Abstract
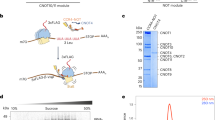
The CCR4–NOT deadenylase complex is a master regulator of translation and mRNA stability. Its NOT module orchestrates recruitment of the catalytic subunits to target mRNAs. We report the crystal structure of the human NOT module formed by the CNOT1, CNOT2 and CNOT3 C-terminal (-C) regions. CNOT1-C provides a rigid scaffold consisting of two perpendicular stacks of HEAT-like repeats. CNOT2-C and CNOT3-C heterodimerize through their SH3-like NOT-box domains. The heterodimer is stabilized and tightly anchored to the surface of CNOT1 through an unexpected intertwined arrangement of peptide regions lacking defined secondary structure. These assembly peptides mold onto their respective binding surfaces and form extensive interfaces. Mutagenesis of individual interfaces and perturbation of endogenous protein ratios cause defects in complex assembly and mRNA decay. Our studies provide a structural framework for understanding the recruitment of the CCR4–NOT complex to mRNA targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others

References
Collart, M.A. & Panasenko, O.O. The Ccr4-Not complex. Gene 492, 42–53 (2012).
Wahle, E. & Winkler, G.S. RNA decay machines: deadenylation by the Ccr4-Not and Pan2-Pan3 complexes. Biochim. Biophys. Acta 1829, 561–570 (2013).
Bawankar, P., Loh, B., Wohlbold, L., Schmidt, S. & Izaurralde, E. NOT10 and C2orf29/NOT11 form a conserved module of the CCR4-NOT complex that docks onto the NOT1 N-terminal domain. RNA Biol. 10, 228–244 (2013).
Chekulaeva, M. et al. miRNA repression involves GW182-mediated recruitment of CCR4–NOT through conserved W-containing motifs. Nat. Struct. Mol. Biol. 18, 1218–1226 (2011).
Cooke, A., Prigge, A. & Wickens, M. Translational repression by deadenylases. J. Biol. Chem. 285, 28506–28513 (2010).
Zekri, L., Kuzuoğlu-Öztürk, D. & Izaurralde, E. GW182 proteins cause PABP dissociation from silenced miRNA targets in the absence of deadenylation. EMBO J. 32, 1052–1065 (2013).
Bai, Y. et al. The CCR4 and CAF1 proteins of the CCR4-NOT complex are physically and functionally separated from NOT2, NOT4, and NOT5. Mol. Cell. Biol. 19, 6642–6651 (1999).
Lau, N.C. et al. Human Ccr4-Not complexes contain variable deadenylase subunits. Biochem. J. 422, 443–453 (2009).
Liu, H.Y. et al. The NOT proteins are part of the CCR4 transcriptional complex and affect gene expression both positively and negatively. EMBO J. 17, 1096–1106 (1998).
Basquin, J. et al. Architecture of the nuclease module of the yeast Ccr4-Not complex: the Not1-Caf1-Ccr4 interaction. Mol. Cell 48, 207–218 (2012).
Ito, K., Takahashi, A., Morita, M., Suzuki, T. & Yamamoto, T. The role of the CNOT1 subunit of the CCR4–NOT complex in mRNA deadenylation and cell viability. Protein Cell 2, 755–763 (2011).
Maillet, L., Tu, C., Hong, Y.K., Shuster, E.O. & Collart, M.A. The essential function of Not1 lies within the Ccr4-Not complex. J. Mol. Biol. 303, 131–143 (2000).
Mauxion, F., Prève, B. & Séraphin, B. C2ORF29/CNOT11 and CNOT10 form a new module of the CCR4-NOT complex. RNA Biol. 10, 267–276 (2013).
Petit, A.P. et al. The structural basis for the interaction between CAF1 nuclease and the NOT1 scaffold of the human CCR4-NOT deadenylase complex. Nucleic Acids Res. 40, 11058–11072 (2012).
