Abstract
Template switching (TS) mediates damage bypass via a recombination-related mechanism involving PCNA polyubiquitination and polymerase δ–dependent DNA synthesis. Using two-dimensional gel electrophoresis and EM, here we characterize TS intermediates arising in Saccharomyces cerevisiae at a defined chromosome locus, identifying five major families of intermediates. Single-stranded DNA gaps of 150–200 nt, and not DNA ends, initiate TS by strand invasion. This causes reannealing of the parental strands and exposure of the nondamaged newly synthesized chromatid, which serves as a replication template for the other blocked nascent strand. Structures resembling double Holliday junctions, postulated to be central double-strand break–repair intermediates but so far visualized only in meiosis, mediate late stages of TS before being processed to hemicatenanes. Our results reveal the DNA transitions accounting for recombination-mediated DNA-damage tolerance in mitotic cells and replication under conditions of genotoxic stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Branzei, D. & Foiani, M. Maintaining genome stability at the replication fork. Nat. Rev. Mol. Cell Biol. 11, 208–219 (2010).
Weinert, T., Kaochar, S., Jones, H., Paek, A. & Clark, A.J. The replication fork's five degrees of freedom, their failure and genome rearrangements. Curr. Opin. Cell Biol. 21, 778–784 (2009).
Sale, J.E. Competition, collaboration and coordination: determining how cells bypass DNA damage. J. Cell Sci. 125, 1633–1643 (2012).
Zhang, H. & Lawrence, C.W. The error-free component of the RAD6/RAD18 DNA damage tolerance pathway of budding yeast employs sister-strand recombination. Proc. Natl. Acad. Sci. USA 102, 15954–15959 (2005).
Prakash, L. Characterization of postreplication repair in Saccharomyces cerevisiae and effects of rad6, rad18, rev3 and rad52 mutations. Mol. Gen. Genet. 184, 471–478 (1981).
Lehmann, A.R. Postreplication repair of DNA in ultraviolet-irradiated mammalian cells. J. Mol. Biol. 66, 319–337 (1972).
Branzei, D., Vanoli, F. & Foiani, M. SUMOylation regulates Rad18-mediated template switch. Nature 456, 915–920 (2008).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Stelter, P. & Ulrich, H.D. Control of spontaneous and damage-induced mutagenesis by SUMO and ubiquitin conjugation. Nature 425, 188–191 (2003).
Pfander, B., Moldovan, G.L., Sacher, M., Hoege, C. & Jentsch, S. SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase. Nature 436, 428–433 (2005).
Papouli, E. et al. Crosstalk between SUMO and ubiquitin on PCNA is mediated by recruitment of the helicase Srs2p. Mol. Cell 19, 123–133 (2005).
Minca, E.C. & Kowalski, D. Multiple Rad5 activities mediate sister chromatid recombination to bypass DNA damage at stalled replication forks. Mol. Cell 38, 649–661 (2010).
Haracska, L., Torres-Ramos, C.A., Johnson, R.E., Prakash, S. & Prakash, L. Opposing effects of ubiquitin conjugation and SUMO modification of PCNA on replicational bypass of DNA lesions in Saccharomyces cerevisiae. Mol. Cell. Biol. 24, 4267–4274 (2004).
Lopes, M., Foiani, M. & Sogo, J.M. Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol. Cell 21, 15–27 (2006).
Heller, R.C. & Marians, K.J. Replication fork reactivation downstream of a blocked nascent leading strand. Nature 439, 557–562 (2006).
Torres-Ramos, C.A., Prakash, S. & Prakash, L. Requirement of RAD5 and MMS2 for postreplication repair of UV-damaged DNA in Saccharomyces cerevisiae. Mol. Cell. Biol. 22, 2419–2426 (2002).
Vanoli, F., Fumasoni, M., Szakal, B., Maloisel, L. & Branzei, D. Replication and recombination factors contributing to recombination-dependent bypass of DNA lesions by template switch. PLoS Genet. 6, e1001205 (2010).
