Abstract
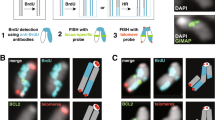
Common fragile sites have been mapped primarily in lymphocytes, but recent analyses show that the setting of these sites relies on cell type–dependent replication programs. Using a new approach, we molecularly mapped common fragile sites in human fibroblasts and showed that commitment to fragility depends on similar replication features in fibroblasts and lymphocytes, although different loci are committed in each cell type. Notably, the common fragile sites that we identified overlapped heretofore unexplained deletion clusters observed in tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Durkin, S.G. & Glover, T.W. Annu. Rev. Genet. 41, 169–192 (2007).
Bignell, G.R. et al. Nature 463, 893–898 (2010).
Negrini, S., Gorgoulis, V.G. & Halazonetis, T.D. Nat. Rev. Mol. Cell Biol. 11, 220–228 (2010).
Iliopoulos, D. et al. Cancer Lett. 232, 27–36 (2006).
El Achkar, E., Gerbault-Seureau, M., Muleris, M., Dutrillaux, B. & Debatisse, M. Proc. Natl. Acad. Sci. USA 102, 18069–18074 (2005).
Chan, K.L., Palmai-Pallag, T., Ying, S. & Hickson, I.D. Nat. Cell Biol. 11, 753–760 (2009).
Naim, V. & Rosselli, F. Nat. Cell Biol. 11, 761–768 (2009).
Letessier, A. et al. Nature 470, 120–123 (2011).
Lukas, C. et al. Nat. Cell Biol. 13, 243–253 (2011).
Méchali, M. Nat. Rev. Mol. Cell Biol. 11, 728–738 (2010).
Ryba, T. et al. Genome Res. 20, 761–770 (2010).
Hansen, R.S. et al. Proc. Natl. Acad. Sci. USA 107, 139–144 (2010).
Smith, D.I., McAvoy, S., Zhu, Y. & Perez, D.S. Semin. Cancer Biol. 17, 31–41 (2007).
Helmrich, A., Stout-Weider, K., Hermann, K., Schrock, E. & Heiden, T. Genome Res. 16, 1222–1230 (2006).
Pasic, I. et al. Cancer Res. 70, 160–171 (2010).
Gilbert, D.M. et al. Cold Spring Harb. Symp. Quant. Biol. 75, 143–153 (2010).
Acknowledgements
We thank R. Rothstein, A. Letessier and H. Técher for critical reading of the manuscript. The M.D. team is supported by Institut National du Cancer (INCa) (2009-1-PLBIO-10-IC-1), by Agence Nationale de la Recherche (ANR-09-GENO-000/repinsCFS) and by Association pour la Recherche sur le Cancer (Subvention Libre n° SL220100601348 and Equipements mi-lourds n° 8514). B.L.T. is supported by a fellowship from INCa.
Author information
Authors and Affiliations
Contributions
B.D. conducted the R-banding analyses; B.L.T., A.M.L. and O.B. carried out and analyzed the FISH experiments; G.A.M. did the Repli-Seq analyses; B.L.T., O.B. and M.D. wrote the manuscript; M.D. and O.B. planned the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2, Supplementary Tables 1–3 and Supplementary Methods (PDF 481 kb)
Rights and permissions
About this article
Cite this article
Le Tallec, B., Dutrillaux, B., Lachages, AM. et al. Molecular profiling of common fragile sites in human fibroblasts. Nat Struct Mol Biol 18, 1421–1423 (2011). https://doi.org/10.1038/nsmb.2155
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2155
This article is cited by
-
Genome-wide high-resolution mapping of mitotic DNA synthesis sites and common fragile sites by direct sequencing
Cell Research (2020)
-
Twin peaks: finding fragile sites with MiDAS-seq
Cell Research (2020)
-
3D genome organization contributes to genome instability at fragile sites
Nature Communications (2020)
-
High-resolution mapping of mitotic DNA synthesis regions and common fragile sites in the human genome through direct sequencing
Cell Research (2020)
-
The prevention and resolution of DNA replication–transcription conflicts in eukaryotic cells
Genome Instability & Disease (2020)