Russell, P., Benson, J.D. & Denis, C.L. Characterization of mutations in NOT2 indicates that it plays an important role in maintaining the integrity of the CCR4-NOT complex. J. Mol. Biol. 322, 27–39 (2002).
Winkler, G.S., Mulder, K.W., Bardwell, V.G., Kalkhoven, E. & Timmers, H.T. Human Ccr4-Not complex is a ligand-dependent repressor of nuclear receptor-mediated transcription. EMBO J. 25, 3089–3099 (2006).
Barckmann, B. & Simonelig, M. Control of maternal mRNA stability in germ cells and early embryos. Biochim. Biophys. Acta 1829, 714–724 (2013).
Braun, J.E., Huntzinger, E., Fauser, M. & Izaurralde, E. GW182 proteins recruit cytoplasmic deadenylase complexes to miRNA targets. Mol. Cell 44, 120–133 (2011).
Fabian, M.R. et al. miRNA-mediated deadenylation is orchestrated by GW182 through two conserved motifs that interact with CCR4–NOT. Nat. Struct. Mol. Biol. 18, 1211–1217 (2011).
Huntzinger, E. et al. The interactions of GW182 proteins with PABP and deadenylases are required for both translational repression and degradation of miRNA targets. Nucleic Acids Res. 41, 978–994 (2013).
Fabian, M.R. et al. Structural basis for the recruitment of the human CCR4–NOT deadenylase complex by tristetraprolin. Nat. Struct. Mol. Biol. 20, 735–739 (2013).
Albert, T.K. et al. Isolation and characterization of human orthologs of yeast CCR4-NOT complex subunits. Nucleic Acids Res. 28, 809–817 (2000).
Zwartjes, C.G., Jayne, S., van den Berg, D.L. & Timmers, H.T. Repression of promoter activity by CNOT2, a subunit of the transcription regulatory Ccr4-Not complex. J. Biol. Chem. 279, 10848–10854 (2004).
Andrade, M.A., Petosa, C., O'Donoghue, S.I., Müller, C.W. & Bork, P. Comparison of ARM and HEAT protein repeats. J. Mol. Biol. 309, 1–18 (2001).
Temme, C. et al. Subunits of the Drosophila CCR4-NOT complex and their roles in mRNA deadenylation. RNA 16, 1356–1370 (2010).
Ito, K. et al. CNOT2 depletion disrupts and inhibits the CCR4–NOT deadenylase complex and induces apoptotic cell death. Genes Cells 16, 368–379 (2011).
Brodersen, D.E. & Nissen, P. The social life of ribosomal proteins. FEBS J. 272, 2098–2108 (2005).
Klein, D.J., Moore, P.B. & Steitz, T.A. The roles of ribosomal proteins in the structure assembly, and evolution of the large ribosomal subunit. J. Mol. Biol. 340, 141–177 (2004).
Demarest, S.J. et al. Mutual synergistic folding in recruitment of CBP/p300 by p160 nuclear receptor coactivators. Nature 415, 549–553 (2002).
Tompa, P. Intrinsically disordered proteins: a 10-year recap. Trends Biochem. Sci. 37, 509–516 (2012).
Dyson, H.J. & Wright, P.E. Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell Biol. 6, 197–208 (2005).
Prakash, S., Tian, L., Ratliff, K.S., Lehotzky, R.E. & Matouschek, A. An unstructured initiation site is required for efficient proteasome-mediated degradation. Nat. Struct. Mol. Biol. 11, 830–837 (2004).
Temme, C., Zaessinger, S., Meyer, S., Simonelig, M. & Wahle, E. A complex containing the CCR4 and CAF1 proteins is involved in mRNA deadenylation in Drosophila. EMBO J. 23, 2862–2871 (2004).
Chicoine, J. et al. Bicaudal-C recruits CCR4-NOT deadenylase to target mRNAs and regulates oogenesis, cytoskeletal organization, and its own expression. Dev. Cell 13, 691–704 (2007).
Suzuki, A., Igarashi, K., Aisaki, K., Kanno, J. & Saga, Y. NANOS2 interacts with the CCR4-NOT deadenylation complex and leads to suppression of specific RNAs. Proc. Natl. Acad. Sci. USA 107, 3594–3599 (2010).
