Abstract
The HIV-1 accessory protein Vpr is required for efficient viral infection of macrophages and promotion of viral replication in T cells. Vpr's biological activities are closely linked to the interaction with human DCAF1, a cellular substrate receptor of the Cullin4–RING E3 ubiquitin ligase (CRL4) of the host ubiquitin–proteasome-mediated protein degradation pathway. The molecular details of how Vpr usurps the protein degradation pathway have not been delineated. Here we present the crystal structure of the DDB1–DCAF1–HIV-1–Vpr–uracil-DNA glycosylase (UNG2) complex. The structure reveals how Vpr engages with DCAF1, creating a binding interface for UNG2 recruitment in a manner distinct from the recruitment of SAMHD1 by Vpx proteins. Vpr and Vpx use similar N-terminal and helical regions to bind the substrate receptor, whereas different regions target the specific cellular substrates. Furthermore, Vpr uses molecular mimicry of DNA by a variable loop for specific recruitment of the UNG2 substrate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Subbramanian, R.A. & Cohen, E.A. Molecular biology of the human immunodeficiency virus accessory proteins. J. Virol. 68, 6831–6835 (1994).
Malim, M.H. & Emerman, M. HIV-1 accessory proteins: ensuring viral survival in a hostile environment. Cell Host Microbe 3, 388–398 (2008).
Kirchhoff, F. Immune evasion and counteraction of restriction factors by HIV-1 and other primate lentiviruses. Cell Host Microbe 8, 55–67 (2010).
Tristem, M., Purvis, A. & Quicke, D.L. Complex evolutionary history of primate lentiviral vpr genes. Virology 240, 232–237 (1998).
Sharifi, H.J., Furuya, A.M. & de Noronha, C.M. The role of HIV-1 Vpr in promoting the infection of nondividing cells and in cell cycle arrest. Curr. Opin. HIV AIDS 7, 187–194 (2012).
Accola, M.A., Ohagen, A. & Göttlinger, H.G. Isolation of human immunodeficiency virus type 1 cores: retention of Vpr in the absence of p6(gag). J. Virol. 74, 6198–6202 (2000).
Re, F., Braaten, D., Franke, E.K. & Luban, J. Human immunodeficiency virus type 1 Vpr arrests the cell cycle in G2 by inhibiting the activation of p34cdc2-cyclin B. J. Virol. 69, 6859–6864 (1995).
Rogel, M.E., Wu, L.I. & Emerman, M. The human immunodeficiency virus type 1 vpr gene prevents cell proliferation during chronic infection. J. Virol. 69, 882–888 (1995).
Belzile, J.P. et al. HIV-1 Vpr-mediated G2 arrest involves the DDB1-CUL4AVPRBP E3 ubiquitin ligase. PLoS Pathog. 3, e85 (2007).
DeHart, J.L. et al. HIV-1 Vpr activates the G2 checkpoint through manipulation of the ubiquitin proteasome system. Virol. J. 4, 57 (2007).
Hrecka, K. et al. Lentiviral Vpr usurps Cul4-DDB1[VprBP] E3 ubiquitin ligase to modulate cell cycle. Proc. Natl. Acad. Sci. USA 104, 11778–11783 (2007).
Le Rouzic, E. et al. HIV1 Vpr arrests the cell cycle by recruiting DCAF1/VprBP, a receptor of the Cul4-DDB1 ubiquitin ligase. Cell Cycle 6, 182–188 (2007).
Schröfelbauer, B., Hakata, Y. & Landau, N.R. HIV-1 Vpr function is mediated by interaction with the damage-specific DNA-binding protein DDB1. Proc. Natl. Acad. Sci. USA 104, 4130–4135 (2007).
Tan, L., Ehrlich, E. & Yu, X.F. DDB1 and Cul4A are required for human immunodeficiency virus type 1 Vpr-induced G2 arrest. J. Virol. 81, 10822–10830 (2007).
