cytomegalovirus infection in the United States, 1988–1994. Clin. Infect. Dis. 43, 1143–1151 (2006).
Mocarski, E.S. Betaherpes viral genes and their functions. in Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis (eds. Arvin, A. et al.), 204–230 (Cambridge University Press, 2007).
Fortunato, E.A., McElroy, A.K., Sanchez, I. & Spector, D.H. Exploitation of cellular signaling and regulatory pathways by human cytomegalovirus. Trends Microbiol. 8, 111–119 (2000).
Mocarski, E.S., Shenk, T. & Pass, R.F. Cytomegaloviruses. in Fields Virology 5th edn, Vol. 2 (eds. Knipe, D.M. & Howley, P.M.) 2701–2772 (Lippincott Williams & Wilkins, 2007).
Campbell, L.A. & Rosenfeld, M.E. Infection and atherosclerosis development. Arch. Med. Res. 46, 339–350 (2015).
DuRose, J.B., Li, J., Chien, S. & Spector, D.H. Infection of vascular endothelial cells with human cytomegalovirus under fluid shear stress reveals preferential entry and spread of virus in flow conditions simulating atheroprone regions of the artery. J. Virol. 86, 13745–13755 (2012).
Schuessler, A., Walker, D.G. & Khanna, R. Cytomegalovirus as a novel target for immunotherapy of glioblastoma multiforme. Front. Oncol. 4, 275 (2014).
Horváth, R. et al. The possible role of human cytomegalovirus (HCMV) in the origin of atherosclerosis. J. Clin. Virol. 16, 17–24 (2000).
Fenwick, M.L. & Walker, M.J. Suppression of the synthesis of cellular macromolecules by herpes simplex virus. J. Gen. Virol. 41, 37–51 (1978).
Sydiskis, R.J. & Roizman, B. Polysomes and protein synthesis in cells infected with a DNA virus. Science 153, 76–78 (1966).
Kwong, A.D. & Frenkel, N. Herpes simplex virus-infected cells contain a function(s) that destabilizes both host and viral mRNAs. Proc. Natl. Acad. Sci. USA 84, 1926–1930 (1987).
Hertel, L. & Mocarski, E.S. Global analysis of host cell gene expression late during cytomegalovirus infection reveals extensive dysregulation of cell cycle gene expression and induction of pseudomitosis independent of US28 function. J. Virol. 78, 11988–12011 (2004).
Gatherer, D. et al. High-resolution human cytomegalovirus transcriptome. Proc. Natl. Acad. Sci. USA 108, 19755–19760 (2011).
Stern-Ginossar, N. et al. Decoding human cytomegalovirus. Science 338, 1088–1093 (2012).
Poulos, M.G., Batra, R., Charizanis, K. & Swanson, M.S. Developments in RNA splicing and disease. Cold Spring Harb. Perspect. Biol. 3, a000778 (2011).
Ulitsky, I. et al. Extensive alternative polyadenylation during zebrafish development. Genome Res. 22, 2054–2066 (2012).
Batra, R. et al. Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol. Cell 56, 311–322 (2014).
Scotti, M.M. & Swanson, M.S. RNA mis-splicing in disease. Nat. Rev. Genet. 17, 19–32 (2016).
Charizanis, K. et al. Muscleblind-like 2-mediated alternative splicing in the developing brain and dysregulation in myotonic dystrophy. Neuron 75, 437–450 (2012).
Jenal, M. et al. The poly(A)-binding protein nuclear 1 suppresses alternative cleavage and polyadenylation sites. Cell 149, 538–553 (2012).
de Klerk, E. et al. Poly(A) binding protein nuclear 1 levels affect alternative polyadenylation. Nucleic Acids Res. 40, 9089–9101 (2012).
Richter, J.D. CPEB: a life in translation. Trends Biochem. Sci. 32, 279–285 (2007).
Scorilas, A. Polyadenylate polymerase (PAP) and 3′ end pre-mRNA processing: function, assays, and association with disease. Crit. Rev. Clin. Lab. Sci. 39, 193–224 (2002).
Subtelny, A.O., Eichhorn, S.W., Chen, G.R., Sive, H. & Bartel, D.P. Poly(A)-tail profiling reveals an embryonic switch in translational control. Nature 508, 66–71 (2014).
Chang, H., Lim, J., Ha, M. & Kim, V.N. TAIL-seq: genome-wide determination of poly(A) tail length and 3′ end modifications. Mol. Cell 53, 1044–1052 (2014).
