Abstract
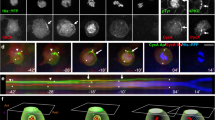
Frizzled planar cell polarity (PCP) signaling regulates cell motility in several tissues, including ommatidial rotation in Drosophila melanogaster. The Nemo kinase (Nlk in vertebrates) has also been linked to cell-motility regulation and ommatidial rotation but its mechanistic role(s) during rotation remain obscure. We show that nemo functions throughout the entire rotation movement, increasing the rotation rate. Genetic and molecular studies indicate that Nemo binds both the core PCP factor complex of Strabismus–Prickle, as well as the E-cadherin–β-catenin (E-cadherin–Armadillo in Drosophila) complex. These two complexes colocalize and, like Nemo, also promote rotation. Strabismus (also called Vang) binds and stabilizes Nemo asymmetrically within the ommatidial precluster; Nemo and β-catenin then act synergistically to promote rotation, which is mediated in vivo by Nemo's phosphorylation of β-catenin. Our data suggest that Nemo serves as a conserved molecular link between core PCP factors and E-cadherin–β-catenin complexes, promoting cell motility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wolff, T. & Ready, D.F. Pattern formation in the Drosophila retina. in The Development of Drosophila melanogaster (ed. Martinez-Arias, M.B.A.) 1277–1326 (Cold Spring Harbor Press, 1993).
Mlodzik, M. Planar polarity in the Drosophila eye: a multifaceted view of signaling specificity and cross-talk. EMBO J. 18, 6873–6879 (1999).
Seifert, J.R. & Mlodzik, M. Frizzled/PCP signalling: a conserved mechanism regulating cell polarity and directed motility. Nat. Rev. Genet. 8, 126–138 (2007).
Wang, Y. & Nathans, J. Tissue/planar cell polarity in vertebrates: new insights and new questions. Development 134, 647–658 (2007).
Mirkovic, I. & Mlodzik, M. Cooperative activities of Drosophila DE-cadherin and DN-cadherin regulate the cell motility process of ommatidial rotation. Development 133, 3283–3293 (2006).
Solnica-Krezel, L. Conserved patterns of cell movements during vertebrate gastrulation. Curr. Biol. 15, R213–R228 (2005).
Choi, K.-W. & Benzer, S. Rotation of photoreceptor clusters in the developing Drosophila eye requires the nemo gene. Cell 78, 125–136 (1994).
Fiehler, R.W. & Wolff, T. Nemo is required in a subset of photoreceptors to regulate the speed of ommatidial rotation. Dev. Biol. 313, 533–544 (2008).
Winter, C.G. et al. Drosophila Rho-associated kinase (Drok) links Frizzled-mediated planar cell polarity signaling to the actin cytoskeleton. Cell 105, 81–91 (2001).
Brown, K.E. & Freeman, M. Egfr signalling defines a protective function for ommatidial orientation in the Drosophila eye. Development 130, 5401–5412 (2003).
Gaengel, K. & Mlodzik, M. Egfr signaling regulates ommatidial rotation and cell motility in the Drosophila eye via MAPK/Pnt signaling and the Ras effector Canoe/AF6. Development 130, 5413–5423 (2003).
Strutt, H. & Strutt, D. EGF signaling and ommatidial rotation in the Drosophila eye. Curr. Biol. 13, 1451–1457 (2003).
Chou, Y.H. & Chien, C.T. Scabrous controls ommatidial rotation in the Drosophila compound eye. Dev. Cell 3, 839–850 (2002).
Fiehler, R.W. & Wolff, T. Drosophila Myosin II, Zipper, is essential for ommatidial rotation. Dev. Biol. 310, 348–362 (2007).
Classen, A.K., Anderson, K.I., Marois, E. & Eaton, S. Hexagonal packing of Drosophila wing epithelial cells by the planar cell polarity pathway. Dev. Cell 9, 805–817 (2005).
Takeichi, M. Cadherins: a molecular family important in selective cell-cell adhesion. Annu. Rev. Biochem. 59, 237–252 (1990).
Tepass, U. et al. shotgun encodes Drosophila E-cadherin and is preferentially required during cell rearrangement in the neurectoderm and other morphogenetically active epithelia. Genes Dev. 10, 672–685 (1996).
Gumbiner, B.M. Regulation of cadherin-mediated adhesion in morphogenesis. Nat. Rev. Mol. Cell Biol. 6, 622–634 (2005).
Chen, Y.T., Stewart, D.B. & Nelson, W.J. Coupling assembly of the E-cadherin/beta-catenin complex to efficient endoplasmic reticulum exit and basal-lateral membrane targeting of E-cadherin in polarized MDCK cells. J. Cell Biol. 144, 687–699 (1999).
Huber, A.H., Stewart, D.B., Laurents, D.V., Nelson, W.J. & Weis, W.I. The cadherin cytoplasmic domain is unstructured in the absence of beta-catenin. A possible mechanism for regulating cadherin turnover. J. Biol. Chem. 276, 12301–12309 (2001).
