Abstract
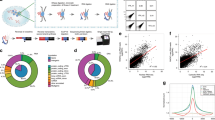
The exon junction complex (EJC) is a central effector of the fate of mRNAs, linking nuclear processing to mRNA transport, translation and surveillance. However, little is known about its transcriptome-wide targets. We used cross-linking and immunoprecipitation methods coupled to high-throughput sequencing (CLIP-seq) in human cells to identify the binding sites of the DEAD-box helicase eIF4AIII, an EJC core component. CLIP reads form peaks that are located mainly in spliced mRNAs. Most expressed exons harbor peaks either in the canonical EJC region, located ~24 nucleotides upstream of exonic junctions, or in other noncanonical regions. Notably, both of these types of peaks are preferentially associated with unstructured and purine-rich sequences containing the motif GAAGA, which is a potential binding site for EJC-associated factors. Therefore, EJC positions vary spatially and quantitatively between exons. This transcriptome-wide mapping of human eIF4AIII reveals unanticipated aspects of the EJC and broadens its potential impact on post-transcriptional regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Moore, M.J. & Proudfoot, N.J. Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 136, 688–700 (2009).
Le Hir, H., Izaurralde, E., Maquat, L.E. & Moore, M.J. The spliceosome deposits multiple proteins 20–24 nucleotides upstream of mRNA exon-exon junctions. EMBO J. 19, 6860–6869 (2000).
Tange, T.Ø., Nott, A. & Moore, M.J. The ever-increasing complexities of the exon junction complex. Curr. Opin. Cell Biol. 16, 279–284 (2004).
Le Hir, H. & Andersen, G.R. Structural insights into the exon junction complex. Curr. Opin. Struct. Biol. 18, 112–119 (2008).
Andersen, C.B. et al. Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science 313, 1968–1972 (2006).
Ballut, L. et al. The exon junction core complex is locked onto RNA by inhibition of eIF4AIII ATPase activity. Nat. Struct. Mol. Biol. 12, 861–869 (2005).
Bono, F., Ebert, J., Lorentzen, E. & Conti, E. The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell 126, 713–725 (2006).
Nielsen, K.H. et al. Mechanism of ATP turnover inhibition in the EJC. RNA 15, 67–75 (2009).
Gehring, N.H., Lamprinaki, S., Hentze, M.W. & Kulozik, A.E. The hierarchy of exon-junction complex assembly by the spliceosome explains key features of mammalian nonsense-mediated mRNA decay. PLoS Biol. 7, e1000120 (2009).
Merz, C., Urlaub, H., Will, C.L. & Luhrmann, R. Protein composition of human mRNPs spliced in vitro and differential requirements for mRNP protein recruitment. RNA 13, 116–128 (2007).
Reichert, V.L., Le Hir, H., Jurica, M.S. & Moore, M.J. 5′ exon interactions within the human spliceosome establish a framework for exon junction complex structure and assembly. Genes Dev. 16, 2778–2791 (2002).
Zhang, Z. & Krainer, A.R. Splicing remodels messenger ribonucleoprotein architecture via eIF4A3-dependent and -independent recruitment of exon junction complex components. Proc. Natl. Acad. Sci. USA 104, 11574–11579 (2007).
Barbosa, I. et al. Human CWC22 escorts the helicase eIF4AIII to spliceosomes and promotes exon junction complex assembly. Nat. Struct. Mol. Biol. 19, 983–990 (2012).
Gehring, N.H., Lamprinaki, S., Kulozik, A.E. & Hentze, M.W. Disassembly of exon junction complexes by PYM. Cell 137, 536–548 (2009).
Hwang, J. & Maquat, L.E. Nonsense-mediated mRNA decay (NMD) in animal embryogenesis: to die or not to die, that is the question. Curr. Opin. Genet. Dev. 21, 422–430 (2011).
Rebbapragada, I. & Lykke-Andersen, J. Execution of nonsense-mediated mRNA decay: what defines a substrate? Curr. Opin. Cell Biol. 21, 394–402 (2009).
Isken, O. & Maquat, L.E. The multiple lives of NMD factors: balancing roles in gene and genome regulation. Nat. Rev. Genet. 9, 699–712 (2008).