Wen, X., Duus, K.M., Friedrich, T.D. & de Noronha, C.M. The HIV1 protein Vpr acts to promote G2 cell cycle arrest by engaging a DDB1 and Cullin4A-containing ubiquitin ligase complex using VprBP/DCAF1 as an adaptor. J. Biol. Chem. 282, 27046–27057 (2007).
Blanchet, F.P., Mitchell, J.P. & Piguet, V. β-TrCP dependency of HIV-1 Vpu-induced downregulation of CD4 and BST-2/tetherin. Curr. HIV Res. 10, 307–314 (2012).
Strebel, K. HIV accessory proteins versus host restriction factors. Curr. Opin. Virol. 3, 692–699 (2013).
Feng, Y., Baig, T.T., Love, R.P. & Chelico, L. Suppression of APOBEC3-mediated restriction of HIV-1 by Vif. Front. Microbiol. 5, 450 (2014).
Laguette, N. et al. Premature activation of the SLX4 complex by Vpr promotes G2/M arrest and escape from innate immune sensing. Cell 156, 134–145 (2014).
Romani, B., Shaykh Baygloo, N., Aghasadeghi, M.R. & Allahbakhshi, E. HIV-1 Vpr protein enhances proteasomal degradation of MCM10 DNA replication factor through the Cul4-DDB1[VprBP] E3 ubiquitin ligase to induce G2/M cell cycle arrest. J. Biol. Chem. 290, 17380–17389 (2015).
Hrecka, K. et al. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 474, 658–661 (2011).
Ahn, J. et al. HIV/simian immunodeficiency virus (SIV) accessory virulence factor Vpx loads the host cell restriction factor SAMHD1 onto the E3 ubiquitin ligase complex CRL4DCAF1. J. Biol. Chem. 287, 12550–12558 (2012).
DeLucia, M., Mehrens, J., Wu, Y. & Ahn, J. HIV-2 and SIVmac accessory virulence factor Vpx down-regulates SAMHD1 enzyme catalysis prior to proteasome-dependent degradation. J. Biol. Chem. 288, 19116–19126 (2013).
Goldstone, D.C. et al. HIV-1 restriction factor SAMHD1 is a deoxynucleoside triphosphate triphosphohydrolase. Nature 480, 379–382 (2011).
Schwefel, D. et al. Structural basis of lentiviral subversion of a cellular protein degradation pathway. Nature 505, 234–238 (2014).
Schwefel, D. et al. Molecular determinants for recognition of divergent SAMHD1 proteins by the lentiviral accessory protein Vpx. Cell Host Microbe 17, 489–499 (2015).
Wu, Y. et al. Structural basis of clade-specific engagement of SAMHD1 (sterile α motif and histidine/aspartate-containing protein 1) restriction factors by lentiviral viral protein X (Vpx) virulence factors. J. Biol. Chem. 290, 17935–17945 (2015).
Bouhamdan, M. et al. Human immunodeficiency virus type 1 Vpr protein binds to the uracil DNA glycosylase DNA repair enzyme. J. Virol. 70, 697–704 (1996).
Ahn, J. et al. HIV-1 Vpr loads uracil DNA glycosylase-2 onto DCAF1, a substrate recognition subunit of a cullin 4A-ring E3 ubiquitin ligase for proteasome-dependent degradation. J. Biol. Chem. 285, 37333–37341 (2010).
Krokan, H.E. & Bjørås, M. Base excision repair. Cold Spring Harb. Perspect. Biol. 5, a012583 (2013).
Kennedy, E.M., Amie, S.M., Bambara, R.A. & Kim, B. Frequent incorporation of ribonucleotides during HIV-1 reverse transcription and their attenuated repair in macrophages. J. Biol. Chem. 287, 14280–14288 (2012).
Yan, N., O'Day, E., Wheeler, L.A., Engelman, A. & Lieberman, J. HIV DNA is heavily uracilated, which protects it from autointegration. Proc. Natl. Acad. Sci. USA 108, 9244–9249 (2011).