Torres-Ramos, C.A., Prakash, S. & Prakash, L. Requirement of yeast DNA polymerase δ in post-replicational repair of UV-damaged DNA. J. Biol. Chem. 272, 25445–25448 (1997).
Karras, G.I. & Jentsch, S. The RAD6 DNA damage tolerance pathway operates uncoupled from the replication fork and is functional beyond S phase. Cell 141, 255–267 (2010).
Liberi, G. et al. Rad51-dependent DNA structures accumulate at damaged replication forks in sgs1 mutants defective in the yeast ortholog of BLM RecQ helicase. Genes Dev. 19, 339–350 (2005).
Ashton, T.M., Mankouri, H.W., Heidenblut, A., McHugh, P.J. & Hickson, I.D. Pathways for Holliday junction processing during homologous recombination in Saccharomyces cerevisiae. Mol. Cell. Biol. 31, 1921–1933 (2011).
Szakal, B. & Branzei, D. Premature Cdk1/Cdc5/Mus81 pathway activation induces aberrant replication and deleterious crossover. EMBO J. 32, 1155–1167 (2013).
Mankouri, H.W., Ashton, T.M. & Hickson, I.D. Holliday junction-containing DNA structures persist in cells lacking Sgs1 or Top3 following exposure to DNA damage. Proc. Natl. Acad. Sci. USA 108, 4944–4949 (2011).
Osman, F. & Whitby, M.C. Exploring the roles of Mus81-Eme1/Mms4 at perturbed replication forks. DNA Repair (Amst.) 6, 1004–1017 (2007).
Sung, P. & Klein, H. Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nat. Rev. Mol. Cell Biol. 7, 739–750 (2006).
Branzei, D. Ubiquitin family modifications and template switching. FEBS Lett. 585, 2810–2817 (2011).
Ciccia, A. et al. Polyubiquitinated PCNA recruits the ZRANB3 translocase to maintain genomic integrity after replication stress. Mol. Cell 47, 396–409 (2012).
Blastyák, A. et al. Yeast Rad5 protein required for postreplication repair has a DNA helicase activity specific for replication fork regression. Mol. Cell 28, 167–175 (2007).
Burkovics, P., Sebesta, M., Balogh, D., Haracska, L. & Krejci, L. Strand invasion by HLTF as a mechanism for template switch in fork rescue. Nucleic Acids Res. 42, 1711–1720 (2014).
Glineburg, M.R., Chavez, A., Agrawal, V., Brill, S.J. & Johnson, F.B. Resolution by unassisted Top3 points to template switch recombination intermediates during DNA replication. J. Biol. Chem. 288, 33193–33204 (2013).
Lehmann, A.R. & Fuchs, R.P. Gaps and forks in DNA replication: rediscovering old models. DNA Repair (Amst.) 5, 1495–1498 (2006).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Tateishi, S., Sakuraba, Y., Masuyama, S., Inoue, H. & Yamaizumi, M. Dysfunction of human Rad18 results in defective postreplication repair and hypersensitivity to multiple mutagens. Proc. Natl. Acad. Sci. USA 97, 7927–7932 (2000).
Hishida, T., Kubota, Y., Carr, A.M. & Iwasaki, H. RAD6–RAD18–RAD5-pathway-dependent tolerance to chronic low-dose ultraviolet light. Nature 457, 612–615 (2009).
Daigaku, Y., Davies, A.A. & Ulrich, H.D. Ubiquitin-dependent DNA damage bypass is separable from genome replication. Nature 465, 951–955 (2010).
Alvaro, D., Lisby, M. & Rothstein, R. Genome-wide analysis of Rad52 foci reveals diverse mechanisms impacting recombination. PLoS Genet. 3, e228 (2007).
Mozlin, A.M., Fung, C.W. & Symington, L.S. Role of the Saccharomyces cerevisiae Rad51 paralogs in sister chromatid recombination. Genetics 178, 113–126 (2008).