Licatalosi, D.D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Katz, Y., Wang, E.T., Airoldi, E.M. & Burge, C.B. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nat. Methods 7, 1009–1015 (2010).
Huelga, S.C. et al. Integrative genome-wide analysis reveals cooperative regulation of alternative splicing by hnRNP proteins. Cell Rep. 1, 167–178 (2012).
Gehman, L.T. et al. The splicing regulator Rbfox1 (A2BP1) controls neuronal excitation in the mammalian brain. Nat. Genet. 43, 706–711 (2011).
Belzile, J.P., Stark, T.J., Yeo, G.W. & Spector, D.H. Human cytomegalovirus infection of human embryonic stem cell-derived primitive neural stem cells is restricted at several steps but leads to the persistence of viral DNA. J. Virol. 88, 4021–4039 (2014).
Luo, W. et al. The conserved intronic cleavage and polyadenylation site of CstF-77 gene imparts control of 3′ end processing activity through feedback autoregulation and by U1 snRNP. PLoS Genet. 9, e1003613 (2013).
Bava, F.A. et al. CPEB1 coordinates alternative 3′-UTR formation with translational regulation. Nature 495, 121–125 (2013).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Kim, Y. et al. Human cytomegalovirus UL18 utilizes US6 for evading the NK and T-cell responses. PLoS Pathog. 4, e1000123 (2008).
Batra, R., Manchanda, M. & Swanson, M.S. Global insights into alternative polyadenylation regulation. RNA Biol. 12, 597–602 (2015).
Mayr, C. & Bartel, D.P. Widespread shortening of 3′UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 138, 673–684 (2009).
Sandberg, R., Neilson, J.R., Sarma, A., Sharp, P.A. & Burge, C.B. Proliferating cells express mRNAs with shortened 3′ untranslated regions and fewer microRNA target sites. Science 320, 1643–1647 (2008).
Homa, N.J. et al. Epstein-Barr virus induces global changes in cellular mRNA isoform usage that are important for the maintenance of latency. J. Virol. 87, 12291–12301 (2013).
Weill, L., Belloc, E., Bava, F.A. & Méndez, R. Translational control by changes in poly(A) tail length: recycling mRNAs. Nat. Struct. Mol. Biol. 19, 577–585 (2012).
Lin, C.L., Evans, V., Shen, S., Xing, Y. & Richter, J.D. The nuclear experience of CPEB: implications for RNA processing and translational control. RNA 16, 338–348 (2010).
Mendez, R. et al. Phosphorylation of CPE binding factor by Eg2 regulates translation of c-mos mRNA. Nature 404, 302–307 (2000).
Mendez, R. & Richter, J.D. Translational control by CPEB: a means to the end. Nat. Rev. Mol. Cell Biol. 2, 521–529 (2001).
Fox, C.A., Sheets, M.D. & Wickens, M.P. Poly(A) addition during maturation of frog oocytes: distinct nuclear and cytoplasmic activities and regulation by the sequence UUUUUAU. Genes Dev. 3, 2151–2162 (1989).
Cepeda, V., Esteban, M. & Fraile-Ramos, A. Human cytomegalovirus final envelopment on membranes containing both trans-Golgi network and endosomal markers. Cell. Microbiol. 12, 386–404 (2010).
Walker, J.D., Maier, C.L. & Pober, J.S. Cytomegalovirus-infected human endothelial cells can stimulate allogeneic CD4+ memory T cells by releasing antigenic exosomes. J. Immunol. 182, 1548–1559 (2009).
Wurdinger, T. et al. Extracellular vesicles and their convergence with viral pathways. Adv. Virol. 2012, 767694 (2012).
Plazolles, N. et al. Pivotal advance: the promotion of soluble DC-SIGN release by inflammatory signals and its enhancement of cytomegalovirus-mediated cis-infection of myeloid dendritic cells. J. Leukoc. Biol. 89, 329–342 (2011).
Schorey, J.S., Cheng, Y., Singh, P.P. & Smith, V.L. Exosomes and other extracellular vesicles in host-pathogen interactions. EMBO Rep. 16, 24–43 (2015).
Burns, D.M. & Richter, J.D. CPEB regulation of human cellular senescence, energy metabolism, and p53 mRNA translation. Genes Dev. 22, 3449–3460 (2008).
Groppo, R. & Richter, J.D. CPEB control of NF-κB nuclear localization and interleukin-6 production mediates cellular senescence. Mol. Cell. Biol. 31, 2707–2714 (2011).