Malim, M.H. APOBEC proteins and intrinsic resistance to HIV-1 infection. Phil. Trans. R. Soc. Lond. B 364, 675–687 (2009).
Hrecka, K. et al. HIV-1 and HIV-2 exhibit divergent interactions with HLTF and UNG2 DNA repair proteins. Proc. Natl. Acad. Sci. USA 113, E3921–E3930 (2016).
Eldin, P. et al. Vpr expression abolishes the capacity of HIV-1 infected cells to repair uracilated DNA. Nucleic Acids Res. 42, 1698–1710 (2014).
Chen, R., Le Rouzic, E., Kearney, J.A., Mansky, L.M. & Benichou, S. Vpr-mediated incorporation of UNG2 into HIV-1 particles is required to modulate the virus mutation rate and for replication in macrophages. J. Biol. Chem. 279, 28419–28425 (2004).
Guenzel, C.A. et al. Recruitment of the nuclear form of uracil DNA glycosylase into virus particles participates in the full infectivity of HIV-1. J. Virol. 86, 2533–2544 (2012).
Weil, A.F. et al. Uracil DNA glycosylase initiates degradation of HIV-1 cDNA containing misincorporated dUTP and prevents viral integration. Proc. Natl. Acad. Sci. USA 110, E448–E457 (2013).
Kaiser, S.M. & Emerman, M. Uracil DNA glycosylase is dispensable for human immunodeficiency virus type 1 replication and does not contribute to the antiviral effects of the cytidine deaminase Apobec3G. J. Virol. 80, 875–882 (2006).
Mol, C.D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: protein mimicry of DNA. Cell 82, 701–708 (1995).
Xu, C. & Min, J. Structure and function of WD40 domain proteins. Protein Cell 2, 202–214 (2011).
Parker, J.B. et al. Enzymatic capture of an extrahelical thymine in the search for uracil in DNA. Nature 449, 433–437 (2007).
Morellet, N., Bouaziz, S., Petitjean, P. & Roques, B.P. NMR structure of the HIV-1 regulatory protein VPR. J. Mol. Biol. 327, 215–227 (2003).
Selig, L. et al. Uracil DNA glycosylase specifically interacts with Vpr of both human immunodeficiency virus type 1 and simian immunodeficiency virus of sooty mangabeys, but binding does not correlate with cell cycle arrest. J. Virol. 71, 4842–4846 (1997).
Li, T., Chen, X., Garbutt, K.C., Zhou, P. & Zheng, N. Structure of DDB1 in complex with a paramyxovirus V protein: viral hijack of a propeller cluster in ubiquitin ligase. Cell 124, 105–117 (2006).
Li, T., Robert, E.I., van Breugel, P.C., Strubin, M. & Zheng, N. A promiscuous α-helical motif anchors viral hijackers and substrate receptors to the CUL4–DDB1 ubiquitin ligase machinery. Nat. Struct. Mol. Biol. 17, 105–111 (2010).
Fischer, E.S. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147, 1024–1039 (2011).
Yeh, J.I. et al. Damaged DNA induced UV-damaged DNA-binding protein (UV-DDB) dimerization and its roles in chromatinized DNA repair. Proc. Natl. Acad. Sci. USA 109, E2737–E2746 (2012).
Fischer, E.S. et al. Structure of the DDB1–CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014).
Scrima, A. et al. Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex. Cell 135, 1213–1223 (2008).
Gérard, F.C. et al. Defining the interactions and role of DCAF1/VPRBP in the DDB1-cullin4A E3 ubiquitin ligase complex engaged by HIV-1 Vpr to induce a G2 cell cycle arrest. PLoS One 9, e89195 (2014).
Connor, R.I., Chen, B.K., Choe, S. & Landau, N.R. Vpr is required for efficient replication of human immunodeficiency virus type-1 in mononuclear phagocytes. Virology 206, 935–944 (1995).
