Abstract
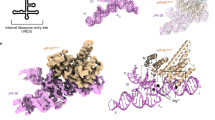
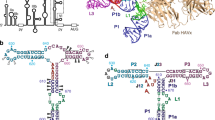
Many viruses bypass canonical cap-dependent translation in host cells by using internal ribosomal entry sites (IRESs) in their transcripts; IRESs hijack initiation factors for the assembly of initiation complexes. However, it is currently unknown how IRES RNAs recognize initiation factors that have no endogenous RNA binding partners; in a prominent example, the IRES of encephalomyocarditis virus (EMCV) interacts with the HEAT-1 domain of eukaryotic initiation factor 4G (eIF4G). Here we report the solution structure of the J-K region of this IRES and show that its stems are precisely organized to position protein-recognition bulges. This multisite interaction mechanism operates on an all-or-nothing principle in which all domains are required. This preorganization is accomplished by an 'adjuster module': a pentaloop motif that acts as a dual-sided docking station for base-pair receptors. Because subtle changes in the orientation abrogate protein capture, our study highlights how a viral RNA acquires affinity for a target protein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Filbin, M.E. & Kieft, J.S. Toward a structural understanding of IRES RNA function. Curr. Opin. Struct. Biol. 19, 267–276 (2009).
Gebauer, F. & Hentze, M.W. IRES unplugged. Science 351, 228 (2016).
Weingarten-Gabbay, S. et al. Systematic discovery of cap-independent translation sequences in human and viral genomes. Science http://dx.doi.org/10.1126/science.aad4939 (2016).
Xue, S. et al. RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 517, 33–38 (2015).
Berry, K.E., Waghray, S., Mortimer, S.A., Bai, Y. & Doudna, J.A. Crystal structure of the HCV IRES central domain reveals strategy for start-codon positioning. Structure 19, 1456–1466 (2011).
Hashem, Y. et al. Hepatitis-C-virus-like internal ribosome entry sites displace eIF3 to gain access to the 40S subunit. Nature 503, 539–543 (2013).
Lukavsky, P.J. Structure and function of HCV IRES domains. Virus Res. 139, 166–171 (2009).
Bochkov, Y.A. & Palmenberg, A.C. Translational efficiency of EMCV IRES in bicistronic vectors is dependent upon IRES sequence and gene location. Biotechniques 41, 283–292 (2006).
Hundsdoerfer, P., Thoma, C. & Hentze, M.W. Eukaryotic translation initiation factor 4GI and p97 promote cellular internal ribosome entry sequence-driven translation. Proc. Natl. Acad. Sci. USA 102, 13421–13426 (2005).
Spriggs, K.A. et al. Canonical initiation factor requirements of the Myc family of internal ribosome entry segments. Mol. Cell. Biol. 29, 1565–1574 (2009).
Knight-Jones, T.J. & Rushton, J. The economic impacts of foot and mouth disease: what are they, how big are they and where do they occur? Prev. Vet. Med. 112, 161–173 (2013).
Dibrov, S.M. et al. Hepatitis C virus translation inhibitors targeting the internal ribosomal entry site. J. Med. Chem. 57, 1694–1707 (2014).
Pestova, T.V., Hellen, C.U. & Shatsky, I.N. Canonical eukaryotic initiation factors determine initiation of translation by internal ribosomal entry. Mol. Cell. Biol. 16, 6859–6869 (1996).
Pestova, T.V., Shatsky, I.N. & Hellen, C.U. Functional dissection of eukaryotic initiation factor 4F: the 4A subunit and the central domain of the 4G subunit are sufficient to mediate internal entry of 43S preinitiation complexes. Mol. Cell. Biol. 16, 6870–6878 (1996).
Lomakin, I.B., Hellen, C.U. & Pestova, T.V. Physical association of eukaryotic initiation factor 4G (eIF4G) with eIF4A strongly enhances binding of eIF4G to the internal ribosomal entry site of encephalomyocarditis virus and is required for internal initiation of translation. Mol. Cell. Biol. 20, 6019–6029 (2000).
Duke, G.M., Hoffman, M.A. & Palmenberg, A.C. Sequence and structural elements that contribute to efficient encephalomyocarditis virus RNA translation. J. Virol. 66, 1602–1609 (1992).
Kolupaeva, V.G., Pestova, T.V., Hellen, C.U. & Shatsky, I.N. Translation eukaryotic initiation factor 4G recognizes a specific structural element within the internal ribosome entry site of encephalomyocarditis virus RNA. J. Biol. Chem. 273, 18599–18604 (1998).
