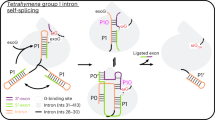
Abstract
Bacterial group II introns are large catalytic RNAs related to nuclear spliceosomal introns and eukaryotic retrotransposons. They self-splice, yielding mature RNA, and integrate into DNA as retroelements. A fully active group II intron forms a ribonucleoprotein complex comprising the intron ribozyme and an intron-encoded protein that performs multiple activities including reverse transcription, in which intron RNA is copied into the DNA target. Here we report cryo-EM structures of an endogenously spliced Lactococcus lactis group IIA intron in its ribonucleoprotein complex form at 3.8-Å resolution and in its protein-depleted form at 4.5-Å resolution, revealing functional coordination of the intron RNA with the protein. Remarkably, the protein structure reveals a close relationship between the reverse transcriptase catalytic domain and telomerase, whereas the active splicing center resembles the spliceosomal Prp8 protein. These extraordinary similarities hint at intricate ancestral relationships and provide new insights into splicing and retromobility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Lambowitz, A.M. & Zimmerly, S. Group II introns: mobile ribozymes that invade DNA. Cold Spring Harb. Perspect. Biol. 3, a003616 (2011).
Lambowitz, A.M. & Belfort, M. Mobile bacterial group II introns at the crux of eukaryotic evolution. Microbiol Spectr. 3 MDNA3-0050-2014 (2015).
Will, C.L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Luan, D.D., Korman, M.H., Jakubczak, J.L. & Eickbush, T.H. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell 72, 595–605 (1993).
Zimmerly, S., Guo, H., Perlman, P.S. & Lambowitz, A.M. Group II intron mobility occurs by target DNA-primed reverse transcription. Cell 82, 545–554 (1995).
Nakamura, T.M. & Cech, T.R. Reversing time: origin of telomerase. Cell 92, 587–590 (1998).
Eickbush, T.H. Telomerase and retrotransposons: which came first? Science 277, 911–912 (1997).
Marcia, M. & Pyle, A.M. Visualizing group II intron catalysis through the stages of splicing. Cell 151, 497–507 (2012).
Robart, A.R., Chan, R.T., Peters, J.K., Rajashankar, K.R. & Toor, N. Crystal structure of a eukaryotic group II intron lariat. Nature 514, 193–197 (2014).
Toor, N., Keating, K.S., Taylor, S.D. & Pyle, A.M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008).
Wank, H., SanFilippo, J., Singh, R.N., Matsuura, M. & Lambowitz, A.M. A reverse transcriptase/maturase promotes splicing by binding at its own coding segment in a group II intron RNA. Mol. Cell 4, 239–250 (1999).
Singh, R.N., Saldanha, R.J., D'Souza, L.M. & Lambowitz, A.M. Binding of a group II intron-encoded reverse transcriptase/maturase to its high affinity intron RNA binding site involves sequence-specific recognition and autoregulates translation. J. Mol. Biol. 318, 287–303 (2002).
Watanabe, K. & Lambowitz, A.M. High-affinity binding site for a group II intron-encoded reverse transcriptase/maturase within a stem-loop structure in the intron RNA. RNA 10, 1433–1443 (2004).
Matsuura, M. et al. A bacterial group II intron encoding reverse transcriptase, maturase, and DNA endonuclease activities: biochemical demonstration of maturase activity and insertion of new genetic information within the intron. Genes Dev. 11, 2910–2924 (1997).
Matsuura, M., Noah, J.W. & Lambowitz, A.M. Mechanism of maturase-promoted group II intron splicing. EMBO J. 20, 7259–7270 (2001).
Gupta, K. et al. Quaternary arrangement of an active, native group II intron ribonucleoprotein complex revealed by small-angle X-ray scattering. Nucleic Acids Res. 42, 5347–5360 (2014).
Pyle, A.M. The tertiary structure of group II introns: implications for biological function and evolution. Crit. Rev. Biochem. Mol. Biol. 45, 215–232 (2010).
