Abstract
HIV-1 vaccine design is informed by structural studies elucidating mechanisms by which broadly neutralizing antibodies (bNAbs) recognize and/or accommodate N-glycans on the trimeric envelope glycoprotein (Env). Variability in high-mannose and complex-type Env glycoforms leads to heterogeneity that usually precludes visualization of the native glycan shield. We present 3.5-Å- and 3.9-Å-resolution crystal structures of the HIV-1 Env trimer with fully processed and native glycosylation, revealing a glycan shield of high-mannose and complex-type N-glycans, which we used to define complete epitopes of two bNAbs. Env trimer was complexed with 10-1074 (against the V3-loop) and IOMA, a new CD4-binding site (CD4bs) antibody. Although IOMA derives from VH1-2*02, the germline gene of CD4bs-targeting VRC01-class bNAbs, its light chain lacks the short CDRL3 that defines VRC01-class bNAbs. Thus IOMA resembles 8ANC131-class/VH1-46–derived CD4bs bNAbs, which have normal-length CDRL3s. The existence of bNAbs that combine features of VRC01-class and 8ANC131-class antibodies has implications for immunization strategies targeting VRC01-like bNAbs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lasky, L.A. et al. Neutralization of the AIDS retrovirus by antibodies to a recombinant envelope glycoprotein. Science 233, 209–212 (1986).
Kwong, P.D. et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393, 648–659 (1998).
Julien, J.P. et al. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science 342, 1477–1483 (2013).
Pancera, M. et al. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature 514, 455–461 (2014).
Scharf, L. et al. Broadly neutralizing antibody 8ANC195 recognizes closed and open states of HIV-1 Env. Cell 162, 1379–1390 (2015).
Garces, F. et al. Affinity maturation of a potent family of HIV antibodies is primarily focused on accommodating or avoiding glycans. Immunity 43, 1053–1063 (2015).
Lee, J.H., de Val, N., Lyumkis, D. & Ward, A.B. Model building and refinement of a natively glycosylated HIV-1 Env protein by high-resolution cryoelectron microscopy. Structure 23, 1943–1951 (2015).
Lee, J.H., Ozorowski, G. & Ward, A.B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 351, 1043–1048 (2016).
Stewart-Jones, G.B.E. et al. Trimeric HIV-1-Env structures define glycan shields from clades A, B, and G. Cell 165, 813–826 (2016).
Doores, K.J. The HIV glycan shield as a target for broadly neutralizing antibodies. FEBS J. 282, 4679–4691 (2015).
Behrens, A.J. et al. Composition and antigenic effects of individual glycan sites of a trimeric HIV-1 envelope glycoprotein. Cell Rep. 14, 2695–2706 (2016).
Binley, J.M. et al. Role of complex carbohydrates in human immunodeficiency virus type 1 infection and resistance to antibody neutralization. J. Virol. 84, 5637–5655 (2010).
Diskin, R. et al. Restricting HIV-1 pathways for escape using rationally designed anti-HIV-1 antibodies. J. Exp. Med. 210, 1235–1249 (2013).
Diskin, R. et al. Increasing the potency and breadth of an HIV antibody by using structure-based rational design. Science 334, 1289–1293 (2011).
Kwon, Y.D. et al. Crystal structure, conformational fixation and entry-related interactions of mature ligand-free HIV-1 Env. Nat. Struct. Mol. Biol. 22, 522–531 (2015).
Kong, L. et al. Complete epitopes for vaccine design derived from a crystal structure of the broadly neutralizing antibodies PGT128 and 8ANC195 in complex with an HIV-1 Env trimer. Acta Crystallogr. D Biol. Crystallogr. 71, 2099–2108 (2015).
Scharf, L. et al. Antibody 8ANC195 reveals a site of broad vulnerability on the HIV-1 envelope spike. Cell Rep. 7, 785–795 (2014).
Zhou, T. et al. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science 329, 811–817 (2010).
Zhou, T. et al. Structural repertoire of HIV-1-neutralizing antibodies targeting the CD4 supersite in 14 donors. Cell 161, 1280–1292 (2015).
Zhou, T. et al. Multidonor analysis reveals structural elements, genetic determinants, and maturation pathway for HIV-1 neutralization by VRC01-class antibodies. Immunity 39, 245–258 (2013).
