Abstract
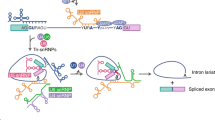
Most human genes produce multiple mRNA isoforms through alternative splicing. However, the biological relevance of most splice variants remains unclear. In this study, we evaluated the functional impact of alternative splicing in cancer cells. We modulated the splicing pattern of 41 cancer-associated splicing events and scored the effects on cell growth, viability and apoptosis, identifying three isoforms essential for cell survival. Specifically, changing the splicing pattern of the spleen tyrosine kinase gene (SYK) impaired cell-cycle progression and anchorage-independent growth. Notably, exposure of cancer cells to epithelial growth factor modulated the SYK splicing pattern to promote the pro-survival isoform that is associated with cancer tissues in vivo. The data suggest that splicing of selected genes is specifically modified during tumor development to allow the expression of isoforms that promote cancer cell survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pan, Q., Shai, O., Lee, L.J., Frey, B.J. & Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Wang, E.T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Matlin, A.J., Clark, F. & Smith, C.W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Castle, J.C. et al. Expression of 24,426 human alternative splicing events and predicted cis regulation in 48 tissues and cell lines. Nat. Genet. 40, 1416–1425 (2008).
Venables, J.P. et al. Cancer-associated regulation of alternative splicing. Nat. Struct. Mol. Biol. 16, 670–676 (2009).
Venables, J.P. Aberrant and alternative splicing in cancer. Cancer Res. 64, 7647–7654 (2004).
Cooper, D.L. & Dougherty, G.J. To metastasize or not? Selection of CD44 splice sites. Nat. Med. 1, 635–637 (1995).
Christofk, H.R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Levanon, E.Y. & Sorek, R. The importance of alternative splicing in the drug discovery process. Targets 2, 109–114 (2003).
Iorns, E., Lord, C.J., Turner, N. & Ashworth, A. Utilizing RNA interference to enhance cancer drug discovery. Nat. Rev. Drug Discov. 6, 556–568 (2007).
Klinck, R. et al. Multiple alternative splicing markers for ovarian cancer. Cancer Res. 68, 657–663 (2008).
Venables, J.P. et al. Identification of alternative splicing markers for breast cancer. Cancer Res. 68, 9525–9531 (2008).
Celotto, A.M. & Graveley, B.R. Exon-specific RNAi: a tool for dissecting the functional relevance of alternative splicing. RNA 8, 718–724 (2002).
Villemaire, J., Dion, I., Elela, S.A. & Chabot, B. Reprogramming alternative pre-messenger RNA splicing through the use of protein-binding antisense oligonucleotides. J. Biol. Chem. 278, 50031–50039 (2003).
Baudot, A.D. et al. The tyrosine kinase Syk regulates the survival of chronic lymphocytic leukemia B cells through PKCdelta and proteasome-dependent regulation of Mcl-1 expression. Oncogene 28, 3261–3273 (2009).
Gobessi, S. et al. Inhibition of constitutive and BCR-induced Syk activation downregulates Mcl-1 and induces apoptosis in chronic lymphocytic leukemia B cells. Leukemia 23, 686–697 (2009).
Adams, K.W. & Cooper, G.M. Rapid turnover of mcl-1 couples translation to cell survival and apoptosis. J. Biol. Chem. 282, 6192–6200 (2007).
Bingle, C.D. et al. Exon skipping in Mcl-1 results in a bcl-2 homology domain 3 only gene product that promotes cell death. J. Biol. Chem. 275, 22136–22146 (2000).
Wang, L. et al. Alternative splicing disrupts a nuclear localization signal in spleen tyrosine kinase that is required for invasion suppression in breast cancer. Cancer Res. 63, 4724–4730 (2003).
Zhou, F., Hu, J., Ma, H., Harrison, M.L. & Geahlen, R.L. Nucleocytoplasmic trafficking of the Syk protein tyrosine kinase. Mol. Cell. Biol. 26, 3478–3491 (2006).
Shaulian, E. AP-1—the Jun proteins: oncogenes or tumor suppressors in disguise? Cell. Signal. 22, 894–899 (2010).