Dai, L. et al. A three-dimensional model of a group II intron RNA and its interaction with the intron-encoded reverse transcriptase. Mol. Cell 30, 472–485 (2008).
Saldanha, R. et al. RNA and protein catalysis in group II intron splicing and mobility reactions using purified components. Biochemistry 38, 9069–9083 (1999).
Blocker, F.J. et al. Domain structure and three-dimensional model of a group II intron-encoded reverse transcriptase. RNA 11, 14–28 (2005).
Stoddard, B.L. Homing endonucleases from mobile group I introns: discovery to genome engineering. Mob. DNA 5, 7 (2014).
Rambo, R.P. & Doudna, J.A. Assembly of an active group II intron-maturase complex by protein dimerization. Biochemistry 43, 6486–6497 (2004).
Bryan, T.M., Goodrich, K.J. & Cech, T.R. Tetrahymena telomerase is active as a monomer. Mol. Biol. Cell 14, 4794–4804 (2003).
Jiang, J. et al. Structure of Tetrahymena telomerase reveals previously unknown subunits, functions, and interactions. Science 350, aab4070 (2015).
Golas, M.M. et al. 3D cryo-EM structure of an active step I spliceosome and localization of its catalytic core. Mol. Cell 40, 927–938 (2010).
Newman, A.J. & Nagai, K. Structural studies of the spliceosome: blind men and an elephant. Curr. Opin. Struct. Biol. 20, 82–89 (2010).
Galej, W.P., Nguyen, T.H., Newman, A.J. & Nagai, K. Structural studies of the spliceosome: zooming into the heart of the machine. Curr. Opin. Struct. Biol. 25, 57–66 (2014).
Nguyen, T.H. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015).
Gillis, A.J., Schuller, A.P. & Skordalakes, E. Structure of the Tribolium castaneum telomerase catalytic subunit TERT. Nature 455, 633–637 (2008).
Mitchell, M., Gillis, A., Futahashi, M., Fujiwara, H. & Skordalakes, E. Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA. Nat. Struct. Mol. Biol. 17, 513–518 (2010).
Gu, S.Q. et al. Genetic identification of potential RNA-binding regions in a group II intron-encoded reverse transcriptase. RNA 16, 732–747 (2010).
Galej, W.P., Oubridge, C., Newman, A.J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013).
Toor, N., Rajashankar, K., Keating, K.S. & Pyle, A.M. Structural basis for exon recognition by a group II intron. Nat. Struct. Mol. Biol. 15, 1221–1222 (2008).
San Filippo, J. & Lambowitz, A.M. Characterization of the C-terminal DNA-binding/DNA endonuclease region of a group II intron-encoded protein. J. Mol. Biol. 324, 933–951 (2002).
Cui, X., Matsuura, M., Wang, Q., Ma, H. & Lambowitz, A.M. A group II intron-encoded maturase functions preferentially in cis and requires both the reverse transcriptase and X domains to promote RNA splicing. J. Mol. Biol. 340, 211–231 (2004).
Noah, J.W. & Lambowitz, A.M. Effects of maturase binding and Mg2+ concentration on group II intron RNA folding investigated by UV cross-linking. Biochemistry 42, 12466–12480 (2003).
Hang, J., Wan, R., Yan, C. & Shi, Y. Structural basis of pre-mRNA splicing. Science 349, 1191–1198 (2015).
Yan, C. et al. Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349, 1182–1191 (2015).
Singh, N.N. & Lambowitz, A.M. Interaction of a group II intron ribonucleoprotein endonuclease with its DNA target site investigated by DNA footprinting and modification interference. J. Mol. Biol. 309, 361–386 (2001).
Aizawa, Y., Xiang, Q., Lambowitz, A.M. & Pyle, A.M. The pathway for DNA recognition and RNA integration by a group II intron retrotransposon. Mol. Cell 11, 795–805 (2003).
Noah, J.W. et al. Atomic force microscopy reveals DNA bending during group II intron ribonucleoprotein particle integration into double-stranded DNA. Biochemistry 45, 12424–12435 (2006).