Fabre, F., Chan, A., Heyer, W.D. & Gangloff, S. Alternate pathways involving Sgs1/Top3, Mus81/ Mms4, and Srs2 prevent formation of toxic recombination intermediates from single-stranded gaps created by DNA replication. Proc. Natl. Acad. Sci. USA 99, 16887–16892 (2002).
Runge, K.W. & Zakian, V.A. Introduction of extra telomeric DNA sequences into Saccharomyces cerevisiae results in telomere elongation. Mol. Cell. Biol. 9, 1488–1497 (1989).
Bzymek, M., Thayer, N.H., Oh, S.D., Kleckner, N. & Hunter, N. Double Holliday junctions are intermediates of DNA break repair. Nature 464, 937–941 (2010).
Neelsen, K.J., Chaudhuri, A.R., Follonier, C., Herrador, R. & Lopes, M. Visualization and interpretation of eukaryotic DNA replication intermediates in vivo by electron microscopy. Methods Mol. Biol. 1094, 177–208 (2014).
Cromie, G.A. et al. Single Holliday junctions are intermediates of meiotic recombination. Cell 127, 1167–1178 (2006).
Liberi, G. et al. Methods to study replication fork collapse in budding yeast. Methods Enzymol. 409, 442–462 (2006).
Lopes, M., Cotta-Ramusino, C., Liberi, G. & Foiani, M. Branch migrating sister chromatid junctions form at replication origins through Rad51/Rad52-independent mechanisms. Mol. Cell 12, 1499–1510 (2003).
Allers, T. & Lichten, M. A method for preparing genomic DNA that restrains branch migration of Holliday junctions. Nucleic Acids Res. 28, e6 (2000).
Joo, C., McKinney, S.A., Lilley, D.M. & Ha, T. Exploring rare conformational species and ionic effects in DNA Holliday junctions using single-molecule spectroscopy. J. Mol. Biol. 341, 739–751 (2004).
Cejka, P., Plank, J.L., Bachrati, C.Z., Hickson, I.D. & Kowalczykowski, S.C. Rmi1 stimulates decatenation of double Holliday junctions during dissolution by Sgs1–Top3. Nat. Struct. Mol. Biol. 17, 1377–1382 (2010).
Gonzalez-Huici, V. et al. DNA bending facilitates the error-free DNA damage tolerance pathway and upholds genome integrity. EMBO J. 33, 327–340 (2014).
Follonier, C., Oehler, J., Herrador, R. & Lopes, M. Friedreich's ataxia–associated GAA repeats induce replication-fork reversal and unusual molecular junctions. Nat. Struct. Mol. Biol. 20, 486–494 (2013).
Sogo, J.M., Lopes, M. & Foiani, M. Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science 297, 599–602 (2002).
Szostak, J.W., Orr-Weaver, T.L., Rothstein, R.J. & Stahl, F.W. The double-strand-break repair model for recombination. Cell 33, 25–35 (1983).
Karras, G.I. et al. Noncanonical role of the 9-1-1 clamp in the error-free DNA damage tolerance pathway. Mol. Cell 49, 536–546 (2013).
Wu, L. & Hickson, I.D. The Bloom's syndrome helicase suppresses crossing over during homologous recombination. Nature 426, 870–874 (2003).
Ira, G., Malkova, A., Liberi, G., Foiani, M. & Haber, J.E. Srs2 and Sgs1-Top3 suppress crossovers during double-strand break repair in yeast. Cell 115, 401–411 (2003).
Bugreev, D.V., Yu, X., Egelman, E.H. & Mazin, A.V. Novel pro- and anti-recombination activities of the Bloom's syndrome helicase. Genes Dev. 21, 3085–3094 (2007).
Hu, Y. et al. RECQL5/Recql5 helicase regulates homologous recombination and suppresses tumor formation via disruption of Rad51 presynaptic filaments. Genes Dev. 21, 3073–3084 (2007).
Robert, T., Dervins, D., Fabre, F. & Gangloff, S. Mrc1 and Srs2 are major actors in the regulation of spontaneous crossover. EMBO J. 25, 2837–2846 (2006).