Lee, Y.J. & Glaunsinger, B.A. Aberrant herpesvirus-induced polyadenylation correlates with cellular messenger RNA destruction. PLoS Biol. 7, e1000107 (2009).
Fraile-Ramos, A., Cepeda, V., Elstak, E. & van der Sluijs, P. Rab27a is required for human cytomegalovirus assembly. PLoS One 5, e15318 (2010).
Silva, M.C., Yu, Q.C., Enquist, L. & Shenk, T. Human cytomegalovirus UL99-encoded pp28 is required for the cytoplasmic envelopment of tegument-associated capsids. J. Virol. 77, 10594–10605 (2003).
Sanchez, V., Sztul, E. & Britt, W.J. Human cytomegalovirus pp28 (UL99) localizes to a cytoplasmic compartment which overlaps the endoplasmic reticulum-golgi-intermediate compartment. J. Virol. 74, 3842–3851 (2000).
Tomtishen, J.P. III. Human cytomegalovirus tegument proteins (pp65, pp71, pp150, pp28). Virol. J. 9, 22 (2012).
Liu, S.T. et al. Synaptic vesicle-like lipidome of human cytomegalovirus virions reveals a role for SNARE machinery in virion egress. Proc. Natl. Acad. Sci. USA 108, 12869–12874 (2011).
Albrecht, T., Cavallo, T., Cole, N.L. & Graves, K. Cytomegalovirus: development and progression of cytopathic effects in human cell culture. Lab. Invest. 42, 1–7 (1980).
Cavallo, T., Graves, K., Cole, N.L. & Albrecht, T. Cytomegalovirus: an ultrastructural study of the morphogenesis of nuclear inclusions in human cell culture. J. Gen. Virol. 56, 97–104 (1981).
Noraz, N., Gozlan, J., Corbeil, J., Brunner, T. & Spector, S.A. HIV-induced apoptosis of activated primary CD4+ T lymphocytes is not mediated by Fas-Fas ligand. AIDS 11, 1671–1680 (1997).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 37, e123 (2009).
Wu, J., Anczuków, O., Krainer, A.R., Zhang, M.Q. & Zhang, C. OLego: fast and sensitive mapping of spliced mRNA-Seq reads using small seeds. Nucleic Acids Res. 41, 5149–5163 (2013).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Acknowledgements
The authors thank members of the laboratories of G.W.Y. and D.H.S. for critical reading of the manuscript and extend particular acknowledgment to M.T. Lovci (UCSD) for initial assistance with bioinformatics. We thank H. Chang (laboratory of N. Kim, Seoul National University) for help with the Tailseeker algorithm, and S. Azoubel Lima and A. Pasquinelli (UCSD) for sharing their modified TAIL-seq protocol before publication. We especially thank C.S. Morello (UCSD) for assistance with HSV-2 infections, and the laboratory of S.A. Spector (UCSD) for HIV-1-infected materials. This work was partially supported by grants from the National Institutes of Health (HG004659, HG007005 and NS075449 to G.W.Y.) and from the California Institute of Regenerative Medicine (RB3-05219 to G.W.Y. and D.H.S.). T.J.S. and E.C.W. are supported in part by the University of California, San Diego, Genetics Training Program through an institutional training grant from the National Institute of General Medical Sciences, T32 GM008666. E.C.W. is supported as an NSF Graduate Research Fellow. R.B. is supported as a Myotonic Dystrophy Foundation postdoctoral fellow. G.W.Y. is supported as an Alfred P. Sloan Research Fellow.
Author information
Authors and Affiliations
Contributions
R.B., T.J.S., D.H.S. and G.W.Y. designed the study and wrote the manuscript; A.E.C. and J.-P.B. maintained HCMV viral stocks, measured titers and performed HCMV infections; R.B., T.J.S. and A.E.C. performed siRNA treatments. T.J.S., A.E.C. and R.B. performed western blots. R.B. performed immunofluorescence and microscopy. T.J.S. and T.J.B. made lentivirus preparations. R.B., T.J.S. and E.C.W. made RNA-seq libraries. R.B., T.J.S., S.C.H., B.A.Y. and S.S. performed bioinformatics analysis. R.B. and T.J.S. performed RT–PCR splicing and APA assays. H.H. and R.B. prepared TAIL-seq libraries, as overseen by S.A. C.G.-B. performed microarray hybridizations. S.C.H., J.P.D. and M.A. performed microarray analysis. R.B., T.J.S. and B.T.R. performed RT–PCR splicing and APA assays. F.R. provided the antisense oligonucleotides.