Jardine, J. et al. Rational HIV immunogen design to target specific germline B cell receptors. Science 340, 711–716 (2013).
Jardine, J.G. et al. Priming a broadly neutralizing antibody response to HIV-1 using a germline-targeting immunogen. Science 349, 156–161 (2015).
McGuire, A.T. et al. Antigen modification regulates competition of broad and narrow neutralizing HIV antibodies. Science 346, 1380–1383 (2014).
McGuire, A.T. et al. Specifically modified Env immunogens activate B-cell precursors of broadly neutralizing HIV-1 antibodies in transgenic mice. Nat. Commun. 7, 10618 (2016).
Jardine, J.G. et al. HIV-1 broadly neutralizing antibody precursor B cells revealed by germline-targeting immunogen. Science 351, 1458–1463 (2016).
McGuire, A.T. et al. Engineering HIV envelope protein to activate germline B cell receptors of broadly neutralizing anti-CD4 binding site antibodies. J. Exp. Med. 210, 655–663 (2013).
Dosenovic, P. et al. Immunization for HIV-1 broadly neutralizing antibodies in human Ig knockin mice. Cell 161, 1505–1515 (2015).
Wu, X. et al. Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329, 856–861 (2010).
West, A.P. Jr., Diskin, R., Nussenzweig, M.C. & Bjorkman, P.J. Structural basis for germ-line gene usage of a potent class of antibodies targeting the CD4-binding site of HIV-1 gp120. Proc. Natl. Acad. Sci. USA 109, E2083–E2090 (2012).
Mouquet, H. et al. Complex-type N-glycan recognition by potent broadly neutralizing HIV antibodies. Proc. Natl. Acad. Sci. USA 109, E3268–E3277 (2012).
Scheid, J.F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009).
Bonsignori, M. et al. Maturation pathway from germline to broad HIV-1 neutralizer of a CD4-mimic antibody. Cell 165, 449–463 (2016).
Scheid, J.F. et al. Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333, 1633–1637 (2011).
Sanders, R.W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Go, E.P. et al. Characterization of glycosylation profiles of HIV-1 transmitted/founder envelopes by mass spectrometry. J. Virol. 85, 8270–8284 (2011).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Kong, L. et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat. Struct. Mol. Biol. 20, 796–803 (2013).
Pejchal, R. et al. A potent and broad neutralizing antibody recognizes and penetrates the HIV glycan shield. Science 334, 1097–1103 (2011).
Scharf, L. et al. Structural basis for germline antibody recognition of HIV-1 immunogens. eLife 5, e13783 (2016).
Adolf-Bryfogle, J., Xu, Q., North, B., Lehmann, A. & Dunbrack, R.L. Jr. PyIgClassify: a database of antibody CDR structural classifications. Nucleic Acids Res. 43, D432–D438 (2015).
North, B., Lehmann, A. & Dunbrack, R.L. Jr. A new clustering of antibody CDR loop conformations. J. Mol. Biol. 406, 228–256 (2011).
Lee, J.H. et al. Antibodies to a conformational epitope on gp41 neutralize HIV-1 by destabilizing the Env spike. Nat. Commun. 6, 8167 (2015).
McLellan, J.S. et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480, 336–343 (2011).
Pancera, M. et al. Structural basis for diverse N-glycan recognition by HIV-1–neutralizing V1-V2–directed antibody PG16. Nat. Struct. Mol. Biol. 20, 804–813 (2013).
Crooks, E.T. et al. Vaccine-elicited tier 2 HIV-1 neutralizing antibodies bind to quaternary epitopes involving glycan-deficient patches proximal to the CD4 binding site. PLoS Pathog. 11, e1004932 (2015).
Murugan, R., Imkeller, K., Busse, C.E. & Wardemann, H. Direct high-throughput amplification and sequencing of immunoglobulin genes from single human B cells. Eur. J. Immunol. 45, 2698–2700 (2015).
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT-PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008).
von Boehmer, L. et al. Antigen-specific antibodies from mouse memory B cells. Nat. Protoc. (in the press) (2016).
Montefiori, D.C. Evaluating neutralizing antibodies against HIV, SIV, and SHIV in luciferase reporter gene assays. Curr. Protoc. Immunol. 64, 12.11 (2005).
