Abstract
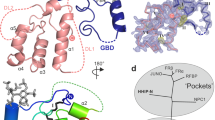
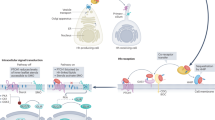
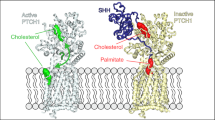
Hedgehog (Hh) morphogens have fundamental roles in development, whereas dysregulation of Hh signaling leads to disease. Multiple cell-surface receptors are responsible for transducing and/or regulating Hh signals. Among these, the Hedgehog-interacting protein (Hhip) is a highly conserved, vertebrate-specific inhibitor of Hh signaling. We have solved a series of crystal structures for the human HHIP ectodomain and Desert hedgehog (DHH) in isolation, as well as HHIP in complex with DHH (HHIP–DHH) and Sonic hedgehog (Shh) (HHIP–Shh), with and without Ca2+. The interaction determinants, confirmed by biophysical studies and mutagenesis, reveal previously uncharacterized and distinct functions for the Hh Zn2+ and Ca2+ binding sites—functions that may be common to all vertebrate Hh proteins. Zn2+ makes a key contribution to the Hh–HHIP interface, whereas Ca2+ is likely to prevent electrostatic repulsion between the two proteins, suggesting an important modulatory role. This interplay of several metal binding sites suggests a tuneable mechanism for regulation of Hh signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ingham, P.W. & McMahon, A.P. Hedgehog signaling in animal development: paradigms and principles. Genes Dev. 15, 3059–3087 (2001).
Varjosalo, M. & Taipale, J. Hedgehog: functions and mechanisms. Genes Dev. 22, 2454–2472 (2008).
Roessler, E. et al. Mutations in the human Sonic hedgehog gene cause holoprosencephaly. Nat. Genet. 14, 357–360 (1996).
Dellovade, T., Romer, J.T., Curran, T. & Rubin, L.L. The hedgehog pathway and neurological disorders. Annu. Rev. Neurosci. 29, 539–563 (2006).
Yauch, R.L. et al. A paracrine requirement for hedgehog signalling in cancer. Nature 455, 406–410 (2008).
Mann, R.K. & Beachy, P.A. Novel lipid modifications of secreted protein signals. Annu. Rev. Biochem. 73, 891–923 (2004).
Zeng, X. et al. A freely diffusible form of Sonic hedgehog mediates long-range signalling. Nature 411, 716–720 (2001).
Panáková, D., Sprong, H., Marois, E., Thiele, C. & Eaton, S. Lipoprotein particles are required for Hedgehog and Wingless signalling. Nature 435, 58–65 (2005).
Pepinsky, R.B. et al. Identification of a palmitic acid-modified form of human Sonic hedgehog. J. Biol. Chem. 273, 14037–14045 (1998).
Lum, L. & Beachy, P.A. The Hedgehog response network: sensors, switches, and routers. Science 304, 1755–1759 (2004).
Wilson, C.W. & Chuang, P.T. New “hogs” in Hedgehog transport and signal reception. Cell 125, 435–438 (2006).
Beckett, K., Franch-Marro, X. & Vincent, J.P. Glypican-mediated endocytosis of Hedgehog has opposite effects in flies and mice. Trends Cell Biol. 18, 360–363 (2008).
Chuang, P.T. & McMahon, A.P. Vertebrate Hedgehog signalling modulated by induction of a Hedgehog-binding protein. Nature 397, 617–621 (1999).
Kang, J.S., Zhang, W. & Krauss, R.S. Hedgehog signaling: cooking with Gas1. Sci. STKE 2007, pe50 (2007).
Coulombe, J., Traiffort, E., Loulier, K., Faure, H. & Ruat, M. Hedgehog interacting protein in the mature brain: membrane-associated and soluble forms. Mol. Cell. Neurosci. 25, 323–333 (2004).
Parmantier, E. et al. Schwann cell-derived Desert hedgehog controls the development of peripheral nerve sheaths. Neuron 23, 713–724 (1999).
Bourikas, D. et al. Sonic hedgehog guides commissural axons along the longitudinal axis of the spinal cord. Nat. Neurosci. 8, 297–304 (2005).
Olsen, C.L., Hsu, P.P., Glienke, J., Rubanyi, G.M. & Brooks, A.R. Hedgehog-interacting protein is highly expressed in endothelial cells but down-regulated during angiogenesis and in several human tumors. BMC Cancer 4, 43 (2004).
Tada, M. et al. Down-regulation of hedgehog-interacting protein through genetic and epigenetic alterations in human hepatocellular carcinoma. Clin. Cancer Res. 14, 3768–3776 (2008).
Taniguchi, H. et al. Intrahepatic mRNA levels of type I interferon receptor and interferon-stimulated genes in genotype 1b chronic hepatitis C. Association between IFNAR1 mRNA level and sustained response to interferon therapy. Intervirology 50, 32–39 (2007).
Tojo, M., Kiyosawa, H., Iwatsuki, K. & Kaneko, F. Expression of a sonic hedgehog signal transducer, hedgehog-interacting protein, by human basal cell carcinoma. Br. J. Dermatol. 146, 69–73 (2002).