Corresponding author
Ethics declarations
Competing interests
F.R. is a paid employee of Ionis Pharmaceuticals, which provided ASO reagents.
Integrated supplementary information
Supplementary Figure 1 Viral gene expression differs among cell types.
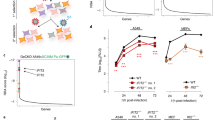
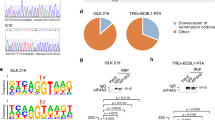
(a) The percent of mapped sequencing reads between the human and viral genomes for HFFs, ECs and NPCs at 48 and 96 h post HCMV TB40E-infection (hpi) from RNA-seq data (n=1 per condition). (b) Pearson correlation matrix of HCMV gene expression values in HFFs, ECs, and NPCs at 48 and 96 hpi. (c) Scatter plot comparisons of HCMV (TB40E) gene levels (Log2 RPKM). Significantly under-represented genes in NPCs at 96 hpi are highlighted in green. (d) Comparison of the highest fold-change differences between 96 hpi NPCs and 96 hpi TB40E HFFs and ECs (genes labeled green in c). Associated temporal gene classes are indicated. D.E. is Delayed Early.
Supplementary Figure 2 Host alternative polyadenylation changes occur during herpesvirus infections.
(a) RNA-seq coverage of MARCH6 3′UTR in HCMV infected NPCs (96 hpi) and HFFs (96 hpi). Canonical predicted CPEB1 recognition sites (CPE; specifically, (U)UUUUAU or UUUUAAU) are indicated by triangles below the 3’UTR. (b) qRT-PCR validation of decreased distal 3′UTR usage in MARCH6 mRNA transcript in HCMV infection (96 hpi, mean +/- standard deviation; n=3). (c) Global distribution of changes in 3′UTR length among shortening events during HCMV infection in three cell types. (d) Bar plot showing altered 3′UTR isoform usage determined by the MISO algorithm 8 hours post HSV-2 infection in HFFs and HIV-1 infected T-cells (left). Venn diagram showing overlap in APA events in HCMV and HSV-2 infections (right). (e) RNA-seq coverage of PCGF3 3′UTR in HSV-2 infected HFFs (8 hpi). (f) qPCR validation of 3′UTR shortening in PCGF3 and ANKH. Error bars = mean +/- standard deviation; n = 3 replicates. (g) RNA-seq coverage of PICALM showing alternatively spliced exon (exclusion) in HSV-2 infection 8 hpi.
Supplementary Figure 3 HCMV infection causes host gene-expression changes.
(a) Heatmap of host gene expression changes (all genes) in HFFs, ECs, and NPCs at 48 and 96 hpi with HCMV TB40E. (b) Venn diagram showing dynamics of host gene expression changes from 48 to 96 hpi with HCMV TB40E. The grey circles are the host genes with altered expression between mock vs. HCMV at 48hpi and black circles represent genes altered at 96 hpi in respective cell types. (c) Venn diagrams of differentially expressed host genes in HCMV TB40E infected HFFs (black), ECs (green), and NPCs (red) at 48 and 96 hpi. (d) qRT-PCR showing CPEB1 upregulation in NPC cells (differentiated from HUES9 and H9 human embryonic stem cells) at 96 hpi with HCMV TB40E. Error bars are mean +/- STD; n=3. (e) Immunoblot showing CPEB1 upregulation in NPCs (from H9) 96 hpi with HCMV TB40E. (f) Hierarchical clustering of log2 RPKM values of all known and predicted RNA binding proteins (RBPs). CPEB1 is shown in the box. Color bar shows log2 scale. Raw values and visualization are also shown in Supplementary Data Set 4b.
Supplementary Figure 4 CPEB1 is specifically upregulated during HCMV infection.