Sternberg, S.H., LaFrance, B., Kaplan, M. & Doudna, J.A. Conformational control of DNA target cleavage by CRISPR-Cas9. Nature 527, 110–113 (2015).
Dlakić, M. & Mushegian, A. Prp8, the pivotal protein of the spliceosomal catalytic center, evolved from a retroelement-encoded reverse transcriptase. RNA 17, 799–808 (2011).
de Lange, T. A loopy view of telomere evolution. Front. Genet. 6, 321 (2015).
Pei, J., Kim, B.H. & Grishin, N.V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli . J. Microsc. 146, 113–136 (1987).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Scheres, S.H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Popenda, M. et al. Automated 3D structure composition for large RNAs. Nucleic Acids Res. 40, e112 (2012).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Kelley, L.A. & Sternberg, M.J. Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009).
Trabuco, L.G., Villa, E., Schreiner, E., Harrison, C.B. & Schulten, K. Molecular dynamics flexible fitting: a practical guide to combine cryo-electron microscopy and X-ray crystallography. Methods 49, 174–180 (2009).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We are thankful to X.(E.) Dong for useful discussions. We thank the Tsinghua University Branch of the National Center for Protein Sciences (Beijing) for providing the cryo-EM and computational facility support. The high-performance computation was completed on the Explorer 100 cluster system at the Tsinghua National Laboratory for Informational Science and Technology. Work was supported by NIH grants GM39422 and GM44844 (to M.B.), NIH grant GM61576 (to R.K.A.) and by the National Science Foundation of China grant 31270765 (to H.-W.W.).
Author information
Authors and Affiliations
Contributions
M.B. and H.-W.W. conceived the project; G.Q. and C.L.P. purified the RNP complex and performed activity assays; J.W., H.S. and H.-W.W. performed EM analysis and obtained the cryo-EM maps; R.K.A. directed the molecular interpretation of the cryo-EM map; P.S.K. and R.K.A. analyzed the cryo-EM maps and generated the atomic model of the RNP complex; P.S.K., G.Q., H.-W.W. and R.K.A. analyzed the molecular structure; and G.Q., P.S.K., R.K.A., M.B. and H.-W.W. analyzed the overall data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 RNP purification, components and activity.
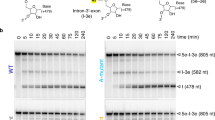
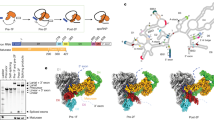
(a) Intein-mediated RNP production and purification. The Ll.LtrB intron RNA (red) (Exons E1 and E2 in green) and associated intron encoded protein (LtrA in yellow) were co-expressed with nisin induction. The RNP was captured and purified on chitin resin column through the chitin binding domain (CBD) that is fused to the LtrA C-terminus, released by DTT cleavage of the intein (I), and further purified by sucrose gradient sedimentation. (b) Product purity. LtrB intron RNA (902 nt) and associated LtrA protein (~70 kD) were analyzed, respectively, on a 1.2% denaturing formaldehyde agarose gel (left, lane 1, R) and on 12% denaturing SDS-PAGE gels (right, lanes 1, P), stained with Coomassie (blue) and silver (brown). Silver staining also revealed a higher molecular weight product, likely corresponding to intron RNA which stains with silver, not seen in a purified LtrA control (lanes 2). (c) RNA analysis. RNA in purified spliced RNPs (lanes 3 and 5) or extracted from the RNP (lane 4) were assayed by primer extension and the cDNA analyzed on an 8% urea- PAGE gel. RNPs were either wild-type (wt) or a splicing-defective mutant of the catalytic triad (Triad). A primer, specific to the 5' end of the intron (black arrow and P), was used to reveal precursor (pre-mRNA) and spliced intron, from which 68-nt and 41-nt cDNA products, respectively, were generated. Lane 1 corresponds to total RNA isolated from cells expressing the group II intron RNP and lane 2 to no RNA. The purified RNP resulted in predominantly a 41-nt band with a minor amount of precursor (lane 3), indicating that most of the pre-mRNA had been removed. In contrast, the splicing defective Triad RNP yielded a primer extension product that corresponded exclusively to the intron-containing precursor (lane 5). (d) Activity assays with target DNA. Purified RNPs incubated with target DNA (T, 52nt) were assayed for DNA binding (left) and reverse splicing activity (right) on a 4% native-PAGE and a 10% urea-PAGE gel, respectively. Lanes 1-3 contain no RNP, wild-type RNP and RNP from the catalytic triad mutant, respectively. Only the wt RNP, and not the catalytic mutant Triad, yielded shifted target DNA (left, black triangles). Likewise, only the wt RNP was active in reverse splicing (right) on the target DNA (IS=integration site, CS=cleavage site), producing a high molecular-weight integration product (black triangle) and the corresponding reverse splicing product (RSP) and cleavage product (CP).