Aricescu, A.R., Lu, W. & Jones, E.Y. A time- and cost-efficient system for high-level protein production in mammalian cells. Acta Crystallogr. D Biol. Crystallogr. 62, 1243–1250 (2006).
Harding, M.M. Geometry of metal-ligand interactions in proteins. Acta Crystallogr. D Biol. Crystallogr. 57, 401–411 (2001).
Hall, T.M., Porter, J.A., Beachy, P.A. & Leahy, D.J. A potential catalytic site revealed by the 1.7-crystal structure of the amino-terminal signalling domain of Sonic hedgehog. Nature 378, 212–216 (1995).
Day, E.S. et al. Zinc-dependent structural stability of human Sonic hedgehog. Biochemistry 38, 14868–14880 (1999).
McLellan, J.S. et al. The mode of Hedgehog binding to Ihog homologues is not conserved across different phyla. Nature 455, 979–983 (2008).
Fuse, N. et al. Sonic hedgehog protein signals not as a hydrolytic enzyme but as an apparent ligand for patched. Proc. Natl. Acad. Sci. USA 96, 10992–10999 (1999).
Orioli, I.M. et al. Identification of novel mutations in SHH and ZIC2 in a South American (ECLAMC) population with holoprosencephaly. Hum. Genet. 109, 1–6 (2001).
Ribeiro, L.A. & Richieri-Costa, A. Single median maxillary central incisor, hypophyseal tumor, and SHH mutation. Am. J. Med. Genet. A. 136A, 346–347 (2005).
Hofer, A.M. & Brown, E.M. Extracellular calcium sensing and signalling. Nat. Rev. Mol. Cell Biol. 4, 530–538 (2003).
Springer, T.A., Zhu, J. & Xiao, T. Structural basis for distinctive recognition of fibrinogen γC peptide by the platelet integrin αIIbβ3. J. Cell Biol. 182, 791–800 (2008).
Chang, V.T. et al. Glycoprotein structural genomics: solving the glycosylation problem. Structure 15, 267–273 (2007).
Walter, T.S. et al. A procedure for setting up high-throughput nanolitre crystallization experiments. Crystallization workflow for initial screening, automated storage, imaging and optimization. Acta Crystallogr. D Biol. Crystallogr. 61, 651–657 (2005).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Terwilliger, T.C. Automated side-chain model building and sequence assignment by template matching. Acta Crystallogr. D Biol. Crystallogr. 59, 45–49 (2003).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D Biol. Crystallogr. 60, 2210–2221 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Hooft, R.W., Vriend, G., Sander, C. & Abola, E.E. Errors in protein structures. Nature 381, 272 (1996).
Stuart, D.I., Levine, M., Muirhead, H. & Stammers, D.K. Crystal structure of cat muscle pyruvate kinase at a resolution of 2.6. J. Mol. Biol. 134, 109–142 (1979).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Lawrence, M.C. & Colman, P.M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Trombetta, E.S. & Parodi, A.J. Quality control and protein folding in the secretory pathway. Annu. Rev. Cell Dev. Biol. 19, 649–676 (2003).
O'Callaghan, C.A. et al. BirA enzyme: production and application in the study of membrane receptor-ligand interactions by site-specific biotinylation. Anal. Biochem. 266, 9–15 (1999).
Acknowledgements
We thank the staff of European Synchrotron Radiation Facility beamline ID 29 and Diamond beamlines I02 and I03 for assistance with data collection, T. Walter for help with crystallization, G. Sutton for help with MALS experiments, and S. Graham and D. Stuart for discussions. We acknowledge the use of the crystallization facilities provided by the Medical Research Council (MRC)–funded Oxford Protein Production Facility (OPPF). The work was funded by the Wellcome Trust. A.R.A. and C.A.O'C. are funded by the MRC. E.Y.J. is funded by Cancer Research UK. C.S. is funded by the Wellcome Trust.
Author information
Authors and Affiliations
Contributions
C.S. designed the project; B.B. and C.S. produced the constructs and crystallized the proteins; B.B. and C.A.O'C. performed the SPR experiments; K.H. and C.S. collected X-ray data; C.S. solved the structures; B.B, A.R.A., C.A.O'C., E.Y.J. and C.S. analyzed the data and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 1534 kb)
Rights and permissions
About this article
Cite this article
Bishop, B., Aricescu, A., Harlos, K. et al. Structural insights into hedgehog ligand sequestration by the human hedgehog-interacting protein HHIP. Nat Struct Mol Biol 16, 698–703 (2009). https://doi.org/10.1038/nsmb.1607
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1607
This article is cited by
-
Hedgehog-Interacting Protein is a multimodal antagonist of Hedgehog signalling
Nature Communications (2021)
-
SLITRK5 is a negative regulator of hedgehog signaling in osteoblasts
Nature Communications (2021)
-
GLI3: a mediator of genetic diseases, development and cancer
Cell Communication and Signaling (2020)
-
DHH pathogenic variants involved in 46,XY disorders of sex development differentially impact protein self-cleavage and structural conformation
Human Genetics (2020)
-
Hedgehog Interacting Protein (Hhip) Regulates Insulin Secretion in Mice Fed High Fat Diets
Scientific Reports (2019)