Igreja, C. & Izaurralde, E. CUP promotes deadenylation and inhibits decapping of mRNA targets. Genes Dev. 25, 1955–1967 (2011).
Leppek, K. et al. Roquin promotes constitutive mRNA decay via a conserved class of stem-loop recognition motifs. Cell 153, 869–881 (2013).
Diebold, M.-L., Fribourg, S., Koch, M., Metzger, T. & Romier, C. Deciphering correct strategies for multiprotein complex assembly by co-expression: application to complexes as large as the histone octamer. J. Struct. Biol. 175, 178–188 (2011).
Doublié, S. Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530 (1997).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Sheldrick, G.M. Experimental phasing with SHELXC/D/E: combining chaintracing with density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 479–485 (2010).
Terwilliger, T.C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D Biol. Crystallogr. 65, 582–601 (2009).
Cowtan, K. The Buccaneer software for automated model building. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Zhang, K.Y.J., Cowtan, K. & Main, P. Combining constraints for electron-density modification. Methods Enzymol. 277, 53–64 (1997).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Behm-Ansmant, I., Gatfield, D., Rehwinkel, J., Hilgers, V. & Izaurralde, E. A conserved role for cytoplasmic poly(A)-binding protein 1 (PABPC1) in nonsense-mediated mRNA decay. EMBO J. 26, 1591–1601 (2007).
Acknowledgements
We are grateful to J. Su (Max Planck Institute for Developmental Biology) for providing the cDNA for the CNOT1 fragment (residues 1565–2371) and to E. Wahle (Martin Luther University Halle-Wittenberg) for the kind gift of anti–Dm NOT1–NOT3 antibodies. This work was supported by the Max Planck Society, by grants from the Deutsche Forschungsgemeinschaft (DFG, FOR855 and the Gottfried Wilhelm Leibniz Program awarded to E.I.) and the European Union Seventh Framework Program through a Marie Curie Fellowship to S.J. (FP7, 275343).
Author information
Authors and Affiliations
Contributions
A.B., Y.C., T.R. and S.J. contributed equally to this work. A.B. and Y.C. purified, crystallized and solved the structures of CNOT3 and CNOT2 oligomers and of Ct NOT1. A.B. and Y.C. cloned, expressed and established the purification protocol for the ternary complex. T.R. purified and crystallized the ternary complex. T.R. and S.J. solved the structure of the ternary complex. A.B., Y.C., T.R., S.J. and O.W. collected and analyzed diffraction data. L.W. performed pulldowns and coimmunoprecipitations in human cells. D.K.-Ö. performed coimmunoprecipitations and functional assays in S2 cells. E.I. conceived of the project. E.I. and O.W. supervised the project. All authors contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CNOT1, CNOT2 and CNOT3 interact via their C-termini.
(a,b) Interaction between GFP-tagged CNOT2 (full-length or fragments) and HA-tagged CNOT1 (a) and CNOT3 (b). GFP-tagged MBP served as a negative control. In all panels, cell lysates were treated with RNase A. (c,d) Interaction between GFP-tagged CNOT3 (full-length or fragments) and HA-tagged CNOT2 (c) and CNOT1 (d). (e,f) Interaction between GFP-tagged CNOT1 (full-length or fragments) and HA-tagged CNOT2 (e) and CNOT3 (f). In panel (f), the interaction with CNOT3 was analyzed in the absence (lanes 2 and 6) or presence (lanes 4 and 8) of CNOT2. (g) Interaction between MBP-CNOT2 fragments and His6-tagged CNOT1 (residues 1565–2371). (h) Interaction between MBP-CNOT2 fragments and GST-CNOT3 (residues 589–753). (i) Interaction between MBP-CNOT2 (residues 344–540) and GST-CNOT3 fragments. (j) Interaction between MBP-CNOT1 (residues 1595–2376) and GST-CNOT2 (residues 344–540) or GST-CNOT3 (residues 589–753).
