Abstract
Elongation factor 4 (EF4) is a key quality-control factor in translation. Despite its high conservation throughout evolution, EF4 deletion in various organisms has not yielded a distinct phenotype. Here we report that genetic ablation of mitochondrial EF4 (mtEF4) in mice causes testis-specific dysfunction in oxidative phosphorylation, leading to male infertility. Deletion of mtEF4 accelerated mitochondrial translation at the cost of producing unstable proteins. Somatic tissues overcame this defect by activating mechanistic (mammalian) target of rapamycin (mTOR), thereby increasing rates of cytoplasmic translation to match rates of mitochondrial translation. However, in spermatogenic cells, the mTOR pathway was downregulated as part of the developmental program, and the resulting inability to compensate for accelerated mitochondrial translation caused cell-cycle arrest and apoptosis. We detected the same phenotype and molecular defects in germline-specific mtEF4-knockout mice. Thus, our study demonstrates cross-talk between mtEF4-dependent quality control in mitochondria and cytoplasmic mTOR signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Wallace, D.C. Mitochondria, bioenergetics, and the epigenome in eukaryotic and human evolution. Cold Spring Harb. Symp. Quant. Biol. 74, 383–393 (2009).
Smits, P., Smeitink, J. & van den Heuvel, L. Mitochondrial translation and beyond: processes implicated in combined oxidative phosphorylation deficiencies. J. Biomed. Biotechnol. 2010, 737385 (2010).
Gray, M.W. Mitochondrial evolution. Cold Spring Harb. Perspect. Biol. 4, a011403 (2012).
Zhang, X. et al. MicroRNA directly enhances mitochondrial translation during muscle differentiation. Cell 158, 607–619 (2014).
Kapp, L.D. & Lorsch, J.R. The molecular mechanics of eukaryotic translation. Annu. Rev. Biochem. 73, 657–704 (2004).
Dever, T.E. & Green, R. The elongation, termination, and recycling phases of translation in eukaryotes. Cold Spring Harb. Perspect. Biol. 4, a013706 (2012).
Qin, Y. et al. The highly conserved LepA is a ribosomal elongation factor that back-translocates the ribosome. Cell 127, 721–733 (2006).
Yamamoto, H. et al. EF-G and EF4: translocation and back-translocation on the bacterial ribosome. Nat. Rev. Microbiol. 12, 89–100 (2014).
Bauerschmitt, H., Funes, S. & Herrmann, J.M. The membrane-bound GTPase Guf1 promotes mitochondrial protein synthesis under suboptimal conditions. J. Biol. Chem. 283, 17139–17146 (2008).
Margus, T., Remm, M. & Tenson, T. Phylogenetic distribution of translational GTPases in bacteria. BMC Genomics 8, 15 (2007).
Han, B. & Qin, Y. Bioinformatics analysis reveals that LepA C-terminal domain is highly conserved in domain architectures and phylogenetic distribution. Scientia Sinica Chimica 42, 24–31 (2012).
Zhang, D. & Qin, Y. The paradox of elongation factor 4: highly conserved, yet of no physiological significance? Biochem. J. 452, 173–181 (2013).
Lomelí, H., Ramos-Mejía, V., Gertsenstein, M., Lobe, C.G. & Nagy, A. Targeted insertion of Cre recombinase into the TNAP gene: excision in primordial germ cells. Genesis 26, 116–117 (2000).
Wang, H. et al. Atg7 is required for acrosome biogenesis during spermatogenesis in mice. Cell Res. 24, 852–869 (2014).
Fernandez-Capetillo, O. et al. H2AX is required for chromatin remodeling and inactivation of sex chromosomes in male mouse meiosis. Dev. Cell 4, 497–508 (2003).
Naughton, C.K., Jain, S., Strickland, A.M., Gupta, A. & Milbrandt, J. Glial cell-line derived neurotrophic factor-mediated RET signaling regulates spermatogonial stem cell fate. Biol. Reprod. 74, 314–321 (2006).
Sun, F. & Handel, M.A. A mutation in Mtap2 is associated with arrest of mammalian spermatocytes before the first meiotic division. Genes (Basel) 2, 21–35 (2011).
Zhang, D. et al. EF4 disengages the peptidyl-tRNA CCA end and facilitates back-translocation on the 70S ribosome. Nat. Struct. Mol. Biol. 23, 125–131 (2016).
Liu, H., Pan, D., Pech, M. & Cooperman, B.S. Interrupted catalysis: the EF4 (LepA) effect on back-translocation. J. Mol. Biol. 396, 1043–1052 (2010).
Liu, H. et al. The conserved protein EF4 (LepA) modulates the elongation cycle of protein synthesis. Proc. Natl. Acad. Sci. USA 108, 16223–16228 (2011).