Wisdom, R., Johnson, R.S. & Moore, C. c-Jun regulates cell cycle progression and apoptosis by distinct mechanisms. EMBO J. 18, 188–197 (1999).
Bennett, B.L. et al. SP600125, an anthrapyrazolone inhibitor of Jun N-terminal kinase. Proc. Natl. Acad. Sci. USA 98, 13681–13686 (2001).
Akgul, C. Mcl-1 is a potential therapeutic target in multiple types of cancer. Cell. Mol. Life Sci. 66, 1326–1336 (2009).
Michels, J., Johnson, P.W. & Packham, G. Mcl-1. Int. J. Biochem. Cell Biol. 37, 267–271 (2005).
Bae, J., Leo, C.P., Hsu, S.Y. & Hsueh, A.J. MCL-1S, a splicing variant of the antiapoptotic BCL-2 family member MCL-1, encodes a proapoptotic protein possessing only the BH3 domain. J. Biol. Chem. 275, 25255–25261 (2000).
Shieh, J.J., Liu, K.T., Huang, S.W., Chen, Y.J. & Hsieh, T.Y. Modification of alternative splicing of Mcl-1 pre-mRNA using antisense morpholino oligonucleotides induces apoptosis in basal cell carcinoma cells. J. Invest. Dermatol. 129, 2497–2506 (2009).
Mócsai, A., Ruland, J. & Tybulewicz, V.L. The SYK tyrosine kinase: a crucial player in diverse biological functions. Nat. Rev. Immunol. 10, 387–402 (2010).
Coopman, P.J. et al. The Syk tyrosine kinase suppresses malignant growth of human breast cancer cells. Nature 406, 742–747 (2000).
Moroni, M. et al. Progressive loss of Syk and abnormal proliferation in breast cancer cells. Cancer Res. 64, 7346–7354 (2004).
Zyss, D. et al. The Syk tyrosine kinase localizes to the centrosomes and negatively affects mitotic progression. Cancer Res. 65, 10872–10880 (2005).
Faruki, S., Geahlen, R.L. & Asai, D.J. Syk-dependent phosphorylation of microtubules in activated B-lymphocytes. J. Cell Sci. 113, 2557–2565 (2000).
Peters, J.D., Furlong, M.T., Asai, D.J., Harrison, M.L. & Geahlen, R.L. Syk, activated by cross-linking the B-cell antigen receptor, localizes to the cytosol where it interacts with and phosphorylates alpha-tubulin on tyrosine. J. Biol. Chem. 271, 4755–4762 (1996).
Feldman, A.L. et al. Overexpression of Syk tyrosine kinase in peripheral T-cell lymphomas. Leukemia 22, 1139–1143 (2008).
Buchner, M. et al. Spleen tyrosine kinase is overexpressed and represents a potential therapeutic target in chronic lymphocytic leukemia. Cancer Res. 69, 5424–5432 (2009).
Nakashima, H. et al. Clinical significance of nuclear expression of spleen tyrosine kinase (Syk) in gastric cancer. Cancer Lett. 236, 89–94 (2006).
Luangdilok, S. et al. Syk tyrosine kinase is linked to cell motility and progression in squamous cell carcinomas of the head and neck. Cancer Res. 67, 7907–7916 (2007).
Goodman, P.A., Wood, C.M., Vassilev, A., Mao, C. & Uckun, F.M. Spleen tyrosine kinase (Syk) deficiency in childhood pro-B cell acute lymphoblastic leukemia. Oncogene 20, 3969–3978 (2001).
Blaustein, M. et al. Concerted regulation of nuclear and cytoplasmic activities of SR proteins by AKT. Nat. Struct. Mol. Biol. 12, 1037–1044 (2005).
Shin, C. & Manley, J.L. Cell signalling and the control of pre-mRNA splicing. Nat. Rev. Mol. Cell Biol. 5, 727–738 (2004).
Mandal, M. et al. Growth factors regulate heterogeneous nuclear ribonucleoprotein K expression and function. J. Biol. Chem. 276, 9699–9704 (2001).