Saulière, J. et al. The exon junction complex differentially marks spliced junctions. Nat. Struct. Mol. Biol. 17, 1269–1271 (2010).
Ashton-Beaucage, D. et al. The exon junction complex controls the splicing of MAPK and other long intron-containing transcripts in Drosophila. Cell 143, 251–262 (2010).
Michelle, L. et al. Proteins associated with the exon junction complex also control the alternative splicing of apoptotic regulators. Mol. Cell. Biol. 32, 954–967 (2012).
Roignant, J.Y. & Treisman, J.E. Exon junction complex subunits are required to splice Drosophila MAP kinase, a large heterochromatic gene. Cell 143, 238–250 (2010).
Hachet, O. & Ephrussi, A. Splicing of oskar RNA in the nucleus is coupled to its cytoplasmic localization. Nature 428, 959–963 (2004).
Haremaki, T., Sridharan, J., Dvora, S. & Weinstein, D.C. Regulation of vertebrate embryogenesis by the exon junction complex core component Eif4a3. Dev. Dyn. 239, 1977–1987 (2010).
Silver, D.L. et al. The exon junction complex component Magoh controls brain size by regulating neural stem cell division. Nat. Neurosci. 13, 551–558 (2010).
Albers, C.A. et al. Compound inheritance of a low-frequency regulatory SNP and a rare null mutation in exon-junction complex subunit RBM8A causes TAR syndrome. Nat. Genet. 44, 435–439 (2012).
Darnell, R.B. HITS-CLIP: panoramic views of protein-RNA regulation in living cells. Wiley Interdiscip Rev. RNA 1, 266–286 (2010).
Ule, J., Jensen, K., Mele, A. & Darnell, R.B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Zhang, C. & Darnell, R.B. Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data. Nat. Biotechnol. 29, 607–614 (2011).
Fejes, A.P. et al. FindPeaks 3.1: a tool for identifying areas of enrichment from massively parallel short-read sequencing technology. Bioinformatics 24, 1729–1730 (2008).
Patel, A.A. & Steitz, J.A. Splicing double: insights from the second spliceosome. Nat. Rev. Mol. Cell Biol. 4, 960–970 (2003).
Hirose, T., Shu, M.D. & Steitz, J.A. Splicing of U12-type introns deposits an exon junction complex competent to induce nonsense-mediated mRNA decay. Proc. Natl. Acad. Sci. USA 101, 17976–17981 (2004).
Dostie, J. & Dreyfuss, G. Translation is required to remove Y14 from mRNAs in the cytoplasm. Curr. Biol. 12, 1060–1067 (2002).
Lejeune, F., Ishigaki, Y., Li, X. & Maquat, L.E. The exon junction complex is detected on CBP80-bound but not eIF4E-bound mRNA in mammalian cells: dynamics of mRNP remodeling. EMBO J. 21, 3536–3545 (2002).
Dennis, G. Jr. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, 3 (2003).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sanford, J.R. et al. Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts. Genome Res. 19, 381–394 (2009).
Bailey, T.L., Boden, M., Whitington, T. & Machanick, P. The value of position-specific priors in motif discovery using MEME. BMC Bioinformatics 11, 179 (2010).
Long, J.C. & Caceres, J.F. The SR protein family of splicing factors: master regulators of gene expression. Biochem. J. 417, 15–27 (2009).
Markham, N.R. & Zuker, M. DINAMelt web server for nucleic acid melting prediction. Nucleic Acids Res. 33, W577–W581 (2005).
Markham, N.R. & Zuker, M. UNAFold: software for nucleic acid folding and hybridization. Methods Mol. Biol. 453, 3–31 (2008).
Mishler, D.M., Christ, A.B. & Steitz, J.A. Flexibility in the site of exon junction complex deposition revealed by functional group and RNA secondary structure alterations in the splicing substrate. RNA 14, 2657–2670 (2008).
Budiman, M.E. et al. Eukaryotic initiation factor 4a3 is a selenium-regulated RNA-binding protein that selectively inhibits selenocysteine incorporation. Mol. Cell 35, 479–489 (2009).
Le Hir, H. & Seraphin, B. EJCs at the heart of translational control. Cell 133, 213–216 (2008).