Sheskin, D. Handbook of Parametric and Nonparametric Statistical Procedures 3rd edn. (Chapman & Hall/CRC, 2004).
Galimidi, R.P. et al. Intra-spike crosslinking overcomes antibody evasion by HIV-1. Cell 160, 433–446 (2015).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D Biol. Crystallogr. 66, 133–144 (2010).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Jaroszewski, L., Li, Z., Cai, X.H., Weber, C. & Godzik, A. FFAS server: novel features and applications. Nucleic Acids Res. 39, W38–W44 (2011).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol. Crystallogr. 60, 2256–2268 (2004).
Kleywegt, G.J. Use of non-crystallographic symmetry in protein structure refinement. Acta Crystallogr. D Biol. Crystallogr. 52, 842–857 (1996).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Acknowledgements
We thank L. Stamatatos, M. Crispin, and A.-J. Behrens for helpful discussions; J. Vielmetter, J. Nhan, and the Caltech Protein Expression Center for producing proteins; P. Gnanapragasam for performing in-house neutralization assays; J. Kaiser for assistance with data processing; K. Vellizon and N. Thomas for assistance with FACS sorting; T. Oliveira for bioinformatic analyses; M. Murphy and H. Wang for assistance in making figures; the beamline staff at SSRL; A. Hurley (Rockefeller University Hospital) and C. Lehmann and G. Kremer (Infectious Disease Division at the University Hospital Cologne) for supporting the collection of patient material; and members of the Bjorkman and Nussenzweig laboratories for critical reading of the manuscript. This research was supported by National Institute Of Allergy And Infectious Diseases of the National Institutes of Health grant HIVRAD P01 AI100148 (P.J.B.) (the content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health), the Bill and Melinda Gates Foundation (Collaboration for AIDS Vaccine Discovery Grant OPP1124068 (M.C.N. and P.J.B.)), Comprehensive Antibody-Vaccine Immune Monitoring Consortium grant 1032144 (M.S.S.), the Rockefeller University Center for Clinical and Translational Science grant UL1 TR000043/KL2TR000151 from the National Center for Advancing Translational Sciences (L.v.B.), and the Molecular Observatory at Caltech, supported by the Gordon and Betty Moore Foundation. F.K. is supported by the Heisenberg-Program of the DFG (KL 2389/2-1), the European Research Council (ERC-StG639961), and the German Center for Infection Research (DZIF), partner site Bonn-Cologne, Cologne, Germany. Operations at the Stanford Synchrotron Radiation Lightsource are supported by the US Department of Energy and the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
H.B.G., L.v.B., A.P.W. Jr., M.C.N., and P.J.B. conceived the experiments; H.B.G. solved and analyzed crystal structures; L.v.B. performed single-cell experiments and isolated, cloned, and analyzed R1 antibodies; A.P.W. Jr. performed computational and bioinformatics analyses of VRC01-class, IOMA-class, and 8ANC131-class antibodies; H.B.G. and M.S. purified proteins for crystallography and optimized crystals; A.G. and J.G. produced monoclonal antibodies; M.S.S. conducted in vitro neutralization assays; F.K. and G.F. were the principal investigators for the clinical protocols and provided serum and PBMCs of subject R1; H.B.G., L.v.B., A.P.W., M.C.N. and P.J.B. wrote the paper, on which all principal investigators commented.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Patient R1 serum analysis and isolation and characterization of IOMA.
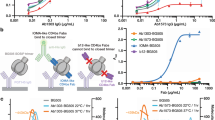
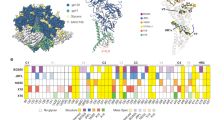
(a) Neutralizing activity of purified serum IgG from patient R1 against a panel of 18 cross-clade HIV-1 viruses. Median inhibitory concentration (IC50) of serum IgG indicated in red (1-50 μg/mL); orange (51-100 μg/mL); yellow (101-300 μg/mL); or white (>500 μg/mL). (b) Neutralizing activity of total R1 serum IgG versus gp140YU2-absorbed R1 serum IgG against two HIV-1 viruses. (c) Representative flow cytometry plots for single cell-sorted 2cc-specific IgG+ B cells, pre-gated on total lymphocytes and singlets. (d) ELISAs measuring the binding of IOMA to wild-type gp140YU2 (left) and to the CD4bs knockout gp140YU2 D368R (right). Black lines with solid dots represent positive control antibody 10-1074; black lines with empty dot represent negative control antibody mGO53; blue lines represent IOMA; the yellow line represents the CD4bs antibody b12. (e) Clonal members of IOMA. Mature IOMA sequences are indicated in red. A dot indicates amino acid identity; a dash indicates an unclear residue in a sequence. Heavy chain clonal members (top). Light chain clonal members (bottom).