Drees, F., Pokutta, S., Yamada, S., Nelson, W.J. & Weis, W.I. Alpha-catenin is a molecular switch that binds E-cadherin-beta-catenin and regulates actin-filament assembly. Cell 123, 903–915 (2005).
Imamura, Y., Itoh, M., Maeno, Y., Tsukita, S. & Nagafuchi, A. Functional domains of alpha-catenin required for the strong state of cadherin-based cell adhesion. J. Cell Biol. 144, 1311–1322 (1999).
Weis, W.I. & Nelson, W.J. Re-solving the cadherin-catenin-actin conundrum. J. Biol. Chem. 281, 35593–35597 (2006).
Stappert, J. & Kemler, R. A short core region of E-cadherin is essential for catenin binding and is highly phosphorylated. Cell Adhes. Commun. 2, 319–327 (1994).
Daugherty, R.L. & Gottardi, C.J. Phospho-regulation of beta-catenin adhesion and signaling functions. Physiology (Bethesda) 22, 303–309 (2007).
Kuroda, S. et al. Role of IQGAP1, a target of the small GTPases Cdc42 and Rac1, in regulation of E-cadherin- mediated cell-cell adhesion. Science 281, 832–835 (1998).
Nieset, J.E. et al. Characterization of the interactions of alpha-catenin with alpha-actinin and beta-catenin/plakoglobin. J. Cell Sci. 110, 1013–1022 (1997).
Yap, A.S., Niessen, C.M. & Gumbiner, B.M. The juxtamembrane region of the cadherin cytoplasmic tail supports lateral clustering, adhesive strengthening, and interaction with p120ctn. J. Cell Biol. 141, 779–789 (1998).
Ishitani, T. et al. The TAK1-NLK-MAPK-related pathway antagonizes signalling between beta-catenin and transcription factor TCF. Nature 399, 798–802 (1999).
Meneghini, M.D. et al. MAP kinase and Wnt pathways converge to downregulate an HMG-domain repressor in Caenorhabditis elegans. Nature 399, 793–797 (1999).
Zeng, Y.A. & Verheyen, E.M. Nemo is an inducible antagonist of Wingless signaling during Drosophila wing development. Development 131, 2911–2920 (2004).
Verheyen, E.M. et al. The tissue polarity gene nemo carries out multiple roles in patterning during Drosophila development. Mech. Dev. 101, 119–132 (2001).
Zeng, Y.A., Rahnama, M., Wang, S., Sosu-Sedzorme, W. & Verheyen, E.M. Drosophila Nemo antagonizes BMP signaling by phosphorylation of Mad and inhibition of its nuclear accumulation. Development 134, 2061–2071 (2007).
Ishitani, T. et al. Nemo-like kinase suppresses Notch signalling by interfering with formation of the Notch active transcriptional complex. Nat. Cell Biol. 12, 278–285 (2010).
Weber, U., Pataki, C., Mihaly, J. & Mlodzik, M. Combinatorial signaling by the Frizzled/PCP and Egfr pathways during planar cell polarity establishment in the Drosophila eye. Dev. Biol. 316, 110–123 (2008).
Basler, K., Siegrist, P. & Hafen, E. The spatial and temporal expression pattern of sevenless is exclusively controlled by gene-internal elements. EMBO J. 8, 2381–2386 (1989).
Wolff, T. & Rubin, G.M. strabismus, a novel gene that regulates tissue polarity and cell fate decisions in Drosophila. Development 125, 1149–1159 (1998).
Strutt, D., Johnson, R., Cooper, K. & Bray, S. Asymmetric localization of frizzled and the determination of Notch-dependent cell fate in the Drosophila eye. Curr. Biol. 12, 813–824 (2002).
Nelson, W.J. & Nusse, R. Convergence of Wnt, beta-catenin, and cadherin pathways. Science 303, 1483–1487 (2004).
Pai, L.M., Orsulic, S., Bejsovec, A. & Peifer, M. Negative regulation of Armadillo, a Wingless effector in Drosophila. Development 124, 2255–2266 (1997).
Mirkovic, I., Charish, K., Gorski, S.M., McKnight, K. & Verheyen, E.M. Drosophila nemo is an essential gene involved in the regulation of programmed cell death. Mech. Dev. 119, 9–20 (2002).
Djiane, A., Yogev, S. & Mlodzik, M. The apical determinants aPKC and dPatj regulate Frizzled-dependent planar cell polarity in the Drosophila eye. Cell 121, 621–631 (2005).
Wu, J., Klein, T.J. & Mlodzik, M. Subcellular localization of frizzled receptors, mediated by their cytoplasmic tails, regulates signaling pathway specificity. PLoS Biol. 2, 1004–1014 (2004).
Courbard, J.R., Djiane, A., Wu, J. & Mlodzik, M. The apical/basal-polarity determinant Scribble cooperates with the PCP core factor Stbm/Vang and functions as one of its effectors. Dev. Biol. 333, 67–77 (2009).