(a) Immunoblot showing CPEB4 protein levels in mock and TB40E HCMV infection of HFFs at 48 and 96 hpi. Beta actin was used as a loading control. (b) Immunoblot showing CPEB2 and CPEB4 protein levels in mock and TB40E infected HFFs at 48hpi. GAPDH was used as a loading control. (c) Immunoblot analysis of hnRNP proteins and core 3′UTR machinery factors in mock (M) and HCMV (V) treated HFFs (MOI 5) at 48 hpi. β-actin was used as a loading control. (d) Immunoblot analysis of CSTF-64 at 24 hours post mock treatment or HCMV infection in presence or absence of CSTF-64 shRNA treatment. (e) qRT-PCR analysis of distal over proximal PCGF3 3′UTRs at 24 and 72 hpi with HCMV after control or CSTF-64 shRNA treatment. Error bars are mean +/- STD; n=3 (f) qPCR showing fold change over mock mRNA levels for CPEB1 in mock, interferon-gamma (IFN-g), UV-inactivated TB40E, and live TB40E HCMV treated HFFs. (g) RT-PCR of exon exclusion levels in SPAG9 and ITGA6 in HCMV treated HFFs compared to IFN-g, UV-inactivated virus, and mock treatment. (h) Immunoblot of CPEB1 in mock or HCMV (TB40E) infected HFFs treated with non-targeting siRNA (NT1) and two different CPEB1 siRNAs (si1 and si2) at 48 hpi. CH160 antibody was used to identify the immediate early HCMV infection markers IE72 and IE86. β-actin was used as a loading control. The antibody used recognized CPEB1 as two bands and both bands were depleted with siRNA treatments. (i) Bar chart showing cell viability (measured by presto blue reagent) in uninfected cells that are untreated or treated with NT siRNA or siCPEB1, and in HCMV infected cells at 48 hpi that are treated with NT siRNA or siCPEB1 siRNA. Error bars show standard deviation between three replicates.
Supplementary Figure 5 CPEB1-related alternative polyadenylation changes in HCMV infection and their enrichment in extracellular exosomal genes.
(a) Scatter plot of mean distal APA site indices (psi values or posterior mean values in Supplementary Data Set 3) of siNT in mock (x-axis) vs. siNT in HCMV-infected (y-axis) cells. Red dots show 3′UTR shortening (76%) and blue dots show 3′UTR lengthening (24%) during HCMV infection (Colored dots have Bayes Factor > 10000); this color codes also applies to (b) and (c). (b) Scatter plot of mean distal APA site indices in GFP OE (x-axis) vs. CPEB1 OE (y-axis). Data shows 3′UTR shortening (77%) and 3′UTR lengthening (23%) with CPEB1 OE. (c) Venn diagram of APA changes in CPEB1 OE and HCMV (+) HFFs measured by RNA-seq. (d) Scatter plot of mean distal APA site indices in siNT in HCMV (x-axis) vs. siCPEB1 in HCMV-infected (y-axis) cells. Data shows 3′UTR shortening (39%) and 3′UTR lengthening (61%) with CPEB1 depletion in HCMV infection. (e) RNA-seq coverage of RPS6KA3 (Ribosomal S6 protein kinase associated with growth) showing 3 ‘UTR shortening (delta-psi = -0.56, Bayes Factor >1012) during infection and reversal with siCPEB1 (delta-psi = 0.15, Bayes Factor = ~5 x 105). 3′UTR shortening is also observed in CPEB1 OE compared with GFP OE (delta-psi = -0.39, Bayes Factor >1012). Known APA sites, target scan (TS) miRNA binding sites and 100-vertebrate nucleotide conservation tracks from UCSC are shown below. (f) Venn diagram showing overlap between APA changes in siNT in mock vs. siNT in HCMV and siNT in HCMV vs. siCPEB1 in HCMV-infection at 48hpi in HFFs. 196 genes are reversed (lengthened) in siCPEB1 compared to HCMV. (g) qRT-PCR analysis of distal 3′UTR usage in PCGF3 and ANKH in mock, HCMV infection (48 hpi), GFP OE, or CPEB1 OE. Error bars represent standard deviation between three replicates. We validated total 3/4 targets tested based on targets common in HCMV infection and CPEB1 OE. (h) DAVID gene ontology (GO) analysis shows progressive enrichment of extracellular exosome category in APA altered genes in siNT in mock vs. siNT in HCMV, siNT in HCMV vs. siCPEB1 in HCMV-infected, and GFP OE vs. CPEB1 OE. Y- axis is –log2 P value with FDR correction applied.
Supplementary Figure 6 siRNA- and antisense oligonucleotide–mediated CPEB1 depletion affects HCMV genes and cell morphology.