Supplementary Figure 2 Negative-staining EM and random conical tilt reconstruction of the Ll.LtrB group II intron RNP.
(a, b) One pair of electron micrographs of an untilted (a) and tilted (b) negatively stained specimen of the group II intron RNP complex. Four tilt pair particles are labeled with colored boxes, one color labeling one molecule. (c) Two-dimensional class averages of the particle images picked from micrographs of untilted specimen. Two major populated class averages are marked in cyan and lavender respectively. (d) The top row shows the RCT 3D reconstruction volumes from the tilted particle images corresponding to the two major class averages shown in (c). The bottom row shows 3D reconstructions obtained by refining the dataset merged from all the untilted and tilted particle images against the two RCT reconstruction volumes, respectively.
Supplementary Figure 3 Cryo-EM of the Ll.LtrB group II intron RNP and LtrA-depleted intron.
(a) A representative raw micrograph of frozen-hydrated intron RNP particles generated from the 300 kV Titan Krios microscope. Few typical particles are identified by green square boxes. (b) Computed power spectrum of the micrograph in panel A. (c) Outline of the image processing steps to obtain the 3.8 Å resolution cryo-EM map of the group II intron RNP complex and the 4.5 Å resolution map of the LtrA-depleted intron RNA. Reference-free 2D classification of 608,019 particle images generated ~200 2D class averages of the group II intron RNP comprising good particles free of ice contaminants and other junks. A 3D classification of the selected particles into four classes yielded three classes with strong LtrA protein occupancy that were merged as a total particle dataset of 450,296 particles and one class lacking density for the LtrA protein as a dataset of 102,522 particles. The first round of 3D auto-refinement of the merged dataset yielded a reconstruction at 4.2 Å resolution. A soft mask (shown in light grey semitransparency) around the most rigid portion of the RNP complex was used for further refinement to generate a final 3D reconstruction at 3.8 Å resolution. Local resolution maps of the 3D reconstructions without and with a soft mask during the refinement are shown on the right of the 4.2-Å and 3.8-Å resolution maps, respectively. The volumes are sliced to show the central portion of the local resolution distribution. A refinement of the LtrA-depleted intron dataset yielded a reconstruction at 4.5 Å resolution. The Euler angle distributions of particle images used for the intron RNP and the LtrA-depleted intron RNA reconstructions are shown next to corresponding EM maps. (d) Corrected gold-standard FSC curves of the final 3D reconstructions of RNP or LtrA-depleted intron RNA with or without a soft mask, respectively, after post-processing in RELION 1.3.
Supplementary Figure 4 Structure comparisons of the group IIA intron in the LtrA-bound state with that in the LtrA-depleted state and the group IIB intron.
(a) Comparison of group IIA intron structure in LtrA-bound state (color coding of RNA domains are according to Fig. 1b) and IEP-depleted state (dark slate gray) shows a RMSD value of 2.4 Å. Regions corresponding to helices Id1 and Id3ii encompassing EBS2 and EBS1, respectively, are enlarged on upper left and right corners. (b) Comparison between tertiary structures of individual domains of the group IIA intron (this study) and group IIB intron (PDB ID: 4R0D). Domains of the group IIA intron are color coded and labeled as in Fig. 2, whereas that of the group IIB intron are shown as light grey color. Differences are displayed for each domain between the two introns. The β-β' interaction site, which is unique to group IIA intron, is boxed in red in the upper left panel. (c) Difference in relative positions of RNA helices of domain I, Id(iv) and domain IV with respect to domain V (red) between the group IIA and group IIB introns. The relative movements in group IIA intron helices are indicated by black arrows. RNA helices from group IIA and group IIB introns are shown in matching colors, except that the IIB intron helices are shown as semitransparent tubes. (d) Fourier shell correlation between atomic coordinates with the corresponding EM densities. RNP, intron-LtrA complex; and DV, domain V of the intron RNA.