Gangloff, S., McDonald, J.P., Bendixen, C., Arthur, L. & Rothstein, R. The yeast type I topoisomerase Top3 interacts with Sgs1, a DNA helicase homolog: a potential eukaryotic reverse gyrase. Mol. Cell. Biol. 14, 8391–8398 (1994).
Branzei, D. et al. Ubc9- and Mms21-mediated sumoylation counteracts recombinogenic events at damaged replication forks. Cell 127, 509–522 (2006).
Sollier, J. et al. The Saccharomyces cerevisiae Esc2 and Smc5–6 proteins promote sister chromatid junction-mediated intra-S repair. Mol. Biol. Cell 20, 1671–1682 (2009).
Acknowledgements
This work was supported by European Research Council (ERC), Associazione Italiana per la Ricerca sul Cancro (AIRC) and Fondazione Telethon grants to D.B., Swiss National Science Foundation grants PP00P3_135292 and 31003A_146924 to K.Z., C.F. and M.L., and AIRC and Fondazione Telethon grants to M.F. We thank the Center for Microscopy and Image Analysis of the University of Zürich for technical assistance with the EM experiments, I. Psakhye for critical reading of the manuscript, W. Carotenuto at IFOM for initial assistance in drawing the model, members of our laboratories for helpful discussions and FIRC for various support.
Author information
Authors and Affiliations
Contributions
M.G. designed and executed the experiments, acquired the EM images, analyzed the data and made the figures. K.Z. and C.F. acquired a subset of EM images and helped with EM data analysis. M.F. conceived the project and discussed the results. M.L. conceived the project, supervised the EM part, analyzed the EM data and commented on the manuscript. D.B. conceived and supervised the project, designed the experiments, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Features of the minichromosome YLpFAT7.1 and minichromosome-derived template switch–intermediate families.
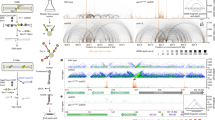
(a) Schematic representation of the minichromosome YLpFAT7.1 with the location of the leu2d probe. (b) Undigested or BglII digested genomic DNA samples from wild-type (wt, CY11340), sgs1Δ (CY11357), rad51Δ (CY12388), sgs1Δ rad51Δ (CY12390) strains carrying the minichromosome YLpFAT7.1 were analyzed by gel electrophoresis. The minichromosome was visualized by Southern blotting using the leu2d probe. (c) Genomic DNA from wt (CY11340) and sgs1Δ (CY11357) strains was digested with a restriction enzymes mix containing PacI, XhoI, SgrAI, BssHII, SpeI, BclI and PmlI, and an aliquot of the digestion mix was examined on a first dimension gel (0.35% agarose TBE 1X). After ethidium bromide staining, the minichromosome band is visible at the expected molecular weight and the endogenous genomic DNA is cleaved to an average length smaller than 5kb. (d) Number and percentage of X-shaped DNA structures found in the wt and sgs1Δ samples that were assigned as minichromosome-derived structures. (e) Box plot showing the distribution of the ssDNA lengths in F1*-F1-F2 molecules in wt and sgs1Δ cells. Center line, median; box limits, 25th and 75th percentiles; whiskers, 10th and 90th percentiles; dots, outliers; (n), number of samples (molecules) in the data set. The median (M) and average (A) values derived from calculations are displayed. NS, P> 0.05 by t-test with two tails (f) Chart representing the distribution of the JM families in wt and sgs1Δ cells.
Supplementary Figure 2 Representative EM pictures of F1 family of molecules.
(a-c) Total views (left panels) of representative F1 family DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points and red arrows the ssDNA discontinuities on one branch in immediate proximity to the junction point. Schematic representations of the junction points are shown with a black/grey code to indicate the contributions of the DNA duplexes to the junction and with a black/red code to indicate the ssDNA regions in red and dsDNA regions in black. Scale bars are shown.
Supplementary Figure 3 Representative EM pictures of F2 family of molecules.