(a) PolyA tail length assessment of spike-ins with known tail lengths (0, 8, 16, 32, 64, 128 As) show accurate tail length determination by the Tailseeker2 program. (b) Bar plot showing enriched GO categories in genes decreased in polyA tail lengths post siRNA-mediated CPEB1 depletion in HCMV infected HFFs. (c) Phase contrast images of HFFs under the following conditions: NT siRNA-Mock; NT siRNA-HCMV; CPEB1 siRNA-HCMV; UL99 (pp28 protein) overexpression-CPEB1 siRNA-HCMV; and CPEB1 overexpression-CPEB1 siRNA-HCMV (top). Phase contrast and GFP overlay of UL99-overexpression and CPEB1 siRNA-HCMV, and CPEB1 overexpression-CPEB1 siRNA-HCMV HFFs (bottom) at lower (left) and higher (right) magnifications. GFP is present in the same transcript as UL99 or CPEB1 and translated through an internal ribosome entry site (IRES). CPEB1-GFP lenti is codon optimized and resistant to CPEB1 siRNA. (d) RT-PCR using primers designed on the flanking exons shows activity of 19 different ASOs (including a control ASO in lane 1) designed to target exon 5 of CPEB1 pre-mRNA. ASOs were designed to block the 5’ splice site (5’ ss) or 3’ splice site (ss) of the specified exons. Boxed ASOs 2 and 14 were successful in causing exon skipping. Use of ASO 2 and 14 lead to a decrease in overall CPEB1 mRNA levels. (e) Quantitative RT-PCR with primers designed to interrogate the 3′UTR of CPEB1, showing CPEB1 levels during infection and their reduction by different ASOs and their combinations. Combination of ASOs 14 and 2 (blocking both 5’ and 3’ ss of exon 5) was the most effective in reducing CPEB1 mRNA levels. The error bars are standard error around the mean of the replicates. (f) Immunoblot showing CPEB1, UL57, and pp28 levels in mock, HCMV infected non-targeting (NT) and smart pool (SP) siRNA treated, and ASO-treated HFFs. GAPDH was used as a loading control and UV-treated HCMV was used as a negative control. RT-PCR in the lower panel shows SPAG9 AS for respective samples.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1117 kb)
Supplementary Data Set 1
HCMV gene expression calculations (XLSX 21 kb)
Supplementary Data Set 2
All significantly changing host alternative splicing events from microarrays (XLSX 489 kb)
Supplementary Data Set 3
Significantly changing alternative host polyadenylation events in 2 or more cell types (S3a), Mock vs. HCMV (S3b), HCMV vs. siCPEB1 (S3c), GFP OE vs. CPEB1 OE (S3d), and Mock vs. HSV-2 (S3e) (XLSX 335 kb)
Supplementary Data Set 4
Significant host gene expression changes and clustering groups. (XLSX 3059 kb)
Supplementary Data Set 5
Alternative cassette exon splicing event changes common to CPEB1 OE and HCMV infected cells (XLSX 122 kb)
Supplementary Data Set 6
Significant host gene alternative cassette splicing changes upon CPEB1 KD and CPEB1 OE from RNA-seq (XLSX 1049 kb)
Supplementary Data Set 7
Significant Gene Ontology (GO) enriched categories for APA altered genes (XLSX 46 kb)
Supplementary Data Set 8
Poly(A) tail lengths analyzed by TAIL-seq (XLSX 886 kb)
Supplementary Data Set 9
Antisense oligonucleotides (ASOs) targeting CPEB1 (XLSX 10 kb)
Supplementary Data Set 10
Uncropped immunoblots (PDF 33685 kb)
Supplementary Data Set 11
Primer sequences for RT–PCR and qPCR (XLS 30 kb)
Rights and permissions
About this article
Cite this article
Batra, R., Stark, T., Clark, A. et al. RNA-binding protein CPEB1 remodels host and viral RNA landscapes. Nat Struct Mol Biol 23, 1101–1110 (2016). https://doi.org/10.1038/nsmb.3310
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3310
This article is cited by
-
Unconventional viral gene expression mechanisms as therapeutic targets
Nature (2021)
-
Viral hijacking of the TENT4–ZCCHC14 complex protects viral RNAs via mixed tailing
Nature Structural & Molecular Biology (2020)
-
The sustained expression of Cas9 targeting toxic RNAs reverses disease phenotypes in mouse models of myotonic dystrophy type 1
Nature Biomedical Engineering (2020)
-
Human cytomegalovirus genomics and transcriptomics through the lens of next-generation sequencing: revision and future challenges
Virus Genes (2019)
-
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Nature Communications (2018)