Supplementary Figure 2 Neutralizing activity of IOMA.
Top, Clade-specific neutralizing activity of IOMA against a 118-virus panel. Geometric mean IC50 value including non-neutralized strains is indicated for each set of viruses by a red line; the green line indicates the geometric mean IC50 value calculated for only neutralized strains. The number of neutralized and total strains evaluated for each clade are indicated at the top of the panel. Bottom, Neutralizing activity of IOMA against a 118-virus panel. IC50 and IC80 values (μg/mL) are indicated for each virus. Fields in red indicate neutralization.
Supplementary Figure 3 Sequence alignment of CD4-mimetic bNAbs.
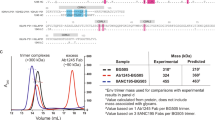
(a) Heavy chains. (b) Light chains. (c) Dendrogram showing relation of VH1-2 (black) and VH1-46 (red) CD4bs bNAbs. The dendrogram was calculated based on nucleotide sequence similarity of their aligned HC V gene segments. Scale bar represents 20% nucleotide substitutions per site.
Supplementary Figure 4 Characterization of BG505 SOSIP.664 protein used for crystallization.
Size exclusion chromatography (SEC) profile showing migration of the BG505 SOSIP.664 used for incubation with IOMA and 10-1074 Fabs to generate samples for crystallization. Inset shows SDS-PAGE analysis under non-reducing conditions of SEC fractions. SEC fractions 7-8 (blue) (larger apparent molecular mass; likely more glycosylated) were used for the 3.9Å IOMA–10-1074–BG505 structure and fractions 11-12 (smaller apparent molecular mass; likely less glycosylated) were used for the 3.5Å IOMA–10-1074–BG505 structure (Table 1).
Supplementary Figure 5 Glycans at individual BG505 SOSIP.664 PNGSs (N88gp120–N160gp120).
Electron density (contoured at 0.8σ) from 2Fo-Fc maps calculated with model phases (labeled as 2Fo) or composite annealed omit maps calculated with phases in which the model was omitted to reduce model bias (Adams, P. D. et al. Acta Crystallogr D Biol Crystallogr 66, 213-221, 2010) (labeled as Omit) shown for PNGSs in the 3.9Å and the 3.5Å IOMA–10-1074–BG505 structures (Table 1). Glycan residues built into the electron density are shown as colored sticks (fucose, red; GlcNAc, blue; mannose, green; galactose, yellow; sialic acid, purple). We identified complex-type N-glycans primarily through the presence of electron density for a core fucose residue. Although the structures were extensively refined (see Methods), coordinates for glycans in our structures should be regarded as approximate: the glycan density was sometimes ambiguous, as expected given the large degree of glycan heterogeneity in HIV-1 Env, one of the most heavily glycosylated proteins known (Doores, K. J. FEBS J 282, 4679-4691, 2015). (b) Schematic versions of crystallographically-identified BG505 SOSIP.664 glycans (fucose, red triangle; GlcNAc, blue square; mannose, green circle; galactose, yellow circle; sialic acid, purple diamond) shown in panel a (labeled as 3.9Å and 3.5Å) compared with glycans from the 4.2Å EM structure of a natively-glycosylated JR-FL (non-SOSIP) Env (Lee, J. H., Ozorowski, G. & Ward, A. B. Science 351, 1043-1048, 2016), the 3.7Å crystal structure of a high mannose-only fully- but not natively-glycosylated BG505 SOSIP.664 (Stewart-Jones, G. B. E. et al. Cell 165, 813-826, 2016), and a mass spectroscopy analysis of natively-glycosylated BG505 SOSIP.664 (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016). Not all glycan species identified at each PNGS by mass spectroscopy (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016) are shown; the major species are depicted in this figure except at PNGSs in which it appears that our crystals contained a subset of BG505 SOSIP.664 protein with underrepresented glycan species (see text). Although the BG505 SOSIP.664 construct (Sanders, R. W. et al. PLoS Pathog 9, e1003618, 2013) was used both for our structures and the mass spectrometry analysis (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016), some differences in glycans at individual PNGSs between the mass spectrometry study and our structures might be expected if the crystals preferentially incorporated a subset of the glycosylation states within the BG505 protein.
