Abstract
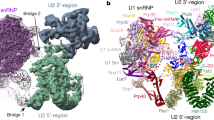
Aquarius is a multifunctional putative RNA helicase that binds precursor-mRNA introns at a defined position. Here we report the crystal structure of human Aquarius, revealing a central RNA helicase core and several unique accessory domains, including an ARM-repeat domain. We show that Aquarius is integrated into spliceosomes as part of a pentameric intron-binding complex (IBC) that, together with the ARM domain, cross-links to U2 snRNP proteins within activated spliceosomes; this suggests that the latter aid in positioning Aquarius on the intron. Aquarius's ARM domain is essential for IBC formation, thus indicating that it has a key protein-protein–scaffolding role. Finally, we provide evidence that Aquarius is required for efficient precursor-mRNA splicing in vitro. Our findings highlight the remarkable structural adaptations of a helicase to achieve position-specific recruitment to a ribonucleoprotein complex and reveal a new building block of the human spliceosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wahl, M.C., Will, C.L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Staley, J.P. & Guthrie, C. Mechanical devices of the spliceosome: motors, clocks, springs, and things. Cell 92, 315–326 (1998).
Bessonov, S. et al. Characterization of purified human Bact spliceosomal complexes reveals compositional and morphological changes during spliceosome activation and first step catalysis. RNA 16, 2384–2403 (2010).
Agafonov, D.E. et al. Semiquantitative proteomic analysis of the human spliceosome via a novel two-dimensional gel electrophoresis method. Mol. Cell. Biol. 31, 2667–2682 (2011).
Bleichert, F. & Baserga, S.J. The long unwinding road of RNA helicases. Mol. Cell 27, 339–352 (2007).
Pyle, A.M. Translocation and unwinding mechanisms of RNA and DNA helicases. Annu. Rev. Biophys. 37, 317–336 (2008).
Chakrabarti, S. et al. Molecular mechanisms for the RNA-dependent ATPase activity of Upf1 and its regulation by Upf2. Mol. Cell 41, 693–703 (2011).
Semlow, D.R. & Staley, J.P. Staying on message: ensuring fidelity in pre-mRNA splicing. Trends Biochem. Sci. 37, 263–273 (2012).
Fabrizio, P. et al. The evolutionarily conserved core design of the catalytic activation step of the yeast spliceosome. Mol. Cell 36, 593–608 (2009).
Fairman-Williams, M.E., Guenther, U.P. & Jankowsky, E. SF1 and SF2 helicases: family matters. Curr. Opin. Struct. Biol. 20, 313–324 (2010).
Bessonov, S., Anokhina, M., Will, C.L., Urlaub, H. & Luhrmann, R. Isolation of an active step I spliceosome and composition of its RNP core. Nature 452, 846–850 (2008).
Gozani, O., Feld, R. & Reed, R. Evidence that sequence-independent binding of highly conserved U2 snRNP proteins upstream of the branch site is required for assembly of spliceosomal complex A. Genes Dev. 10, 233–243 (1996).
Hirose, T. et al. A spliceosomal intron binding protein, IBP160, links position-dependent assembly of intron-encoded box C/D snoRNP to pre-mRNA splicing. Mol. Cell 23, 673–684 (2006).
Ideue, T., Sasaki, Y.T., Hagiwara, M. & Hirose, T. Introns play an essential role in splicing-dependent formation of the exon junction complex. Genes Dev. 21, 1993–1998 (2007).
Korneta, I., Magnus, M. & Bujnicki, J.M. Structural bioinformatics of the human spliceosomal proteome. Nucleic Acids Res. 40, 7046–7065 (2012).
Chang, Y.F., Imam, J.S. & Wilkinson, M.F. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74 (2007).
Tanner, N.K., Cordin, O., Banroques, J., Doere, M. & Linder, P. The Q motif: a newly identified motif in DEAD box helicases may regulate ATP binding and hydrolysis. Mol. Cell 11, 127–138 (2003).
Kuraoka, I. et al. Isolation of XAB2 complex involved in pre-mRNA splicing, transcription, and transcription-coupled repair. J. Biol. Chem. 283, 940–950 (2008).
Andrade, M.A., Petosa, C., O'Donoghue, S.I., Muller, C.W. & Bork, P. Comparison of ARM and HEAT protein repeats. J. Mol. Biol. 309, 1–18 (2001).
Tewari, R., Bailes, E., Bunting, K.A. & Coates, J.C. Armadillo-repeat protein functions: questions for little creatures. Trends Cell Biol. 20, 470–481 (2010).
Chan, S.P., Kao, D.I., Tsai, W.Y. & Cheng, S.C. The Prp19p-associated complex in spliceosome activation. Science 302, 279–282 (2003).
McGrail, J.C., Krause, A. & O'Keefe, R.T. The RNA binding protein Cwc2 interacts directly with the U6 snRNA to link the nineteen complex to the spliceosome during pre-mRNA splicing. Nucleic Acids Res. 37, 4205–4217 (2009).