Lozano, G. & Martínez-Salas, E. Structural insights into viral IRES-dependent translation mechanisms. Curr. Opin. Virol. 12, 113–120 (2015).
Bassili, G. et al. Sequence and secondary structure requirements in a highly conserved element for foot-and-mouth disease virus internal ribosome entry site activity and eIF4G binding. J. Gen. Virol. 85, 2555–2565 (2004).
Clark, A.T., Robertson, M.E., Conn, G.L. & Belsham, G.J. Conserved nucleotides within the J domain of the encephalomyocarditis virus internal ribosome entry site are required for activity and for interaction with eIF4G. J. Virol. 77, 12441–12449 (2003).
Hoffman, M.A. & Palmenberg, A.C. Mutational analysis of the J-K stem-loop region of the encephalomyocarditis virus IRES. J. Virol. 69, 4399–4406 (1995).
Kolupaeva, V.G., Lomakin, I.B., Pestova, T.V. & Hellen, C.U. Eukaryotic initiation factors 4G and 4A mediate conformational changes downstream of the initiation codon of the encephalomyocarditis virus internal ribosomal entry site. Mol. Cell. Biol. 23, 687–698 (2003).
López de Quinto, S. & Martínez-Salas, E. Interaction of the eIF4G initiation factor with the aphthovirus IRES is essential for internal translation initiation in vivo. RNA 6, 1380–1392 (2000).
Correll, C.C. & Swinger, K. Common and distinctive features of GNRA tetraloops based on a GUAA tetraloop structure at 1.4 A resolution. RNA 9, 355–363 (2003).
Fiore, J.L. & Nesbitt, D.J. An RNA folding motif: GNRA tetraloop-receptor interactions. Q. Rev. Biophys. 46, 223–264 (2013).
Hura, G.L. et al. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat. Methods 6, 606–612 (2009).
Reyes, F.E., Schwartz, C.R., Tainer, J.A. & Rambo, R.P. Methods for using new conceptual tools and parameters to assess RNA structure by small-angle X-ray scattering. Methods Enzymol. 549, 235–263 (2014).
Kaminski, A. & Jackson, R.J. The polypyrimidine tract binding protein (PTB) requirement for internal initiation of translation of cardiovirus RNAs is conditional rather than absolute. RNA 4, 626–638 (1998).
Theimer, C.A., Wang, Y., Hoffman, D.W., Krisch, H.M. & Giedroc, D.P. Non-nearest neighbor effects on the thermodynamics of unfolding of a model mRNA pseudoknot. J. Mol. Biol. 279, 545–564 (1998).
Jones, S., Daley, D.T., Luscombe, N.M., Berman, H.M. & Thornton, J.M. Protein-RNA interactions: a structural analysis. Nucleic Acids Res. 29, 943–954 (2001).
Marcotrigiano, J. et al. A conserved HEAT domain within eIF4G directs assembly of the translation initiation machinery. Mol. Cell 7, 193–203 (2001).
Yu, Y., Abaeva, I.S., Marintchev, A., Pestova, T.V. & Hellen, C.U. Common conformational changes induced in type 2 picornavirus IRESs by cognate trans-acting factors. Nucleic Acids Res. 39, 4851–4865 (2011).
Boehr, D.D., Nussinov, R. & Wright, P.E. The role of dynamic conformational ensembles in biomolecular recognition. Nat. Chem. Biol. 5, 789–796 (2009).
Tucker, B.J. & Breaker, R.R. Riboswitches as versatile gene control elements. Curr. Opin. Struct. Biol. 15, 342–348 (2005).
Oberer, M., Marintchev, A. & Wagner, G. Structural basis for the enhancement of eIF4A helicase activity by eIF4G. Genes Dev. 19, 2212–2223 (2005).
Schütz, P. et al. Crystal structure of the yeast eIF4A-eIF4G complex: an RNA-helicase controlled by protein-protein interactions. Proc. Natl. Acad. Sci. USA 105, 9564–9569 (2008).
Akabayov, S.R. & Wagner, G. Backbone resonance assignment of the HEAT1-domain of the human eukaryotic translation initiation factor 4GI. Biomol. NMR Assign. 8, 89–91 (2014).
Hyberts, S.G., Milbradt, A.G., Wagner, A.B., Arthanari, H. & Wagner, G. Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J. Biomol. NMR 52, 315–327 (2012).
Schwieters, C.D. & Clore, G.M. Using small angle solution scattering data in Xplor-NIH structure calculations. Prog. Nucl. Magn. Reson. Spectrosc. 80, 1–11 (2014).