Supplementary Figure 6 Glycans at individual BG505 SOSIP.664 PNGSs (N197gp120–N295gp120).
Electron density (contoured at 0.8σ) from 2Fo-Fc maps calculated with model phases (labeled as 2Fo) or composite annealed omit maps calculated with phases in which the model was omitted to reduce model bias (Adams, P. D. et al. Acta Crystallogr D Biol Crystallogr 66, 213-221, 2010) (labeled as Omit) shown for PNGSs in the 3.9Å and the 3.5Å IOMA–10-1074–BG505 structures (Table 1). Glycan residues built into the electron density are shown as colored sticks (fucose, red; GlcNAc, blue; mannose, green; galactose, yellow; sialic acid, purple). We identified complex-type N-glycans primarily through the presence of electron density for a core fucose residue. Although the structures were extensively refined (see Methods), coordinates for glycans in our structures should be regarded as approximate: the glycan density was sometimes ambiguous, as expected given the large degree of glycan heterogeneity in HIV-1 Env, one of the most heavily glycosylated proteins known (Doores, K. J. FEBS J 282, 4679-4691, 2015). (b) Schematic versions of crystallographically-identified BG505 SOSIP.664 glycans (fucose, red triangle; GlcNAc, blue square; mannose, green circle; galactose, yellow circle; sialic acid, purple diamond) shown in panel a (labeled as 3.9Å and 3.5Å) compared with glycans from the 4.2Å EM structure of a natively-glycosylated JR-FL (non-SOSIP) Env (Lee, J. H., Ozorowski, G. & Ward, A. B. Science 351, 1043-1048, 2016), the 3.7Å crystal structure of a high mannose-only fully- but not natively-glycosylated BG505 SOSIP.664 (Stewart-Jones, G. B. E. et al. Cell 165, 813-826, 2016), and a mass spectroscopy analysis of natively-glycosylated BG505 SOSIP.664 (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016). Not all glycan species identified at each PNGS by mass spectroscopy (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016) are shown; the major species are depicted in this figure except at PNGSs in which it appears that our crystals contained a subset of BG505 SOSIP.664 protein with underrepresented glycan species (see text). Although the BG505 SOSIP.664 construct (Sanders, R. W. et al. PLoS Pathog 9, e1003618, 2013) was used both for our structures and the mass spectrometry analysis (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016), some differences in glycans at individual PNGSs between the mass spectrometry study and our structures might be expected if the crystals preferentially incorporated a subset of the glycosylation states within the BG505 protein.
Supplementary Figure 7 Glycans at individual BG505 SOSIP.664 PNGSs (N301gp120–N363gp120).
Electron density (contoured at 0.8σ) from 2Fo-Fc maps calculated with model phases (labeled as 2Fo) or composite annealed omit maps calculated with phases in which the model was omitted to reduce model bias (Adams, P. D. et al. Acta Crystallogr D Biol Crystallogr 66, 213-221, 2010) (labeled as Omit) shown for PNGSs in the 3.9Å and the 3.5Å IOMA–10-1074–BG505 structures (Table 1). Glycan residues built into the electron density are shown as colored sticks (fucose, red; GlcNAc, blue; mannose, green; galactose, yellow; sialic acid, purple). We identified complex-type N-glycans primarily through the presence of electron density for a core fucose residue. Although the structures were extensively refined (see Methods), coordinates for glycans in our structures should be regarded as approximate: the glycan density was sometimes ambiguous, as expected given the large degree of glycan heterogeneity in HIV-1 Env, one of the most heavily glycosylated proteins known (Doores, K. J. FEBS J 282, 4679-4691, 2015). (b) Schematic versions of crystallographically-identified BG505 SOSIP.664 glycans (fucose, red triangle; GlcNAc, blue square; mannose, green circle; galactose, yellow circle; sialic acid, purple diamond) shown in panel a (labeled as 3.9Å and 3.5Å) compared with glycans from the 4.2Å EM structure of a natively-glycosylated JR-FL (non-SOSIP) Env (Lee, J. H., Ozorowski, G. & Ward, A. B. Science 351, 1043-1048, 2016), the 3.7Å crystal structure of a high mannose-only fully- but not natively-glycosylated BG505 SOSIP.664 (Stewart-Jones, G. B. E. et al. Cell 165, 813-826, 2016), and a mass spectroscopy analysis of natively-glycosylated BG505 SOSIP.664 (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016). Not all glycan species identified at each PNGS by mass spectroscopy (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016) are shown; the major species are depicted in this figure except at PNGSs in which it appears that our crystals contained a subset of BG505 SOSIP.664 protein with underrepresented glycan species (see text). Although the BG505 SOSIP.664 construct (Sanders, R. W. et al. PLoS Pathog 9, e1003618, 2013) was used both for our structures and the mass spectrometry analysis (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016), some differences in glycans at individual PNGSs between the mass spectrometry study and our structures might be expected if the crystals preferentially incorporated a subset of the glycosylation states within the BG505 protein.