Hay, N. & Sonenberg, N. Upstream and downstream of mTOR. Genes Dev. 18, 1926–1945 (2004).
Wrobel, L. et al. Mistargeted mitochondrial proteins activate a proteostatic response in the cytosol. Nature 524, 485–488 (2015).
Morita, M. et al. mTORC1 controls mitochondrial activity and biogenesis through 4E-BP-dependent translational regulation. Cell Metab. 18, 698–711 (2013).
Cunningham, J.T. et al. mTOR controls mitochondrial oxidative function through a YY1-PGC-1α transcriptional complex. Nature 450, 736–740 (2007).
Düvel, K. et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol. Cell 39, 171–183 (2010).
Johnson, S.C. et al. mTOR inhibition alleviates mitochondrial disease in a mouse model of Leigh syndrome. Science 342, 1524–1528 (2013).
Mori, H. et al. Critical role for hypothalamic mTOR activity in energy balance. Cell Metab. 9, 362–374 (2009).
Yuan, H.X., Xiong, Y. & Guan, K.L. Nutrient sensing, metabolism, and cell growth control. Mol. Cell 49, 379–387 (2013).
Inoki, K., Kim, J. & Guan, K.L. AMPK and mTOR in cellular energy homeostasis and drug targets. Annu. Rev. Pharmacol. Toxicol. 52, 381–400 (2012).
Laplante, M. & Sabatini, D.M. mTOR signaling. Cold Spring Harb. Perspect. Biol. 4, a011593 (2012).
Laplante, M. & Sabatini, D.M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
De Martino, C. et al. Morphological, histochemical and biochemical studies on germ cell mitochondria of normal rats. Cell Tissue Res. 196, 1–22 (1979).
Conine, C.C. et al. Argonautes promote male fertility and provide a paternal memory of germline gene expression in C. elegans. Cell 155, 1532–1544 (2013).
Richter-Dennerlein, R., Dennerlein, S. & Rehling, P. Integrating mitochondrial translation into the cellular context. Nat. Rev. Mol. Cell Biol. 16, 586–592 (2015).
Dennerlein, S. & Rehling, P. Human mitochondrial COX1 assembly into cytochrome c oxidase at a glance. J. Cell Sci. 128, 833–837 (2015).
Mick, D.U., Fox, T.D. & Rehling, P. Inventory control: cytochrome c oxidase assembly regulates mitochondrial translation. Nat. Rev. Mol. Cell Biol. 12, 14–20 (2011).
Fernández-Vizarra, E. & Zeviani, M. Nuclear gene mutations as the cause of mitochondrial complex III deficiency. Front. Genet. 6, 134 (2015).
Stroud, D.A. & Ryan, M.T. Stalking the mitochondrial ATP synthase: Ina found guilty by association. EMBO J. 33, 1617–1618 (2014).
Mckenzie, M. & Ryan, M.T. Assembly factors of human mitochondrial complex I and their defects in disease. IUBMB Life 62, 497–502 (2010).
Mimaki, M., Wang, X., McKenzie, M., Thorburn, D.R. & Ryan, M.T. Understanding mitochondrial complex I assembly in health and disease. Biochim. Biophys. Acta 1817, 851–862 (2012).
Houtkooper, R.H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011).
Popow, J. et al. FASTKD2 is an RNA-binding protein required for mitochondrial RNA processing and translation. RNA 21, 1873–1884 (2015).
Tu, Y.T. & Barrientos, A. The human mitochondrial DEAD-box protein DDX28 resides in RNA granules and functions in mitoribosome assembly. Cell Rep. 10, 854–864 (2015).
Krause, W. Computer-assisted semen analysis systems: comparison with routine evaluation and prognostic value in male fertility and assisted reproduction. Hum. Reprod. 10 (suppl. 1), 60–66 (1995).
Boerboom, D. et al. β-catenin stabilization in gonadotropes impairs FSH synthesis in male mice in vivo. Endocrinology 156, 323–333 (2015).
Enders, G.C. & May, J.J. Developmentally regulated expression of a mouse germ cell nuclear antigen examined from embryonic day 11 to adult in male and female mice. Dev. Biol. 163, 331–340 (1994).
Wittig, I., Braun, H.P. & Schägger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Khvorostov, I., Zhang, J. & Teitell, M. Probing for mitochondrial complex activity in human embryonic stem cells. J. Vis. Exp. 16, e724 (2008).
McKee, E.E., Grier, B.L., Thompson, G.S. & McCourt, J.D. Isolation and incubation conditions to study heart mitochondrial protein synthesis. Am. J. Physiol. 258, E492–E502 (1990).