Supplementary Figure 2 Structure-based sequence alignment of the NOT1 superfamily homology (SH) domain.
Secondary structure elements as determined from the Hs CNOT1 structure are shown above the alignment. Residues conserved in all aligned sequences are shown with a yellow background and residues with >70% similarity are highlighted in orange. Residues interacting with CNOT2 and CNOT3 are indicated by purple and green dots, respectively. Residues involved in CNOT1 subdomain interactions are indicated by blue dots. Residues mutated in this study are marked by red asterisks. Residues substituted in mutant M1 and M5 are indicated. The species abbreviations are as follows: Hs (Homo sapiens), Dm (Drosophila melanogaster), Xt (Xenopus tropicalis), Ce (Caenorhabditis elegans), Ct (Chaetomium thermophilum) and Sc (Sacchromyces cerevisiae).
Supplementary Figure 3 CNOT2 and CNOT3 multimerize in solution.
(a,d) MALLS analysis of the CNOT3 and CNOT2 NOT-box domains. The molecular weight of the protein in solution is indicated in the elution profile. (b) Interfaces between the CNOT3 NOT-box α-helices and the β-barrel. Residues along the interfaces are shown as sticks. (c) Hydrophobic core of the CNOT3 NOT-box β-barrel. (e) Structure of the CNOT2 tetramer as observed in the crystal. The tetramer consists of two pairs of dimers (orange-yellow vs. purple-rose) with a perpendicular orientation to each other. (f) Structure-based sequence alignments of the NOT2 and NOT3 NOT-box domains. Secondary structure elements as determined from the CNOT2 and CNOT3 structures are shown above the alignment. Residues conserved in all aligned sequences are shown with a yellow background, and residues with >70% similarity are highlighted in orange. A NOT2-specific insertion is boxed in purple. Black squares mark residues that form the interface between the NOT-Box N-terminal α-helices and the β-barrel. Gray squares mark residues that form the hydrophobic core of the β-barrel. The species abbreviations are the same as those in Supplementary Fig. 2. (g) Pulldowns showing that CNOT2-C and CNOT3-C exclusively form heterodimers in solution. MBP-tagged CNOT2 and His6-tagged CNOT3 were coexpressed in E. coli. The heterodimers were copurified using MBP pulldown followed by Ni-affinity purification (lanes 1 and 2) or Ni-affinity purification followed by MBP pulldown (lanes 3 and 4).
Supplementary Figure 4 Structure of the NOT module.
(a) Time course of a limited proteolysis of the ternary complex between CNOT1 (residues 1565–2371), CNOT2 (residues 344–540) and CNOT3 (residues 607–753) by thermolysin. CNOT1 is digested into a stable C-terminal fragment corresponding to the SH and an N-terminal fragment (which comigrates with CNOT2), while CNOT2 and CNOT3 remain uncleaved. (b) Time course of a limited proteolysis of CNOT2 (residues 344–540) and CNOT3 (residues 607–753) by thermolysin in the absence or presence of CNOT1 SH (1833–2361). In the absence of CNOT1, CNOT2 and CNOT3 are digested into stable C-terminal fragments corresponding approximately to the NOT-boxes and the CSs, but they remain largely uncleaved in the presence of CNOT1 (c,d) Omit electron density map of the CNOT2 (c) and CNOT3 (d) NAR-Cs folded onto the CNOT1 surface. The electron density (black mesh, 2F0-FC of a composite omit map, calculated with Phenix.AutoBuild) is contoured at 1.0 σ. (e,f) Refined electron density of the CNOT2 (e) and CNOT3 (f) NAR-C regions in stereo view. The views correspond to panels (c) and (d), respectively. The electron density (black mesh, 2F0-FC map) is contoured at 1.0 σ. (g) Superposition of Ct NOT1 (blue) and Hs CNOT1 (gray) showing strong structure conservation. (h) Superposition of the CNOT3 NOT-box domain in isolation (gray) and in the complex (green). (i) Superposition of the NOT-box domains of CNOT2 (purple) and CNOT3 (green) as observed in the ternary complex.
