Abstract
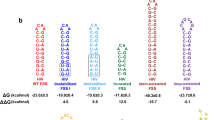
During protein synthesis, the ribosome translates nucleotide triplets in single-stranded mRNA into polypeptide sequences. Strong downstream mRNA secondary structures, which must be unfolded for translation, can slow or even halt protein synthesis. Here we used single-molecule fluorescence resonance energy transfer to determine reaction rates for specific steps within the elongation cycle as the Escherichia coli ribosome encounters stem-loop or pseudoknot mRNA secondary structures. Downstream stem-loops containing 100% GC base pairs decrease the rates of both tRNA translocation within the ribosome and deacylated tRNA dissociation from the ribosomal exit site (E site). Downstream stem-loops or pseudoknots containing both GC and AU pairs also decrease the rate of tRNA dissociation, but they have little effect on tRNA translocation rate. Thus, somewhat unexpectedly, unfolding of mRNA secondary structures is more closely coupled to E-site tRNA dissociation than to tRNA translocation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Zhang, G., Hubalewska, M. & Ignatova, Z. Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nat. Struct. Mol. Biol. 16, 274–280 (2009).
Guisez, Y., Robbens, J., Remaut, E. & Fiers, W. Folding of the Ms2 coat protein in Escherichia coli is modulated by translational pauses resulting from messenger-RNA secondary structure and codon usage: a hypothesis. J. Theor. Biol. 162, 243–252 (1993).
Varenne, S., Buc, J., Lloubes, R. & Lazdunski, C. Translation is a non-uniform process: effect of transfer-RNA availability on the rate of elongation of nascent polypeptide-chains. J. Mol. Biol. 180, 549–576 (1984).
Kimchi-Sarfaty, C. et al. A “silent” polymorphism in the MDR1 gene changes substrate specificity. Science 315, 525–528 (2007).
Namy, O., Moran, S.J., Stuart, D.I., Gilbert, R.J.C. & Brierley, I. A mechanical explanation of RNA pseudoknot function in programmed ribosomal frameshifting. Nature 441, 244–247 (2006).
Giedroc, D.P. & Cornish, P.V. Frameshifting RNA pseudoknots: structure and mechanism. Virus Res. 139, 193–208 (2009).
Gurvich, O.L. et al. Sequences that direct significant levels of frameshifting are frequent in coding regions of Escherichia coli. EMBO J. 22, 5941–5950 (2003).
Brierley, I., Meredith, M.R., Bloys, A.J. & Hagervall, T.G. Expression of a coronavirus ribosomal frameshift signal in Escherichia coli: influence of tRNA anticodon modification on frameshifting. J. Mol. Biol. 270, 360–373 (1997).
Brierley, I. & Dos Ramos, F.J. Programmed ribosomal frameshifting in HIV-1 and the SARS-CoV. Virus Res. 119, 29–42 (2006).
Yusupova, G.Z., Yusupov, M.M., Cate, J.H.D. & Noller, H.F. The path of messenger RNA through the ribosome. Cell 106, 233–241 (2001).
Takyar, S., Hickerson, R.P. & Noller, H.F. mRNA helicase activity of the ribosome. Cell 120, 49–58 (2005).
Spahn, C.M.T. et al. Structure of the 80S ribosome from Saccharomyces cerevisiae tRNA-ribosome and subunit-subunit interactions. Cell 107, 373–386 (2001).
Barbara, P.F. Single-molecule spectroscopy. Acc. Chem. Res. 38, 503 (2005).
Qu, X. et al. The ribosome uses two active mechanisms to unwind messenger RNA during translation. Nature 475, 118–121 (2011).
Wen, J.D. et al. Following translation by single ribosomes one codon at a time. Nature 452, 598–603 (2008).
Brierley, I., Rolley, N.J., Jenner, A.J. & Inglis, S.C. Mutational analysis of the RNA pseudoknot component of a coronavirus ribosomal frameshifting signal. J. Mol. Biol. 220, 889–902 (1991).
Somogyi, P., Jenner, A.J., Brierley, I. & Inglis, S.C. Ribosomal pausing during translation of an RNA pseudoknot. Mol. Cell Biol. 13, 6931–6940 (1993).
Chen, C. et al. Single-molecule fluorescence measurements of ribosomal translocation dynamics. Mol. Cell 42, 367–377 (2011).
Chen, C. et al. Allosteric vs. spontaneous exit-site (E-site) tRNA dissociation early in protein synthesis. Proc. Natl. Acad. Sci. USA 108, 16980–16985 (2011).
Stevens, B. et al. FRET-based identification of mRNAs undergoing translation. PLoS ONE 7, e38344 (2012).
Agirrezabala, X. et al. Visualization of the hybrid state of tRNA binding promoted by spontaneous ratcheting of the ribosome. Mol. Cell 32, 190–197 (2008).
Julian, P. et al. Structure of ratcheted ribosomes with tRNAs in hybrid states. Proc. Natl. Acad. Sci. USA 105, 16924–16927 (2008).
Frank, J. & Gonzalez, R.L. Structure and dynamics of a processive Brownian motor: The translating ribosome. Annu. Rev. Biochem. 79, 381–412 (2010).