Acknowledgements
We are grateful to Y. Zhang, Q. Chen and X. Fu for help and discussions, X. Zhang for proteomics data analyses, L. Sun and Y. Jia for assistance with TEM, and T. Juelich for English editing. This work was supported by grants from the Major State Basic Research Development Program of China (2013CB531200 and 2012CB911000 to Y.Q.), the National Natural Science Foundation of China (31322015 and 31270847 to Y.Q.), the Institute of Biophysics 135 Goal-oriented Project, National Laboratory of Biomacromolecules (Institute of Biophysics, Chinese Academy of Sciences) to Y.Q., the Opening Project of the Zhejiang Provincial Top Key Discipline of Clinical Medicine (LKFJ009) to Y.Q. and the Shanghai Key Laboratory of Molecular Andrology, China to Y.Q. We thank F. Gao (Institute of Zoology, Beijing) for anti-GCNA1.
Author information
Authors and Affiliations
Contributions
Y.Q. conceived the project and designed the experiments. Y.G. and X.B. performed most experiments and processed and analyzed the data. D.Z., C.H., J.Y., W.L., X.C., Z.C., F.S., Z.Z., and F.G. assisted with experiments. Y.Q. wrote the manuscript, which was edited by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Generation and validation of mtEF4-knockout mice.
(a) Alignment of EF4 (E. coli) with mouse, yeast and human EF4. (b) Domain structures of mouse mtEF4 compared to those of EF4 (E. coli). (c) mtEF4 is ubiquitous in mice tissues and organs. (d) Schematic strategy for the generation of mtEF4 KO mouse. (e) Southern blot verification of a single introduction of the targeting construct in the homologous recombined embryonic stem (ES) cell clones. C: control, 129 ES cells. 1-3: positive clones. 5’-: 5’-primer. 3’-: 3’-primer. (f) PCR-based genotyping of KO and gKO mice. (g) qPCR of mtEF4 transcripts from various tissues of WT and KO mice (mean ± s.d., n = 12 mice). (h) Western blotting (WB) of mtEF4 protein from total WT (+/+) and KO (−/−) mice, and gKO mice.
Supplementary Figure 2 Morphology of mitochondria-rich tissues and hormones from WT and total-KO male mice.
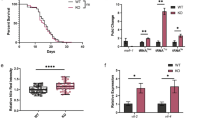
(a, b) Macrograph (a) and Hematoxylin and eosin (H&E) staining (b) of the heart. Scale bar: 200 μm (c) H&E staining of the liver, muscle and cerebrum. Scale bar: 200 μm (d) Quantification of the seminiferous tubule diameters (mean ± s.d., n = 5 mice, **P < 0.01). (e-g) Hormone changes upon mtEF4 ablation. Comparison of WT, KO and gKO mice serum follicle-stimulating hormone (FSH) (e), luteinizing hormone (LH) (f) and testosterone (T) (g) (mean ± s.d., n = 5 mice, *P < 0.05, **P < 0.01).
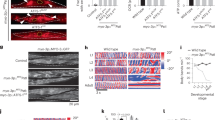
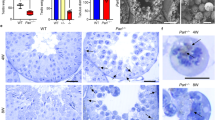
Supplementary Figure 3 Morphology of mitochondria-rich tissues from WT and gKO male mice.
(a-c) H&E staining of the heart, liver, muscle, cerebrum (a), testis (b) and epididymis (epi) (c) from WT and gKO mice. Scale bar: 200 μm. Red frame: zoom-in, scale bar: 50 μm. Red circle: immature spermatogenic cells. (d) The concentration of the spermatozoa in the epididymis. (e) Morphology of the sperms. Hoechst33258 (blue) and mitotracker (red) staining indicates nuclei and mitochondria, respectively. Abnormal mitochondrial sheaths are indicated by white arrows. Scale bar: 50 μm. (f, g) The mobility of the sperms. (h) Computer-assisted sperm analysis (CASA) parameter of the sperms. Average path velocity (VAP), straight line velocity (VSL), curvilinear velocity (VCL), amplitude of lateral head displacement (ALH), beat cross frequency (BCF) (mean ± s.d., n = 10 mice, ***P < 0.001). (i, j) EF4 is co-localized with the ribosome in the mitochondria of mouse testicular tissue. (k, l) Transmission electron microscope (TEM) photographs of gKO testis (k) and the sperms (l). Mt: mitochondria, Nu: nucleus, scale bar: 1 μm. Red frame: zoom-in. Scale bar: 1 μm.
Supplementary Figure 4 Qualitative and quantitative analysis of OXPHOS subunits from mitochondria-rich tissues of WT and KO male mice.