Hogg, R., McGrail, J.C. & O'Keefe, R.T. The function of the NineTeen Complex (NTC) in regulating spliceosome conformations and fidelity during pre-mRNA splicing. Biochem. Soc. Trans. 38, 1110–1115 (2010).
Makarova, O.V. et al. A subset of human 35S U5 proteins, including Prp19, function prior to catalytic step 1 of splicing. EMBO J. 23, 2381–2391 (2004).
Villa, T. & Guthrie, C. The Isy1p component of the NineTeen complex interacts with the ATPase Prp16p to regulate the fidelity of pre-mRNA splicing. Genes Dev. 19, 1894–1904 (2005).
Baserga, S.J., Yang, X.D. & Steitz, J.A. An intact Box C sequence in the U3 snRNA is required for binding of fibrillarin, the protein common to the major family of nucleolar snRNPs. EMBO J. 10, 2645–2651 (1991).
Warkocki, Z. et al. Reconstitution of both steps of Saccharomyces cerevisiae splicing with purified spliceosomal components. Nat. Struct. Mol. Biol. 16, 1237–1243 (2009).
Trowitzsch, S., Bieniossek, C., Nie, Y., Garzoni, F. & Berger, I. New baculovirus expression tools for recombinant protein complex production. J. Struct. Biol. 172, 45–54 (2010).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Sheldrick, G.M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 479–485 (2010).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Dignam, J.D., Lebovitz, R.M. & Roeder, R.G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Fabrizio, P., Laggerbauer, B., Lauber, J., Lane, W.S. & Luhrmann, R. An evolutionarily conserved U5 snRNP-specific protein is a GTP-binding factor closely related to the ribosomal translocase EF-2. EMBO J. 16, 4092–4106 (1997).
Christian, H., Hofele, R.V., Urlaub, H. & Ficner, R. Insights into the activation of the helicase Prp43 by biochemical studies and structural mass spectrometry. Nucleic Acids Res. 42, 1162–1179 (2014).
Chen, Z.A. et al. Architecture of the RNA polymerase II-TFIIF complex revealed by cross-linking and mass spectrometry. EMBO J. 29, 717–726 (2010).
Leitner, A. et al. Expanding the chemical cross-linking toolbox by the use of multiple proteases and enrichment by size exclusion chromatography. Mol. Cell. Proteomics 11, M111 014126 (2012).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Acknowledgements
We are grateful to M. Raabe for assisting with peptide sequencing; K. Gencalp for help with multiangle light scattering measurements; A. Draycheva and M. Thommen for help with fluorescence anisotropy experiments; T.R. de Moura, J. Schmitzová, B. Kastner and K. Hartmuth for advice and helpful discussions; and the teams of beamlines 14.2 (BESSY, Berlin, Germany) and PXII (SLS, Villigen, Switzerland) for support during diffraction data collection.
Author information
Authors and Affiliations
Contributions
The crystallographic research, purification and biochemical characterization of recombinant proteins and the IBC were performed by I.D. under the supervision of V.P.; the biochemical experiments involving analysis of the IBC components in nuclear extracts, in vitro splicing, purification and analysis of spliceosomes were performed by S.B. under the supervision of R.L.; cross-linking experiments and MS analyses were performed by R.H. under the supervision of H.U. K.d.S. and C.L.W. participated in recombinant expression of Aquarius and in experiments with purified spliceosomes, respectively. All authors participated in the interpretation of the experiments and the writing of the paper. This study was supported by the Max-Planck-Society (V.P. and R.L.) and by the German Research Foundation (V.P., Deutsche Forschungsgemeinschaft grant PE 2079/2-1).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Purification of recombinant Aquarius and the IBC.
Related to Figure 2. (a) Purification of recombinant Aquarius. SDS-PAGE analysis of the peak fractions from anion-exchange (Q-Sepharose), affinity (Ni-NTA) and size exclusion (Superdex 200) chromatography is shown on the left. The band corresponding to Aquarius is indicated. Elution profiles of Aquarius (blue line) and molecular weight standards (dashed grey line with molecular masses in kDa) during size exclusion (Superdex 200) chromatography are shown on the right.
(b) Purification of the recombinant IBC. SDS-PAGE analysis of the peak fractions from affinity (Ni-NTA), anion-exchange (MonoQ) and size exclusion (Superose 6) chromatography is shown on the left. Bands corresponding to Aquarius, hSyf1, CCDC16, hIsy1 and CypE are indicated on the right. The IBC migrates as a sharp single peak in size exclusion (Superose 6) chromatography (right panel).
(c) Determination of the IBC's molar mass by multiangle light scattering. The molar mass versus the elution time is plotted. The calculated molar mass of IBC is 378 kDa.
Supplementary Figure 2 Structural features of Aquarius.
Related to Figure 1. (a) Structural comparison of Aquarius and Upf1 (PDB 2XZL, Chakrabarti et al., 2011). SF1 core domains are colored identically in both structures. Specific domains are colored grey.