Supplementary Figure 8 Glycans at individual BG505 SOSIP.664 PNGSs (N386gp120–N637gp41).
Electron density (contoured at 0.8σ) from 2Fo-Fc maps calculated with model phases (labeled as 2Fo) or composite annealed omit maps calculated with phases in which the model was omitted to reduce model bias (Adams, P. D. et al. Acta Crystallogr D Biol Crystallogr 66, 213-221, 2010) (labeled as Omit) shown for PNGSs in the 3.9Å and the 3.5Å IOMA–10-1074–BG505 structures (Table 1). Glycan residues built into the electron density are shown as colored sticks (fucose, red; GlcNAc, blue; mannose, green; galactose, yellow; sialic acid, purple). We identified complex-type N-glycans primarily through the presence of electron density for a core fucose residue. Although the structures were extensively refined (see Methods), coordinates for glycans in our structures should be regarded as approximate: the glycan density was sometimes ambiguous, as expected given the large degree of glycan heterogeneity in HIV-1 Env, one of the most heavily glycosylated proteins known (Doores, K. J. FEBS J 282, 4679-4691, 2015). (b) Schematic versions of crystallographically-identified BG505 SOSIP.664 glycans (fucose, red triangle; GlcNAc, blue square; mannose, green circle; galactose, yellow circle; sialic acid, purple diamond) shown in panel a (labeled as 3.9Å and 3.5Å) compared with glycans from the 4.2Å EM structure of a natively-glycosylated JR-FL (non-SOSIP) Env (Lee, J. H., Ozorowski, G. & Ward, A. B. Science 351, 1043-1048, 2016), the 3.7Å crystal structure of a high mannose-only fully- but not natively-glycosylated BG505 SOSIP.664 (Stewart-Jones, G. B. E. et al. Cell 165, 813-826, 2016), and a mass spectroscopy analysis of natively-glycosylated BG505 SOSIP.664 (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016). Not all glycan species identified at each PNGS by mass spectroscopy (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016) are shown; the major species are depicted in this figure except at PNGSs in which it appears that our crystals contained a subset of BG505 SOSIP.664 protein with underrepresented glycan species (see text). Although the BG505 SOSIP.664 construct (Sanders, R. W. et al. PLoS Pathog 9, e1003618, 2013) was used both for our structures and the mass spectrometry analysis (Behrens, A. J. et al. Cell Reports 14, 2695-2706, 2016), some differences in glycans at individual PNGSs between the mass spectrometry study and our structures might be expected if the crystals preferentially incorporated a subset of the glycosylation states within the BG505 protein.
Supplementary Figure 9 Effects of glycans on neutralization potencies of 10-1074 and IOMA.