Ashton-Beaucage, D. et al. The exon junction complex controls the splicing of MAPK and other long intron-containing transcripts in Drosophila. Cell 143, 251–262 (2010).
Roignant, J.Y. & Treisman, J.E. Exon junction complex subunits are required to splice Drosophila MAP kinase, a large heterochromatic gene. Cell 143, 238–250 (2010).
Nicke, B. et al. Involvement of MINK, a Ste20 family kinase, in Ras oncogene-induced growth arrest in human ovarian surface epithelial cells. Mol. Cell 20, 673–685 (2005).
Hendzel, M.J. et al. Mitosis-specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma 106, 348–360 (1997).
Brosseau, J.P. et al. High-throughput quantification of splicing isoforms. RNA 16, 442–449 (2010).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 8, R19 (2007).
Venables, J.P. et al. Multiple and specific mRNA processing targets for the major human hnRNP proteins. Mol. Cell. Biol. 28, 6033–6043 (2008).
Zhang, J.H., Chung, T.D. & Oldenburg, K.R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Birmingham, A. et al. Statistical methods for analysis of high-throughput RNA interference screens. Nat. Methods 6, 569–575 (2009).
Chung, N. et al. Median absolute deviation to improve hit selection for genome-scale RNAi screens. J. Biomol. Screen. 13, 149–158 (2008).
Acknowledgements
We are indebted to C. Rancourt (Département de Microbiologie, Université de Sherbrooke) for providing cell lines and help in the initial setup phase of the project and to B. Lamontagne and L. Bergeron Jr. for assay setup and help with troubleshooting. We thank the Réseau de Recherche sur le Cancer (Fonds de la Recherche en Santé du Québec; FRSQ) tissue bank for ovarian tumor tissues and L. Volkov for help with flow cytometry. This work was funded by Genome Canada/Génome Québec and National Cancer Institute of Canada grant no. 700529. B.C. is the Canada Research Chair in Functional Genomics. J.-P.P. is the Canada Research Chair on Genomics and Catalytic RNA. S.A.E. is a Chercheur National of the FRSQ.
Author information
Authors and Affiliations
Contributions
D. Garneau, M.B., D. Gendron., S.C., J.-P.B., E.L. and M.D. carried out experiments and analyzed data and J.-F.L., P.T. and J.G.-B. analyzed data and prepared the figures. J.-F.L. developed the ISI design program and the FASE statistics and bioinformatics analysis tools. J.-P.B., K.T., E.L. and P.T. developed the qPCR procedures used to evaluate ISI impact on splicing. H.N. did the histopathological review of tissue specimens. P.P., K.T., R.K., J.-P.P., B.C., R.J.W. and S.A.E. designed experiments, discussed data and participated in the writing of the paper. P.P. and K.T. supervised experiments and analyzed data. P.P. and S.A.E. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1 and 3, and Supplementary Methods (PDF 3282 kb)
Supplementary Table 2
Primary FASE screen results. The results of the primary screen are shown, and the numbers of ISIs that are above the Z-score cut-off (>3) are indicated for each gene and each assay separately. Only ASEs with more than two positive ISIs per assay were retained for the validation phase (phase 2). (XLS 41 kb)
Rights and permissions
About this article
Cite this article
Prinos, P., Garneau, D., Lucier, JF. et al. Alternative splicing of SYK regulates mitosis and cell survival. Nat Struct Mol Biol 18, 673–679 (2011). https://doi.org/10.1038/nsmb.2040
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2040
This article is cited by
-
Pan-cancer analysis of clinical relevance of alternative splicing events in 31 human cancers
Oncogene (2019)
-
MMSET I acts as an oncoprotein and regulates GLO1 expression in t(4;14) multiple myeloma cells
Leukemia (2019)
-
Expression of Syk and MAP4 proteins in ovarian cancer
Journal of Cancer Research and Clinical Oncology (2019)
-
Splicing isoform-specific functional genomic in cancer cells
Applied Cancer Research (2018)
-
Inhibition of ovarian tumor cell invasiveness by targeting SYK in the tyrosine kinase signaling pathway
Oncogene (2018)