(b) Comparison of the ARM domains of Aquarius (left) and β-catenin (right). The armadillo repeats (green and numbered), a helical protrusion (blue) and a helical extension (yellow) of the ARM domain are highlighted.
(c) The structure of Aquarius dissected in the middle along the stalk (blue). The two halves are rotated 90˚ either clockwise (left) or counterclockwise (right) to allow visualization of the stalk. The stalk is shown in ribbon representation.
(d) Comparison of the β-barrels of Aquarius (top) and Upf1 (bottom). The hydrophobic β-barrel core (magenta) is connected to helical insertions (blue) and loops (green).
Supplementary Figure 3 Sequence alignment of Aquarius orthologs and Upf1.
The amino acid sequence of Aquarius from Homo sapiens (H.s.), Danio rerio (D.r.), Drosophila melanogaster (D.m.), Caenorhabditis elegans (C.e.) Arabidopsis thaliana (A.t.) and Schizosaccharomyces pombe (S.p.) were aligned using Clustal Omega (Sievers et al., 2011). The amino acid sequence of Aquarius and human Upf1 were aligned by structural superposition. The PDB access code for the latter is 2GJK. Secondary structure elements are indicated above the sequences (α – alpha helices, β – beta strands, ŋ – 310 helices) and colored according to their domains (indicated at the bottom). Conserved helicase sequence motifs are indicated by black lines with Q or Roman numerals. Mutated residues are highlighted in magenta. Hydrophobic residues of the stalk involved in interactions with other domains are marked with red arrowheads.
Supplementary Figure 4 Purification and biochemical characterization of Aquarius lacking an ARM domain (ΔARM).
Related to Figure 3. (a) SDS-PAGE analysis of the peak fractions from size exclusion chromatography of the truncated Aquarius protein (ΔARM, 417-1485). The position of the protein is indicated on the right. Molecular weight markers (Mw, kDa) are indicated on the left.
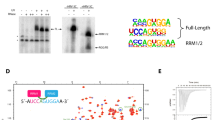
(b) Biochemical properties of ΔARM. Unwinding of a 3’ overhang RNA duplex (left) by truncated (ΔARM) versus full-length Aquarius was analysed by native PAGE. The positions of double- (ds) and single-stranded (ss) RNA are indicated. RNA binding (middle) was analyzed by an electrophoretic mobility shift assay. Positions of the free RNA and protein-RNA complexes are shown. ATPase activity (right) was analysed by thin layer chromatography. The position of ATP and inorganic phosphate (Pi) are indicated.
Supplementary Figure 5 Aquarius does not hydrolyze GTP.
Related to Figure 4. TLC analysis of the ATPase (lanes 1-3) and GTPase (lanes 4-6) activities of recombinant Aquarius in the presence or absence of ssRNA.
Supplementary Figure 6 Immunodepletion of Aquarius and IBC reduces splicing efficiency.
Related to Figure 6. Denaturing PAGE analysis (upper left) of splicing of PM5 pre-mRNA performed with mock-depleted or IBC immunodepleted (ΔIBC) nuclear extract in the presence or absence of purified recombinant IBC or buffer alone (as indicated above the lanes). The positions of the pre-mRNA and splicing intermediates are indicated on the right. The efficiency of the first step of splicing was calculated by dividing the sum of the intensities of the exon and intron lariat bands by the sum of the intensities of all three bands (pre-mRNA, exon, intron lariat) (bottom left). Bar diagram showing the efficiency of step I after 30 min of splicing normalized to the step I efficiency of mock-depleted extract supplemented with only buffer (set to 100%) (upper right). Values ± the standard deviation were determined from three independent experiments. The efficiency of IBC depletion was analysed by western blotting with anti-Aquarius antibodies (bottom right). Anti-U5-116K antibodies were used to demonstrate equal loading.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1 and 2 (PDF 12543 kb)
Supplementary Data Set 1
IBC proteins are cross-linked to U2 snRNP proteins in the spliceosome. (PDF 1053 kb)
Supplementary Data Set 2
Protein-protein interactions within the IBC. (PDF 1409 kb)
Supplementary Data Set 3
Protein composition of affinity-purified spliceosomes formed in the presence of the recombinant IBC. (PDF 826 kb)
Supplementary Data Set 4
Uncropped western blots and gels (PDF 820 kb)
Rights and permissions
About this article
Cite this article
De, I., Bessonov, S., Hofele, R. et al. The RNA helicase Aquarius exhibits structural adaptations mediating its recruitment to spliceosomes. Nat Struct Mol Biol 22, 138–144 (2015). https://doi.org/10.1038/nsmb.2951
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2951
This article is cited by
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)
-
Structural basis of catalytic activation in human splicing
Nature (2023)
-
A clue to the catalytic activation of splicing
Nature (2023)
-
Principles of RNA processing from analysis of enhanced CLIP maps for 150 RNA binding proteins
Genome Biology (2020)
-
DHX9 helicase promotes R-loop formation in cells with impaired RNA splicing
Nature Communications (2018)