Nagafuchi, A., Ishihara, S. & Tsukita, S. The roles of catenins in the cadherin-mediated cell adhesion: functional analysis of E-cadherin-alpha catenin fusion molecules. J. Cell Biol. 127, 235–245 (1994).
Dumstrei, K., Wang, F., Shy, D., Tepass, U. & Hartenstein, V. Interaction between EGFR signaling and DE-cadherin during nervous system morphogenesis. Development 129, 3983–3994 (2002).
Bertet, C., Sulak, L. & Lecuit, T. Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation. Nature 429, 667–671 (2004).
Fernandez-Gonzalez, R., Simoes Sde, M., Roper, J.C., Eaton, S. & Zallen, J.A. Myosin II dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009).
Simões, S.M. et al. Rho-kinase directs Bazooka/Par-3 planar polarity during Drosophila axis elongation. Dev. Cell 19, 377–388 (2010).
Shewan, A.M. et al. Myosin 2 is a key Rho kinase target necessary for the local concentration of E-cadherin at cell-cell contacts. Mol. Biol. Cell 16, 4531–4542 (2005).
Ulrich, F. et al. Slb/Wnt11 controls hypoblast cell migration and morphogenesis at the onset of zebrafish gastrulation. Development 130, 5375–5384 (2003).
Thorpe, C.J. & Moon, R.T. nemo-like kinase is an essential co-activator of Wnt signaling during early zebrafish development. Development 131, 2899–2909 (2004).
Ulrich, F. et al. Wnt11 functions in gastrulation by controlling cell cohesion through Rab5c and E-cadherin. Dev. Cell 9, 555–564 (2005).
Jenny, A., Reynolds-Kenneally, J., Das, G., Burnett, M. & Mlodzik, M. Diego and Prickle regulate Frizzled planar cell polarity signalling by competing for Dishevelled binding. Nat. Cell Biol. 7, 691–697 (2005).
Brand, A.H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Iwai, Y. et al. Axon patterning requires DN-cadherin, a novel neuronal adhesion receptor, in the Drosophila embryonic CNS. Neuron 19, 77–89 (1997).
Billuart, P., Winter, C.G., Maresh, A., Zhao, X. & Luo, L. Regulating axon branch stability: the role of p190 RhoGAP in repressing a retraction signaling pathway. Cell 107, 195–207 (2001).
Acknowledgements
We thank Z. Chen (Baylor College of Medicine), T. Clandinin (Stanford School of Medicine), H. Oda and S. Tsukita (Japan Science and Technology Corporation, Kyoto), P. Rørth (Institute of Molecular and Cell Biology, Singapore), K. Saigo (University of Tokyo), U. Tepass (University of Toronto), N. Tolwinski (Sloan-Kettering Institute), T. Uemura (Kyoto University), W. Weis (Stanford University), T. Wolff (Howard Hughes Medical Institute, Janelia Farm), B. Mollereau (Rockefeller University) and the Bloomington stock center for flies and reagents, S. Okello and Y.A. Zeng for technical assistance, N. Maj for help in generating the rose diagrams, J. Delaney and U. Weber for comments on the manuscript, and all members of the Mlodzik laboratory for helpful discussions. This work was supported by US National Institutes of Health, National Eye Institute grant RO1 EY14597 to M.M., by Natural Sciences and Engineering Research Council of Canada grant RGPIN/203545 and Canadian Institutes of Health Research grant MOP 62895 to E.V., and US National Institutes of Health grant GM076561 to C.J.G.
Author information
Authors and Affiliations
Contributions
I.M. and M.M. designed experiments; I.M., W.J.G., M.R., A.J. and K.G. conducted experiments and analyzed data; A.J., W.J.G. and C.J.G. edited the manuscript; D.B. generated the nmoDB allele; C.J.G. conducted biochemical experiments with β-cat and Nmo; I.M., E.M.V. and M.M. analyzed data and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1, Supplementary Figures 1–8 and Supplementary Methods (PDF 15557 kb)
Rights and permissions
About this article
Cite this article
Mirkovic, I., Gault, W., Rahnama, M. et al. Nemo kinase phosphorylates β-catenin to promote ommatidial rotation and connects core PCP factors to E-cadherin–β-catenin. Nat Struct Mol Biol 18, 665–672 (2011). https://doi.org/10.1038/nsmb.2049
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2049
This article is cited by
-
Tissue fluidity mediated by adherens junction dynamics promotes planar cell polarity-driven ommatidial rotation
Nature Communications (2021)
-
Notch signaling coordinates ommatidial rotation in the Drosophila eye via transcriptional regulation of the EGF-Receptor ligand Argos
Scientific Reports (2019)
-
Monomeric α-catenin links cadherin to the actin cytoskeleton
Nature Cell Biology (2013)
-
The many faces and functions of β-catenin
The EMBO Journal (2012)