Mashiba, M., Collins, D.R., Terry, V.H. & Collins, K.L. Vpr overcomes macrophage-specific restriction of HIV-1 Env expression and virion production. Cell Host Microbe 16, 722–735 (2014).
Srivastava, S. et al. Lentiviral Vpx accessory factor targets VprBP/DCAF1 substrate adaptor for cullin 4 E3 ubiquitin ligase to enable macrophage infection. PLoS Pathog. 4, e1000059 (2008).
Sharova, N. et al. Primate lentiviral Vpx commandeers DDB1 to counteract a macrophage restriction. PLoS Pathog. 4, e1000057 (2008).
Lee, J. & Zhou, P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol. Cell 26, 775–780 (2007).
Angers, S. et al. Molecular architecture and assembly of the DDB1–CUL4A ubiquitin ligase machinery. Nature 443, 590–593 (2006).
DeLaBarre, B. & Brunger, A.T. Considerations for the refinement of low-resolution crystal structures. Acta Crystallogr. D Biol. Crystallogr. 62, 923–932 (2006).
Karplus, P.A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D Biol. Crystallogr. 60, 2210–2221 (2004).
Acknowledgements
We thank J. Skowronski for carefully reading the manuscript and providing us with valuable critical comments and suggestions. We also acknowledge M. Becker and C. Ogata at GM/CA (Argonne National Laboratory), and I. Mathews, C. Smith and A. Gonzalez at the Stanford Synchrotron Radiation Lightsource (SSRL) for their support during data collection. We thank D. Lee for home-source X-ray technical support. This work was supported by NIH grant P50GM082251 (to A.M.G.). G.C. acknowledges support from BioXFEL-STC1231306. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract no. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH.
Author information
Authors and Affiliations
Contributions
A.M.G., J.A. and G.C. conceived the study. Y.W., X.Z., A.M.G., J.A. and G.C. designed the experiments and analyzed the data. Y.W. and G.C. performed crystallization, data collection, structure refinement and analysis. X.Z., M.D. and J.A. performed mutagenesis and structure validation analysis. C.O.B. and A.E.C. performed X-ray data processing. M.D. and J.A. prepared recombinant protein complexes. Y.W., X.Z., A.M.G., J.A. and G.C. wrote the manuscript. All authors discussed the results, commented and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Overall structure of the DDB1–DCAF1–Vpr–UNG2 complex.
(A) The final refined 2Fo-Fc electron density map, contoured at 1.4σ of Vpr, illustrating well-defined side chains.
(B–D) Three views of the final refined 2Fo-Fc electron density map of the Vpr (red)-UNG2 (green) interacting region, contoured at 1.4σ. (B) Electron density for both Vpr and UNG2 residues. (C) Electron density for UNG2 residues. (D) Electron density for Vpr residues.
(E) Two views of the dimeric arrangement of the complex in the ASU.
(F) Schematic depiction of the protein constructs used for crystallization. The color scheme for the individual proteins is used throughout the manuscript.
(G) Crystal contacts across the ASU.
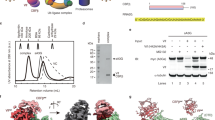
(H) Multi-angle light scattering of the DDB1-DCAF1 (blue), DDB1-DCAF1-Vpr (red) and a mixture of DDB1-DCAF1-Vpr with excess of UNG2 (green). All samples were injected into an analytical Superdex200 gel filtration column. Molecular masses of DDB1-DCAF1, DDB1-DCAF1-Vpr, DDB1-DCAF1-Vpr-UNG2 and UNG2 were estimated to be 168 kDa, 182 kDa, 206 kDa, and 32 kDa, respectively.
Supplementary Figure 2 Vpr-UNG2 interactions.
(A) Ribbon representation of the Vpr structure in the DDB1-DCAF1-Vpr-UNG2 complex.
(B) Electrostatic surface representation of Vpr, illustrating pertinent structural features important for binding UNG2 (green) and DCAF1 (blue). Vpr bridges UNG2 and DCAF1 and no direct contacts between the latter two are present in the structure. The contact area of Vpr with UNG2 (insert loop) is negatively charged, mimicking the negative charge of the DNA phosphate backbone.