Supplementary Figure 5 Structure-based sequence alignment of the CNOT2 and CNOT3.
Secondary structure elements as determined from the CNOT2 and CNOT3 structure are shown above the alignment. Symbols are as described in Supplementary Fig. 2. Residues interacting with the respective partner proteins are indicated by dots above the alignments and are colored blue for CNOT1, purple for CNOT2 and green for CNOT3. Residues forming the lock are marked with black diamonds, and residues at the junction between the symmetric and asymmetric lobes of the ternary complex are marked with pink triangles. The species abbreviations are the same as those in Supplementary Fig. 2.
Supplementary Figure 6 Surface conservation of the trimeric complex.
(a–e) The conservation scores of the individual residues are represented on the surface by color gradients from light (no conservation) to dark colors (100% conservation) for CNOT1 (blue), CNOT2 (purple) and CNOT3 (green). Conservation scores were calculated based on well-balanced multiple alignments covering all eukaryotic strata. (a) View from the CNOT1 surface that binds CNOT2 and CNOT3 (a) or the opposite surface (b). Conservation of the CNOT1 surface contacting CNOT2 and CNOT3 (c). The view is the same as that shown in Fig. 5a. (d) Conservation of CNOT2 surface residues contacting the CNOT3 connector sequence (CS). The CNOT3 residues involved in the interaction with CNOT2 are shown as sticks. (e) Conservation of CNOT3 surface contacting the CNOT2 connector sequence (CS). The CNOT2 residues involved in CNOT3 binding are shown as sticks.
Supplementary Figure 7 Mutagenesis of the NOT1-NOT2-NOT3 interfaces in human and Dm S2 cells.
(a) Interaction of GFP-CNOT1 (either wild-type or the indicated mutants) with endogenous CNOT3 and HA-CNOT2. (b,c) Interaction of GFP-CNOT1 (either wild-type or the indicated mutants) with HA-CNOT7 in human cells. (d) The decay of the adh-hsp70 mRNA was monitored in control cells (expressing MBP) and in cells expressing NOT1 (either wild-type or mutant). Adh-hsp70 mRNA levels were normalized to the levels of long-lived rp49 mRNA and plotted against time. A representative Northern blot is shown in Fig. 7a. The mRNA half-lives (t1/2) ± standard deviations calculated from the decay curves obtained from three independent experiments are indicated. (e) Interaction of GFP-tagged Dm CAF1 with wild-type NOT1 or NOT1ΔN-SD in S2 cells. F-Luc-GFP served as a negative control. (f,g) The decay of adh-hsp70 mRNA was analyzed in control cells (treated with GFP dsRNA and expressing MBP) or in cells depleted of NOT1 or NOT3 and expressing MBP or the indicated proteins. Northern blots corresponding to the decay curves are shown in Fig. 7c and 7d, respectively. Adh-hsp70 mRNA levels were normalized to the levels of rp49 mRNA and plotted against time. The mRNA half-lives (t1/2) ± standard deviations obtained from three independent experiments are indicated.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 22785 kb)
Rights and permissions
About this article
Cite this article
Boland, A., Chen, Y., Raisch, T. et al. Structure and assembly of the NOT module of the human CCR4–NOT complex. Nat Struct Mol Biol 20, 1289–1297 (2013). https://doi.org/10.1038/nsmb.2681
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2681
This article is cited by
-
Regulation by the RNA-binding protein Unkempt at its effector interface
Nature Communications (2024)
-
Structure and assembly of the NOT10:11 module of the CCR4-NOT complex
Communications Biology (2023)
-
RNF219 regulates CCR4-NOT function in mRNA translation and deadenylation
Scientific Reports (2022)
-
The functional characterization of phosphorylation of tristetraprolin at C-terminal NOT1-binding domain
Journal of Inflammation (2021)
-
CAPRI enables comparison of evolutionarily conserved RNA interacting regions
Nature Communications (2019)