Acknowledgements
This work was supported by NIH grants GM047467 and EB002026 (to G.W.) and R01 AI51340 (to C.U.T.H.). This work was also supported by the Japan Society for the Promotion of Science (JSPS) Postdoctoral Fellowships for Research Abroad, the Daiichi-Sankyo Foundation of Life Science, and the Naito Foundation (to S.I.). We would also like to acknowledge the Department of Energy (DOE) Integrated Diffraction Analysis (IDAT) grant, contract no. DE-AC02-05CH11231, for the SAXS data acquisition. The authors wish to thank M. Durney for many helpful discussions.
Author information
Authors and Affiliations
Contributions
S.I., C.U.T.H., V.M.D'S., and G.W. conceived and designed the experiments. S.I. purified the samples and performed the NMR, ITC, and SAXS experiments. S.I. and V.M.D'S. performed the structural analyses. P.K. conducted the toeprinting assay. S.I., C.U.T.H., V.M.D'S. and G.W. interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 NMR signal assignments and structural analysis of SL-J.
(a) Sequence and NMR-derived secondary structure of SL-J. Colors indicate the sequential assignment in (d). (b) Overlay of the Ade H2–C2, H8–C8, and H1′–C1′ regions of 1H-13C HMQC spectra of [13C,15N-Ade/Cyt] SL-J (red) and [13C,15N-Ade] J-K (black). Asterisks indicate signals due to heterogenous termini in the RNA transcript. (c) 1H-1H NOESY spectrum of the imino proton region with the imino proton walk of SL-J that supports the secondary structure of SL-J. The lower and upper stem walks are shown with red and cyan lines, respectively. (d) Sequential walk of H1′–H6 (pyrimidine) or H1′–H8 (purine) connectivity for the assignments. Labels show the intraresidual H1′–H6 or H1′–H8 signals of each residue. Green and cyan, and purple lines represent the sequential walk of the segments shown in the same colors in (a). Asterisks indicate that the signals are below the threshold. (e-m) NOESY strips derived from 13C-edited HMQC-NOESY 3D spectra of [13C,15N-Ade/Cyt] or [13C,15N-Gua/Uri] labeled SL-J for the structural characterization of the bulges. Asterisks indicate that the signals are not in the same plane in the 13C dimension and not connected to the spin system, or that the intensities of the signals are not determined because of overlaps. (n) A NOESY strip derived from 1H-1H NOESY spectrum of SL-J for the structural characterization of the U703 bulge.
Supplementary Figure 2 NMR signal assignments and structural analysis of SL-K.
(a) Sequence and NMR-derived secondary structure of SL-K. Colors indicate the sequential assignment in (d). (b) Overlay of the Ade H2–C2, H8–C8, and H1′–C1′ regions of 1H-13C HMQC spectra of [13C,15N-Ade/Cyt] SL-K (red) and [13C,15N-Ade] J-K (black). Asterisks indicate signals due to heterogenous termini in the RNA transcript. (c) 1H-1H NOESY spectrum of the imino proton region and imino proton walk of SL-K that supports the secondary structure of SL-K. The lower and upper stem walks are shown with red and cyan lines, respectively. The NOE signals between upfield shifted imino protons from U738 and U759 indicate a non-canonical basepair formed by these residues. (d) Sequential walk of H1′–H6 (pyrimidine) or H1′–H8 (purine) connectivity for the assignments. Labels show the intraresidual H1′–H6 or H1′–H8 signals of each residue. Green, cyan, and purple lines represent the sequential walk of the segments shown in the same colors in (a). Asterisks indicate that the signals are below the threshold. (e, g) NOESY strips derived from 1H-1H NOESY spectrum of SL-K for the structural characterization of the bulges. (f, h, j) NOESY strips derived from 13C-edited HMQC-NOESY 3D spectra of [13C,15N-Ade/Cyt] or [13C,15N-Gua/Uri] labeled SL-K for the structural characterization of the bulges. The asterisk in (f) indicates that the signal is not in the same plane in the 13C dimension and not connected to the spin system. (i) The Gua H8–C8 region of 1H-13C HMQC spectrum of [13C,15N-Gua/Uri] SL-K.
Supplementary Figure 3 NMR signal assignments and structural analysis of SL-St.