Nagy, E. & Maquat, L.E. A rule for termination-codon position within intron-containing genes: when nonsense affects RNA abundance. Trends Biochem. Sci. 23, 198–199 (1998).
Bühler, M., Paillusson, A. & Muhlemann, O. Efficient downregulation of immunoglobulin mu mRNA with premature translation-termination codons requires the 5′-half of the VDJ exon. Nucleic Acids Res. 32, 3304–3315 (2004).
Carter, M.S., Li, S. & Wilkinson, M.F. A splicing-dependent regulatory mechanism that detects translation signals. EMBO J. 15, 5965–5975 (1996).
Holbrook, J.A., Neu-Yilik, G., Hentze, M.W. & Kulozik, A.E. Nonsense-mediated decay approaches the clinic. Nat. Genet. 36, 801–808 (2004).
Viegas, M.H., Gehring, N.H., Breit, S., Hentze, M.W. & Kulozik, A.E. The abundance of RNPS1, a protein component of the exon junction complex, can determine the variability in efficiency of the nonsense mediated decay pathway. Nucleic Acids Res. 35, 4542–4551 (2007).
Zetoune, A.B. et al. Comparison of nonsense-mediated mRNA decay efficiency in various murine tissues. BMC Genet. 9, 83 (2008).
Zhang, Z. & Krainer, A.R. Involvement of SR proteins in mRNA surveillance. Mol. Cell 16, 597–607 (2004).
Daguenet, E. et al. Perispeckles are major assembly sites for the exon junction core complex. Mol. Biol. Cell 23, 1765–1782 (2012).
Acknowledgements
We thank C. Tomasetto and J. Stévenin (Institut de Génétique et de Biologie Moléculaire et Cellulaire, Illkirch, France) and J. Marie (Centre de Génétique Moléculaire, Gif-sur-Yvette, France) and E. Izaurralde (Max Planck Institute, Tübingen, Germany) for antibodies, C. Thermes and E. van Dijk (Centre de Génétique Moléculaire, Gif-sur-Yvette, France) at the IMAGIF platform for Illumina deep sequencing and J. Ule (Medical Research Council Laboratory of Molecular Biology, Cambridge, UK) for technical advice. We acknowledge our laboratory for helpful advice, comments and discussions and A. Morillon (Institut Curie, Paris, France) for carefully reading the manuscript. We also thank M. Gratigny and A. Louis (Institut de Biologie de l'Ecole Normale Supérieure, Paris, France) for bioinformatic advice. This work was supported in part by the Centre National de la Recherche Scientifique (ATIP programme blanc 2008 to H.L.H. and ANR-07-JCJC-0097-01 to L.P.), the Institut National de la Santé et de la Recherche Médicale (INSERM; J.S.), the Agence Nationale de la Recherche (2008-BLAN-0323 to H.L.H.) and the Fondation pour la Recherche Médicale (J.S. and H.L.H.). Z.W. is supported by a European Molecular Biology Organization long-term postdoctoral fellowship.
Author information
Authors and Affiliations
Contributions
O.L.T., Y.A., L.P. and J.S. contributed to the development of the CLIP protocol. J.S. performed CLIP experiments. V.M. and H.R.C. performed the bioinformatic analyses. Z.W. performed RIP experiments. E.M. performed protein coimmunoprecitation assays. I.B. purified the antibodies. J.S. and H.L.H. conceived the project. H.L.H. provided resources. J.S. and H.L.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Note (PDF 1709 kb)
Rights and permissions
About this article
Cite this article
Saulière, J., Murigneux, V., Wang, Z. et al. CLIP-seq of eIF4AIII reveals transcriptome-wide mapping of the human exon junction complex. Nat Struct Mol Biol 19, 1124–1131 (2012). https://doi.org/10.1038/nsmb.2420
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2420
This article is cited by
-
Comprehensive mapping of exon junction complex binding sites reveals universal EJC deposition in Drosophila
BMC Biology (2023)
-
Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Nature Communications (2023)
-
Blood-derived lncRNAs as biomarkers for cancer diagnosis: the Good, the Bad and the Beauty
npj Precision Oncology (2022)
-
Exon junction complex shapes the m6A epitranscriptome
Nature Communications (2022)
-
Nanopore sequencing reveals endogenous NMD-targeted isoforms in human cells
Genome Biology (2021)