(C) Stereo-view of the best-fit superposition of the UNG2-DNA (PDB 2OYT)42 complex (brown and purple worms, respectively) and the UNG2-Vpr (green and red worms respectively) portion of the DDB1-DCAF1-Vpr-UNG2 complex structures. DIL: DNA intercalating loop. Conformational changes in UNG2 in these structures involve mainly the DNA binding groove and the DIL.
(D) Two expanded views of (C), illustrating details of the interaction between UNG2 and DNA (brown and purple ribbons, respectively) and UNG2 and Vpr, (green and red ribbons, respectively). The left panel illustrates the close match in distances between residues in helices α1 and α2 and the minor groove phosphate backbone of the DNA. The right panel depicts residues in helices α1 and α2 that mimic the curvature of the minor groove phosphate backbone of the DNA.
(E-F) Two expanded views of the superposition of the Vpr insert loop and the phosphate backbone around the abasic sugar, illustrating Vpr’s structural mimicry.
(G-H) Structure of UNG2 (green ribbon) complexed with Ugi (cyan ribbon)40.
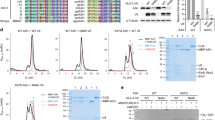
(I) UNG2 base excision and inhibition assay using a ssDNA substrate in the presence and absence of Vpr. The control reaction was carried out without UNG2. DDB1-DCAF1 alone is not able to inhibit UNG2 deamination of the ssDNA substrate, while increasing amounts of DDB1-DCAF1-Vpr added to the reaction inhibits UNG2 activity.
Supplementary Figure 3 DCAF1 interactions with Vpr and DDB1.
(A) View of DCAF1 (blue, ribbon) bridging DDB1 (orange, surface) and Vpr (red, surface). Residues on the DCAF1 loops form the binding interface for Vpr. Residues on DCAF1's helix-loop-helix motif and residues on DCAF1 loops on the opposite site of the Vpr binding site form the DDB1 binding interface.
(B-C) View down onto the DCAF1 WD40 propeller (electrostatic surface representation), illustrating Vpr (red, ribbon) binding. The surface of
the β-propeller is asymmetric; loops connecting the β-strands on one hemi-solenoid of the propeller create the cleft where Vpr binds.
(D) Two views of the interaction between the DCAF1 loops (thick trace, blue; β-strands blue worm) and Vpr (thin trace, red). Pertinent side chain contacts are depicted in the right side panel (stick representation).
(E) Ribbon representation of the interaction between DCAF1 (blue) and DDB1 (orange) in the current structure.
(F) Superposition of DCAF1 (blue) and DDB2 (green) (PDB 3EI1)49 bound to DDB1 (orange).
(G) Structure of the complex between DDB2 (green) and DDB1 (orange) (PDB 3EI1)49.
(H) Detail of the interaction between the helix-loop-helix region of DCAF1 (blue, ribbon) and DDB1 (orange, ribbon). Pertinent side chain interactions are depicted in stick representation.
(I) Pull-down and western blot. Samples are lysates from HEK293 cells, transiently transfected with FLAG-tagged DCAF1 and Myc-tagged DDB1.
(J-K) Two views of the superposition of the DCAF1 (blue)-Vpr (red) portion of the DDB1-DCAF1-Vpr-UNG2 complex with DDB2 (green)-DNA (purple) portion of the DDB1-DDB2-DNA complex (PDB 3EI1)49.
Supplementary Figure 4 Vpr and Vpx structures.
(A-C) Ribbon representation of VprHIV-1 (red), VpxMND (blue) and VpxSM (gray). The hydrophobic cleft in the three-helix bundle of Vpr into which Leu72 and Val74 from UNG2 bind is depicted by a green and black surface. Side chains that fill this pocket in VpxMND include Trp45 and Tyr67 and in VpxSM Trp49 and Tyr71.