(a) Sequence and NMR-derived secondary structure of SL-St. Colors indicate the sequential assignment in (d). (b) Overlay of the Ade H2–C2, H8–C8, and H1′–C1′ regions of 1H-13C HMQC spectra of [13C,15N-Ade/Cyt] SL-St (red) and [13C,15N-Ade] J-K (black). (c) 1H-1H NOESY spectrum of the imino proton region and imino proton walk of SL-St that supports the secondary structure of SL-St. The lower and upper stem walks are represented in red and cyan, respectively. (d) Sequential walk of H1′–H6 (pyrimidine) and H1′–H8 (purine) connectivity for the assignments. Labels show the intraresidual H1′–H6 (pyrimidine) or H1′–H8 (purine) signals of each residue. Green and cyan lines represent the sequential walk of the segments shown in the same colors in (a). Asterisks indicate that the signals are below the threshold. (e-l, n) NOESY strips derived from 13C-edited HMQC-NOESY 3D spectra of [13C,15N-Ade/Cyt] and [13C,15N-Gua/Uri] labeled SL-St for the structural characterization of the bulges. Asterisks indicate that the signals are not in the same plane in the 13C dimension and not connected to the spin system, or that the intensities of the signals are not determined because of overlaps. (m) NOESY strip derived from 1H-1H NOESY spectrum of SL-St for the structural characterization of the bulges.
Supplementary Figure 4 NMR signal assignments and structural analysis of ∆J∆K.
(a) Sequence and NMR-derived secondary structure of ∆J∆K. Colors indicate the sequential assignment in (c). (b) Overlay of the 1H-1H NOESY spectra of [u-2H,{H1′,H2′,H2,H8}-Ade] ∆J∆K (black) and [u-2H,{H1′,H2′,H2,H8}-Ade] J-K (gray). (c) 1H-1H NOESY spectrum of the imino proton region and imino proton walk of ∆J∆K that supports the secondary structure of ∆J∆K. The imino walks are shown with red, cyan, and purple lines, respectively. (d) Sequential walk of H1′–H6 (pyrimidine) or H1′–H8 (purine) connectivity for the assignments. Labels show the intraresidual H1′–H6 or H1′–H8 signals of each residue. Green, cyan and purple lines represent the sequential walk of the segments shown in the same colors in (a). Asterisks indicate that the signals are below the threshold. (e) 1H-1H NOESY spectrum of [u-2H, {H1′,H2′,H2,H8}-Ade] ∆J∆K in H2O. (f) Regions of 1H-1H NOESY spectra of unlabeled ∆J∆K (top) and [u-2H, 1H-Ade/Gua] ∆J∆K (bottom). (g-i) Regions of 1H-1H NOESY spectra showing the NOE connectivities from the ASL to the J and K domains. (j) A region of 1H-1H NOESY spectrum of [u-2H, {H1′,H2′,H2,H8}-Ade] ∆J∆K in H2O. The long range connectivities that shows the register of the ASL to the J and K domains are labeled in red.
Supplementary Figure 5 Mg2+ concentration dependence of the J-K–HEAT-1 interaction.
(a) Results of ITC experiments in which the HEAT-1 domain was titrated to the J-K region at 2, 1, 0.5, and 0 mM of Mg2+. The interaction between the J-K region and the HEAT-1 domain requires divalent ions as seen by the lack of heat of interaction in the absence Mg2+ ions. (b) NMR spectral overlay. Left: Overlay of the 1H-13C HMQC spectra of [13C,15N-Ade] J-K in the absence (black) and presence of 1 mM Mg2+ (red). Right: Overlay of the 1H-1H NOESY spectra of [u-2H, {H1′,H2′,H2,H8}-Ade] J-K in the absence (black) and presence of 1 mM Mg2+ (red). (c) SAXS profiles of the J-K region in the absence (black) and presence of excess (2 mM) Mg2+ (red). (d) Results of the ITC experiment in which Mg2+ was titrated to the J-K region.
Supplementary Figure 6 SAXS, NMR, and ITC data.