Munro, J.B., Sanbonmatsu, K.Y., Spahn, C.M.T. & Blanchard, S.C. Navigating the ribosome's metastable energy landscape. Trends Biochem. Sci. 34, 390–400 (2009).
Fei, J., Kosuri, P., MacDougall, D.D. & Gonzalez, R.L. Coupling of ribosomal L1 stalk and tRNA dynamics during translation elongation. Mol. Cell 30, 348–359 (2008).
Munro, J.B. et al. Spontaneous formation of the unlocked state of the ribosome is a multistep process. Proc. Natl. Acad. Sci. USA 107, 709–714 (2010).
Cornish, P.V. et al. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl. Acad. Sci. USA 106, 2571–2576 (2009).
Giedroc, D.P., Theimer, C.A. & Nixon, P.L. Structure, stability and function of RNA pseudoknots involved in stimulating ribosomal frameshifting. J. Mol. Biol. 298, 167–185 (2000).
Rinnenthal, J., Klinkert, B., Narberhaus, F. & Schwalbe, H. Direct observation of the temperature-induced melting process of the Salmonella fourU RNA thermometer at base-pair resolution. Nucleic Acids Res. 38, 3834–3847 (2010).
Pan, D., Kirillov, S.V. & Cooperman, B.S. Kinetically competent intermediates in the translocation step of protein synthesis. Mol. Cell 25, 519–529 (2007).
Kaur, J., Raj, M. & Cooperman, B.S. Fluorescent labeling of tRNA dihydrouridine residues: mechanism and distribution. RNA 17, 1393–1400 (2011).
Liu, H., Pan, D.L., Pech, M. & Cooperman, B.S. Interrupted catalysis: the EF4 (LepA) effect on back-translocation. J. Mol. Biol. 396, 1043–1052 (2010).
Savelsbergh, A. et al. An elongation factor G-induced ribosome rearrangement precedes tRNA-mRNA translocation. Mol. Cell 11, 1517–1523 (2003).
Ramrath, D.J.F. et al. The complex of tmRNA-SmpB and EF-G on translocating ribosomes. Nature 485, 526–529 (2012).
Ratje, A.H. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010).
Ermolenko, D.N. & Noller, H.F. mRNA translocation occurs during the second step of ribosomal intersubunit rotation. Nat. Struct. Mol. Biol. 18, 457–462 (2011).
Rodnina, M.V. & Wintermeyer, W. GTP consumption of elongation factor Tu during translation of heteropolymeric messenger RNAs. Proc. Natl. Acad. Sci. USA 92, 1945–1949 (1995).
Subramanian, A.R. & Dabbs, E.R. Functional studies on ribosomes lacking protein L1 from mutant Escherichia coli. Eur. J. Biochem. 112, 425–430 (1980).
Pan, D.L., Qin, H.O. & Cooperman, B.S. Synthesis and functional activity of tRNAs labeled with fluorescent hydrazides in the D-loop. RNA 15, 346–354 (2009).
Chiu, J., March, P.E., Lee, R. & Tillett, D. Site-directed, ligase-independent mutagenesis (SLIM): a single-tube methodology approaching 100% efficiency in 4 h. Nucleic Acids Res. 32, e174 (2004).
Odom, O.W. et al. Distances between 3′ ends of ribosomal ribonucleic acids reassembled into Escherichia coli ribosomes. Biochemistry 19, 5947–5954 (1980).
Kapanidis, A.N. et al. Fluorescence-aided molecule sorting: analysis of structure and interactions by alternating-laser excitation of single molecules. Proc. Natl. Acad. Sci. USA 101, 8936–8941 (2004).
McKinney, S.A., Joo, C. & Ha, T. Analysis of single-molecule FRET trajectories using hidden Markov modeling. Biophys. J. 91, 1941–1951 (2006).
Acknowledgements
This work was supported by US National Institutes of Health grant R01GM080376 to B.S.C. and Y.E.G. and American Heart Association Postdoctoral Fellowship 12POST8910014 to C.C.
Author information
Authors and Affiliations
Contributions
C.C., B.S.C. and Y.E.G. designed the experiments. C.C., S.L.B. and M.R. conducted the experiments and analyzed the data. H.Z. and I.F. prepared reagents. C.C., B.S.C. and Y.E.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–4 (PDF 2507 kb)
Rights and permissions
About this article
Cite this article
Chen, C., Zhang, H., Broitman, S. et al. Dynamics of translation by single ribosomes through mRNA secondary structures. Nat Struct Mol Biol 20, 582–588 (2013). https://doi.org/10.1038/nsmb.2544
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2544
This article is cited by
-
Riboformer: a deep learning framework for predicting context-dependent translation dynamics
Nature Communications (2024)
-
Specific length and structure rather than high thermodynamic stability enable regulatory mRNA stem-loops to pause translation
Nature Communications (2022)
-
Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Nature Communications (2022)
-
Diversity and prevalence of ANTAR RNAs across actinobacteria
BMC Microbiology (2021)
-
Octa-repeat domain of the mammalian prion protein mRNA forms stable A-helical hairpin structure rather than G-quadruplexes
Scientific Reports (2019)