(a) Blue native gels (BNG) of heart, liver, muscle and cerebrum mitochondria. (b) In-gel activity (IGA) of complex I and IV. (c) Quantification of IGA from complex I (left) and complex IV (right) (mean ± s.d., n = 4 mice). (d) WB of nuclear and mitochondrial DNA encoded OXPHOS complex subunits. (e) Relative amount (%) of arbitrary units from samples in (d). Tom20: internal control, mean ± s.d., n = 5 mice.
Supplementary Figure 5 Qualitative and quantitative analysis of OXPHOS complexes and subunits from mitochondria-rich tissues of WT and gKO male mice.
(a, b) BNG (a) and IGA (b) of heart, liver, muscle and cerebrum mitochondria show similar amount of OXPHOS complexes in WT and gKO mice. (c) Quantification of IGA from complex I (left) and complex IV (right) (mean ± s.d., n = 4 mice). (d) IGA of OXPHOS complexes from WT and gKO heart, testis and epididymis. (e, f) WB of nDNA (e) and mtDNA (f) encoded OXPHOS complex subunits in the samples as in (d). (g, h) Relative amount (%) of arbitrary units from samples in (e) and (f), respectively. H: heart, T: tesits, E: epididymis, Tom20: internal control, mean ± s.d., n = 5 mice, **P < 0.01, ***P < 0.001.
Supplementary Figure 6 Quantification of mtDNA, mRNA of OXPHOS subunits, and mitochondrial ribosomes.
(a) Relative mtDNA content (mean ± s.d., n = 10 mice, *P < 0.05). (b) Relative quantification of nDNA (blue) and mtDNA (red) encoded OXPHOS subunits. The ratio was defined as the qPCR value of each tested transcript from KO sample versus from WT sample. Actin: internal control, mean ± s.d., n = 10 mice. (c) Quantification of the expression of mitochondrial transcription factor TFAM (mean ± s.d., n = 5 mice, *P < 0.05). (d) Quantification of the expression of mitochondrial small and large ribosomal subunit protein, MRPS18 and MRPL11, respectively. Tom20: internal control, mean ± s.d., n = 3 mice.
Supplementary Figure 7 [35S]methionine pulse-chase-labeling and protein synthesis of isolated mitochondria from WT and gKO testes.
Purified mitochondria were pulse labeled with [35S]-Met for 1 h and subsequently chased by the addition of an excess of unlabeled Met with 3 h and 5 h incubation in the presence (a) or absence (c) of protease inhibitor. The thirteen mitochondrial translation products are indicated. (b, d) The quantification of (a) and (c), respectively. Internal control: Tom20, mean ± s.d., n = 3 mice, ***P < 0.001.
Supplementary Figure 8 mTOR inhibition in the heart results in OXPHOS defects, and deletion of mtEF4 induces downregulation of cytoplasmic translation in sperm cells.
(a) Wet weight of the hearts from male mice treated with DMSO or rapamycin (mean ± s.d., n = 5 mice, **P < 0.01). (b) Quantification of the OXPHOS subunits of the DMSO/rapamycin treated mice. The quantitative value of the sample WT heart treated with DMSO was defined as 100%. Tom20: internal control, mean ± s.d., n = 3 mice, *P < 0.05, **P < 0.01. (c, d) Cytoplasmic polysome patterns of WT (blue) and gKO (red) tissues. On the right, the monosomes (80S) versus polysomes ratio was quantified (mean ± s.d., n = 3 mice, *P<0.05). (e, f) Distribution of ATP5D mRNAs across the density gradients from (c, d) was determined by RT-sqPCR. Red square, increased subunits and monosomes in gKO testis than in WT testis.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1 and 2 (PDF 4760 kb)
Supplementary Table 3
Variation in cellular signaling pathways (XLSX 661 kb)
Supplementary Data Set 1
Uncropped images for the main figures (PDF 1073 kb)
Rights and permissions
About this article
Cite this article
Gao, Y., Bai, X., Zhang, D. et al. Mammalian elongation factor 4 regulates mitochondrial translation essential for spermatogenesis. Nat Struct Mol Biol 23, 441–449 (2016). https://doi.org/10.1038/nsmb.3206
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3206
This article is cited by
-
Mitochondrial regulation during male germ cell development
Cellular and Molecular Life Sciences (2022)
-
Active RNA interference in mitochondria
Cell Research (2021)
-
A novel de novo MTOR gain-of-function variant in a patient with Smith-Kingsmore syndrome and Antiphospholipid syndrome
European Journal of Human Genetics (2019)
-
Current applications of antibody microarrays
Clinical Proteomics (2018)
-
PHA-4/FoxA senses nucleolar stress to regulate lipid accumulation in Caenorhabditis elegans
Nature Communications (2018)