(a) Results of the SAXS analyses. Top left: The intensity plots of SAXS data of the J-K region (black), U-rich loop mutant (red), and J-K–HEAT-1 domain complex (cyan). I is scattering intensity, q is proportional to the scattering angle (q = 4sinθ/λ, where 2q is the angle between the incident x-ray beam and the detector, and λ is x-ray wavelength in Å). Right: The Guinier plots of the J-K region, U-rich loop mutant, and J-K–HEAT-1 domain complex. The linear least square fitting provided the radius of gyration of 28.9 ± 0.1, 29.6 ± 0.1, and 34.9 ± 0.1 Å for the J-K region, U-rich loop mutant, and J-K/HEAT-1 domain complex, respectively. Bottom left: The pairwise distance distribution function (PDDF) plots of the J-K region (black), U-rich loop mutant (red), and J-K–HEAT-1 domain complex (cyan), respectively. (b) NMR titration experiment. 1H-1H NOESY spectra of [u-2H, {H1′,H2′,H2,H8}-Ade] J-K region in the absence (black) and substoichiometric presence (red) of the HEAT-1 domain, shown as in Fig. 5b. Assignments of the intraresidual H1′–H8 NOE signals are labeled with the residue numbers, whereas strong intraresidual NOE correlation between H2 and H1′ are labeled with residue numbers and atom names. Residues whose NOE signal intensities were substantially reduced by the addition of the HEAT-1 domain are labeled in black, as in Fig. 5b. (c) Temperature dependent NMR spectral change of ∆J∆K. Left: The H6–C6 region of 1H-13C HMQC spectrum of ∆J∆K at 35ºC. The H6–C6 signal of U769 is down-field shifted in the 13C dimension, indicating that it adopts the syn conformation. Right: The same region of the spectrum at 45ºC. The H6–C6 signals of U768 (at the bottom of the K domain) and U769 (one of the residue from the closing basepair of ASL domain) are split, whereas the others are not. (d) 1H-1H NOESY spectra of the wild type and U-rich loop mutant of the J-K region. Top: 1H-1H NOESY spectrum of [u-2H, {H1′,H2′,H2,H8}-Ade] J-K. Bottom: 1H-1H NOESY spectrum of [u-2H, {H1′,H2′,H2,H8}-Ade] U-rich loop mutant. H1′ resonances are shown with vertical lines with the intraresidual NOE signals labeled as H2 and H8. Interresidual NOE signals from each H1′ resonance are labeled with residue numbers. Labels and lines are colored red (St), green (J), cyan (K), and gray (ASL), depending on the domains that belong to. In each panel, the secondary structure is shown on the right with the mutated stretch highlighted in orange. (e) Results of ITC experiments where the HEAT-1 domain was titrated to the A771U and C696A–G728U mutant of the J-K region. (f) 1H-1H NOESY spectra of the wild type and C696A–G728U of [u-2H, {H1′,H2′,H2,H8}-Ade] ∆J∆K. Although signals from the ASL domain residues (A770-A775) were not perturbed, the long range NOE between A771 H2 (ASL) and A697 H1′ (J) disappeared (green boxes), indicating that the long range interaction between the ASL and J domains is abrogated by the mutation.
Supplementary Figure 7 Structure model of the J-K–HEAT-1 domain complex and mutagenic analyses.
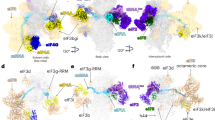
(a) Positions of exposed aromatic residues on the HEAT-1 domain. Conserved and exposed aromatic residues of the eIF4G1 HEAT-1 domain are mapped onto the crystal structure of the homologous eIF4G2 (PDB ID: 1HU3). (b) Results of the ITC experiments of the interaction of F777A, F812A, F949A, and F978A HEAT-1 domain mutants with the J-K region. (c) Structure model of the J-K–HEAT-1 domain complex derived from the SAXS, NMR, and previous mutagenesis assay. Structures of the J-K region and the eIF4G2 HEAT-1 domain (PDB ID: 1HU3) are placed into the SAXS ab initio envelope structure (gray). The basic residues previously identified to be involved in the interaction (R821, K827, R841, and K842) are shown with blue spheres (d,e) The results of assaying formation of binary eIF4G–IRES and ternary eIF4G–eIF4A–IRES complexes by eIF4G(736-1115) (d) and eIF4G(653-1599) (e). The bands labeled as C786 and F.L. represent the fall-off of the reverse transcriptase (RT) at C786 by the binding of eIF4G to the J-K region, and the full-length cDNA where the RT is not stopped by the eIF4G–IRES complex, respectively. (f) The results 48S complex formation assays by eIF4G(736-1115) (lanes 3-6) and eIF4G(653-1599) (lanes 7-10). The 48S formation is evidenced by the bands labeled as 48S(AUG834), which reflects the fall-off of the RT by the 48S complex formed at the first codon(AUG834).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note (PDF 3479 kb)
Source data
Rights and permissions
About this article
Cite this article
Imai, S., Kumar, P., Hellen, C. et al. An accurately preorganized IRES RNA structure enables eIF4G capture for initiation of viral translation. Nat Struct Mol Biol 23, 859–864 (2016). https://doi.org/10.1038/nsmb.3280
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3280
This article is cited by
-
A conserved RNA structural motif for organizing topology within picornaviral internal ribosome entry sites
Nature Communications (2019)
-
Viral RNA structure-based strategies to manipulate translation
Nature Reviews Microbiology (2019)
-
Ribosome-dependent conformational flexibility changes and RNA dynamics of IRES domains revealed by differential SHAPE
Scientific Reports (2018)
-
Evaluation of 15N-detected H–N correlation experiments on increasingly large RNAs
Journal of Biomolecular NMR (2017)