(a-c) Total views (left panels) of representative F2 family of DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points, which have an extended region of homology (double Y-like structures). Black arrows mark the thick DNA filament at the junction point that we infer to be three-stranded for the reasons explained in the main text. Red arrows mark the ssDNA discontinuity on one or two branches of the joint molecule in immediate proximity to the junction point. Black asterisks mark random crosses between DNA filaments. Schematic representations of the junction points are shown with black/grey and black/red codes as in Supplementary Figure 2. Scale bars are shown.
Supplementary Figure 4 Representative EM pictures of F3 family of molecules.
(a-c) Total views (left panels) of representative F3 family of DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points, which have an extended region of homology (bubble-like structures). Lengths of the dsDNA filaments at the junction points of F3 molecules are reported in white in kilobases. Black asterisks mark random crosses between DNA filaments. Schematic representations of the junction points are shown with black/grey and black/red codes as in Supplementary Figure 2. Scale bars are shown.
Supplementary Figure 5 Representative EM pictures of F3* family of molecules.
(a-c) Total views (left panels) of representative F3* family of DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points, which have an extended region of homology (bubble-like structures), and red arrows mark the ssDNA discontinuity on one of the dsDNA filaments present at the junction point. Lengths of the DNA filaments at the junction points are reported in white in kilobases. Schematic representations of the junction points are shown with black/grey and black/red codes as in Supplementary Figure 2. Scale bars are shown.
Supplementary Figure 6 Representative EM pictures of F4 family of molecules.
(a-c) Total views (left panels) of representative F4 family of DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points, which have an extended region of homology (double Y-like structures). Schematic representations of the junction points are shown with black/grey color code as in Supplementary Figure 2. Scale bars are shown.
Supplementary Figure 7 Representative EM pictures of F5 family of molecules.
(a-c) Total views (left panels) of representative F5 family of DNA structures and enlarged views (right panels) of the junction points. Branch sizes until the junction point are reported in kilobases. Two colors, grey and black, in which the length values are displayed, mark the “equal” arms. White arrows mark the junction points and red arrows the ssDNA regions. Schematic representations of the junction points are shown with black/grey and black/red codes as in Supplementary Figure 2. Scale bars are shown.
Supplementary Figure 8 Representative EM pictures of single Holliday junctions and asymmetric F5* and F1* types of molecules.
(a) A representative EM picture of a single HJ-like molecule (sHJ) with a total view of the DNA structure and an enlarged view of the junction point with a schematic representation of the ssDNA regions reported in red is shown. Branch sizes until the junction point are reported in black in kilobases. Schematic representation of the junction point is also shown with a black/grey code to indicate the contributions of two DNA duplexes to the junction. Scale bars are shown. (b-c) Total views of X-shaped DNA structures of F5* family (b) or F1* family (c) and enlarged views of the junction points with schematic representations of the ssDNA regions in red. Branch sizes until the junction point are reported in kilobases. Two colors, blue and black, used to indicate the length values mark the arms belonging to the same DNA duplex involved in the junction. Schematic representations of the junction points are shown with black/grey and black/red codes as in Supplementary Figure 2. Scale bars are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 14969 kb)
Rights and permissions
About this article
Cite this article
Giannattasio, M., Zwicky, K., Follonier, C. et al. Visualization of recombination-mediated damage bypass by template switching. Nat Struct Mol Biol 21, 884–892 (2014). https://doi.org/10.1038/nsmb.2888
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2888
This article is cited by
-
Rad51-mediated replication of damaged templates relies on monoSUMOylated DDK kinase
Nature Communications (2022)
-
BRCA1 deficiency specific base substitution mutagenesis is dependent on translesion synthesis and regulated by 53BP1
Nature Communications (2022)
-
Delayed DNA break repair for genome stability
Nature Cell Biology (2021)
-
Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites
Nature Communications (2021)
-
Participation of the HIM1 gene of yeast Saccharomyces cerevisiae in the error-free branch of post-replicative repair and role Polη in him1-dependent mutagenesis
Current Genetics (2021)