Supplementary Figure 5 Overall fitting of the initial homology model and final model of LtrA domains into the cryo-EM map.
(a) The fingers-palm domain initial homology model (left panel) and final model (middle panel) docked into the corresponding cryo-EM density. Superposition of initial and final models (right panel) shows an RMSD value of 1.9 Å. (b) The thumb domain initial homology model (left panel) and final model (middle panel) docked into the corresponding cryo-EM density. Superposition of initial and final models (right panel) shows an RMSD value of 1.6 Å. (c) The EN domain initial homology model (left panel) and final model after flipping α-helix (middle panel) docked into the corresponding cryo-EM density. It should be noted that the α-helix corresponds to the DBD but was segmented with the EN domain. Superimposition of the aligned segments of the initial homology model and final EN model (right panel) shows an RMSD value of 1.7 Å. Density corresponding to the loop that connects the two-β strands was disordered and hence not modeled. (d) and (e), Overall fittings of NTD and DBD into the corresponding cryo-EM masses. The 3.8 Å resolution cryo-EM map (shown as semitransparent densities in matching colors) was used to model fingers-palm (a), thumb (b) and NTD (d). The dockings of EN (c) and DBD (e) domains were performed in the 5 Å resolution map (corresponding densities shown as semitransparent mass).
Supplementary Figure 6 Structure-based sequence alignment of the RT and EN domains.
Structure based sequence alignments are performed using PROMALS3D 45. (a) Structure based sequence alignment of the RT fingers-palm domains of LtrA and Tribolium castaneum TERT (PDB: 3DU5). Secondary structures are shown with arrows for β-strands and bars for α helices. Identical residues are shaded in grey and indicated underneath with asterisks. Residues involved in TPRT in TERT and corresponding residues in LtrA are highlighted in different colors. Red, catalytic residues; Turquoise, dNTP binding pocket; Green, DNA primer grip; Purple, RNA template interaction. (b) Alignment of the RT thumb domains of LtrA and Saccharomyces cerevisiae Prp8 (PDB: 3ZEF). α-helices are shown as bars, identical residues are shaded in grey and indicated underneath with asterisks. (c) Structure based sequence alignment of the EN domains from LtrA and the putative HNH endonuclease Gmet_0936 protein from Geobacter metallireducens GS-15 (PDB: 4H9D), HNH homing endonuclease I-HmuI (PDB: 1U3E), and E7 toxin (PDB: 1MZ8). Secondary structures are shown with arrows for β-strands and bars for α helices. The typical HNH motif residues are in red and four cysteine residues conserved between LtrA and Geobacter HNH endonuclease and proposed to be in additional metal ion coordination are in magenta. (d) Structure based sequence alignment of RT fingers-palm and thumb domains from LtrA, TERT, Prp8, and HIV-1 RT (PDB: 2HMI). Conserved sequence blocks RT1-RT7 in the fingers- palm domains (rectangular boxes) and the three parallel helices from Prp8 (H1-H3, boxes are in red, orange and green, as in panel b) in thumb domains in LtrA, TERT and HIV-1 RT, are defined using HIV-1 RT and Prp8 as references, respectively. Sequences in α-helices and β-strands are in red and blue, respectively. Insertions between RT sequence blocks, 2a, 4a, 5a, and 7a, and the thumb insertion ti are shown and labeled in red. Sequences of IFD-like motifs, are shaded in grey. Conserved aspartic acid residues at the RT active site are marked by red arrows. Motifs 1, 2, A, B’, C, D, E (tan letters) in Tribolium castaneum TERT are in dark red boxes. Sequences that are not structurally resolved are boxed with dashed grey lines.