(D) DCAF1-Vpx-SAMHD1 structure (PDB 4Z8L)27, illustrating H-bonds between the acidic loop residues of DCAF1 (1090-1095) and Tyr62, Tyr65 and Arg66 of VpxMND (depicted in blue ball and stick) that coordinate interactions with Arg14 from SAMHD1 (depicted in yellow ball and stick)26,27. In contrast, Ala55 and Ala59 of Vpr in the DDB1-DCAF1-Vpr-UNG2 structure, depicted in red, do not form hydrogen bonds with the acidic loop of DCAF1.
(E) Detail of the interactions between the N-terminal tails of VprHIV-1 (red) and VpxMND (blue) and DCAF1 (grey) in the DDB1-DCAF1-Vpr-UNG2 and DCAF1-Vpx-SAMHD1 complex structures, illustrating how SAMHD1's (yellow) N-terminal tail influences the positioning of N-terminus of Vpx. DCAF1 is depicted in surface representation while the other structures are shown in ribbon representation.
(F) Expanded view of the interaction between the insert loop of VpxMND (blue) and SAMHD1 (yellow) in ribbon representation (PDB 4Z8L)27.
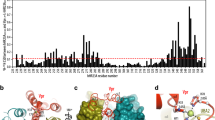
(G) Structure-based sequence alignment of VprHIV-1, VpxMND and VpxSM proteins. The positions of the three helices are illustrated by red, blue and silver bars. Zn2+ coordinating His and Cys residues in all proteins are shown in red. DCAF1 interacting residues and UNG2 (for Vpr) or SAMHD1 interacting residues (for Vpx) are shown in blue and green, respectively.
Supplementary Figure 5 Model of the CRL4–DCAF1 E3 ubiquitin ligase complex.
(A) Superposition of the BPA domain of DDB1 from the DDB1-DCAF1-Vpr-UNG2 structure (orange surface and worm representation) with the BPA domain of the DDB1-CUL4A-RBX1-S5V structure (PDB 2HYE56, grey surface and worm representation) and the BPA domain from the DDB1-CUL4A-Rbx1-DDB2-DNA structure (PDB 4A0L46, blue surface and worm representation). The BPA and BPC domains in all three structures adopt similar conformations, while the BPB domains are rotated with respect to each other46.
(B) Alternative view of (A) illustrating the relative disposition of the BPB domains in all three structures. In the inset, the different directions adopted by the first β-strand of BPB, following the hinge residues (388-392, superimposed) connecting BPA and BPB is shown.
(C-D) Two views of a CUL4A-RBX1-DDB1-DCAF1-Vpr-UNG2 model, based on the superposition depicted in (A) and (B), using PDB 2HYE56, gray; PDB 4A0L46, blue). UNG2 gets positioned close to the RBX1 structure, thus poised for ubiquitination.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 865 kb)
Supplementary Data Set 1
Uncropped western blot images from main figures (PDF 2823 kb)
Rights and permissions
About this article
Cite this article
Wu, Y., Zhou, X., Barnes, C. et al. The DDB1–DCAF1–Vpr–UNG2 crystal structure reveals how HIV-1 Vpr steers human UNG2 toward destruction. Nat Struct Mol Biol 23, 933–940 (2016). https://doi.org/10.1038/nsmb.3284
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3284
This article is cited by
-
PROTAC targeted protein degraders: the past is prologue
Nature Reviews Drug Discovery (2022)
-
HIV-1 Vpr activates host CRL4-DCAF1 E3 ligase to degrade histone deacetylase SIRT7
Virology Journal (2021)
-
Structure of HIV-1 Vpr in complex with the human nucleotide excision repair protein hHR23A
Nature Communications (2021)
-
Meet the IUPAB councilor — Angela M. Gronenborn
Biophysical Reviews (2021)
-
Structural basis of indisulam-mediated RBM39 recruitment to DCAF15 E3 ligase complex
Nature Chemical Biology (2020)