(a) Effects of an intact PNGS at N156gp120 on PGT121-family bNAbs. Geometric mean IC50s (µg/mL) of 10-1074, PGT122, and PGT121 were calculated for HIV-1 strains with or without a PNGS at N156gp120. Only strains containing a PNGS at N332gp120 were included in this analysis. Mean IC50s are calculated treating >50 µg/mL values as 50 µg/mL. PGT122 was ~2-fold more potent against strains including the N156gp120 PNGS, whereas 10-1074 showed ~4-fold greater potency against viral strains lacking the N156gp120 PNGS. PGT121 also was more potent against strains lacking PNGS N156gp120. PGT122 has CDRL3 residue 95RLC, while 10-1074 and PGT121 have Ser and Val at position 95LC, respectively. IC50 values are those reported in the following references: Mouquet, H. et al. Proc Natl Acad Sci USA 109, E3268-3277, 2012; Sok, D. et al. Science Translational Medicine 6, 236ra263, 2014; Walker, L. M. et al. Nature 477, 466–470, 2011; Ferguson, A. L. et al. PLoS One 8, e80562, 2013; Huang, J. et al. Nature 515, 138-142, 2014. (b) Effects of glycan deletions at N137gp120, N156gp120, and N301gp120 on 10-1074 neutralization. In vitro neutralization assays were conducted with HIV-1YU2 pseudoviruses that included all glycans (listed as YU2), had introduced mutations to remove the N137gp120 glycan (YU2 N137S), the N156gp120 glycan (YU2 N156K), the N301gp120 glycan (YU2 N301S), or included all glycans in a high mannose form (YU2 Kif). (c,d) Comparison of glycosylation deletions on neutralization by IOMA and VRC01-class bNAbs (VRC01 and NIH45-46). In vitro neutralization assays were conducted with HIV-1YU2 pseudoviruses that included all glycans (listed as YU2), had introduced mutations to remove the N276gp120 glycan (YU2 N276K), the N197gp120 glycan (YU2 N197K), or included all glycans in a high mannose form (YU2 Kif). Removal of glycans or conversion to high-mannose-only glycans had no effect (PNGS N197gp120) or increased IOMA’s neutralization potency by ~4-fold (PNGS N276gp120 and high-mannose-only glycans). For VRC01-class bNAbs, the same changes produced ~20-fold increased potency (PNGS N197gp120), ~4-11–fold increased potency (PNGS N276gp120), or ~2-fold increased potency (high-mannose-only glycans).
Supplementary Figure 10 Comparisons of IOMA, VH1-2/VRC01-class, and VH1-46-class bNAbs.
(a) Rotation angle and translation distance of VH domains of VH1-2 and VH1-46 CD4bs bNAbs in complex with Env (gp120 or trimer) relative to CD4 in complex with HxBc2 gp120 (PDB code 1GC1). Data points for complexes of VRC01-like bNAbs are shown as blue squares; complexes of 8ANC131-like bNAbs are shown as red diamonds; IOMA is shown as a purple square; VRC01 is shown as a blue circle. (b) Interactions of Env residue D368gp120 with IOMA residue R71HC and indicated bNAbs and with CD4 residue R59CD4. Black dashed lines indicate potential hydrogen bonds; red dashed lines indicate potential hydrogen bonds with non-ideal geometry. (c) Interactions of the CD4-binding loop in gp120 with CDRH2 of IOMA and indicated bNAbs and with the C” strand of CD4. Black dashed lines indicate potential hydrogen bonds; gray dashed lines indicate potential hydrogen bonds with non-ideal geometry. (d) IOMA’s interaction with N-glycan attached to N363gp120. R82BHC interacts with the core pentasaccharide of the N363gp120 glycan. Top: 2Fo-Fc electron density map (contoured at 0.8σ) superimposed over coordinates (glycans colored as in Supplementary Figs. 5, 6, 7, 8). Bottom: coordinates alone.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Note (PDF 4990 kb)
Rights and permissions
About this article
Cite this article
Gristick, H., von Boehmer, L., West Jr, A. et al. Natively glycosylated HIV-1 Env structure reveals new mode for antibody recognition of the CD4-binding site. Nat Struct Mol Biol 23, 906–915 (2016). https://doi.org/10.1038/nsmb.3291
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3291
This article is cited by
-
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Nature Communications (2023)
-
Strategies for HIV-1 vaccines that induce broadly neutralizing antibodies
Nature Reviews Immunology (2023)
-
Protein engineering strategies for rational immunogen design
npj Vaccines (2021)
-
Structural basis of transmembrane coupling of the HIV-1 envelope glycoprotein
Nature Communications (2020)
-
Disruption of the HIV-1 Envelope allosteric network blocks CD4-induced rearrangements
Nature Communications (2020)