Supplementary Figure 7 Comparison of the fingers–palm domain of LtrA with that of TERT and Prp8.
(a) Docking of fingers-palm domain of TERT (PDB code: 3DU6; left panel) and Prp8 (PDB code: 3JB9; right panel) into the cryo-EM density corresponding to the LtrA fingers-palm domain. (b) Structure superimposition of LtrA fingers-palm domains with corresponding domains from TERT (left panel) and Prp8 (right panel) show the RMSD values of 2.0 Å and 2.7 Å, respectively. (c) Rigid-body docking of TERT (left panel) and Prp8 (right panel) fingers domain onto group IIA intron shows a significant steric clash between Prp8 and intron domain Id(iii)a.
Supplementary Figure 8 Comparison of the thumb domain of LtrA with those of Prp8 and TERT.
(a) Docking of the thumb domain of Prp8 (PDB code: 3JB9; right panel) and TERT (PDB code: 3DU6; left panel) into the cryo-EM density corresponding to LtrA’s thumb domain. (b) Structure superimposition of the LtrA thumb domain with corresponding domains from Prp8 (left panel) and TERT (right panel) show the RMSD values of 1.5 Å and 2.8 Å, respectively.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Table 1 and Supplementary Note (PDF 2455 kb)
Supplementary Data Set 1
Uncropped gel image; referring to Figure 1e where the boxed part is shown (PDF 627 kb)
3D EM map and atomic model of the group II intron RNP
The 3.8 Å resolution cryo-EM map, with segmented densities for RNA (yellow) and LtrA protein (blue), is shown while rotating around a vertical axis. Whereas in the second rotation RNA and protein are further segmented into their structural domains, the third rotation shows the final model. The color scheme is same as in Fig. 1. (MP4 7995 kb)
Identification of mRNA density within the group II intron RNP complex
A portion of the 3.8 Å resolution cryo-EM map, comprising the mRNA density bound to EBS1, EBS2 and the interacting segment of the thumb domain of LtrA protein, is shown. The atomic models of EBS1 and EBS2 (cyan), mRNA (magenta), and LtrA (blue) are identified. (MP4 1464 kb)
Comparison between the fingers/palm RT domains of LtrA and TERT
The superimposed atomic models of the finger and palm domains of LtrA protein (magenta and purple) and TERT (light grey) are shown while rotating around a vertical axis. (MP4 2584 kb)
Comparison between the fingers/palm RT domains of LtrA and Prp8
The superimposed atomic models of the finger and palm domains of LtrA protein (magenta and purple) and Prp8 (light grey) are shown while rotating around a vertical axis. (MP4 2583 kb)
Comparison between the fingers/palm RT domains of LtrA and HIV-RT
The superimposed atomic models of the finger and palm domains of LtrA protein (magenta and purple) and HIV-RT (light grey) are shown while rotating around a vertical axis. (MP4 2582 kb)
Comparison between the thumb domains of LtrA and Prp8
The superimposed atomic models of the thumb domains of LtrA protein (blue) and Prp8 (light grey) are shown while rotating around a vertical axis. (MP4 2584 kb)
Rights and permissions
About this article
Cite this article
Qu, G., Kaushal, P., Wang, J. et al. Structure of a group II intron in complex with its reverse transcriptase. Nat Struct Mol Biol 23, 549–557 (2016). https://doi.org/10.1038/nsmb.3220
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3220
This article is cited by
-
Methylation of rRNA as a host defense against rampant group II intron retrotransposition
Mobile DNA (2021)
-
Metal ions and sugar puckering balance single-molecule kinetic heterogeneity in RNA and DNA tertiary contacts
Nature Communications (2020)
-
Visualizing group II intron dynamics between the first and second steps of splicing
Nature Communications (2020)
-
Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories
Mobile DNA (2017)
-
Globular domain structure and function of restriction-like-endonuclease LINEs: similarities to eukaryotic splicing factor Prp8
Mobile DNA (2017)