Abstract
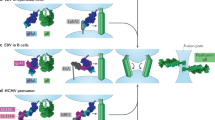
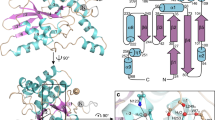
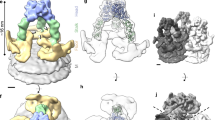
Viral fusion proteins mediate the merger of host and viral membranes during cell entry for all enveloped viruses. Baculovirus glycoprotein gp64 (gp64) is unusual in promoting entry into both insect and mammalian cells and is distinct from established class I and class II fusion proteins. We report the crystal structure of its postfusion form, which explains a number of gp64′s biological properties including its cellular promiscuity, identifies the fusion peptides and shows it to be the third representative of a new class (III) of fusion proteins with unexpected structural homology with vesicular stomatitis virus G and herpes simplex virus type 1 gB proteins. We show that domains of class III proteins have counterparts in both class I and II proteins, suggesting that all these viral fusion machines are structurally more related than previously thought.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Kielian, M. & Rey, F.A. Virus membrane-fusion proteins: more than one way to make a hairpin. Nat. Rev. Microbiol. 4, 67–76 (2006).
Bullough, P.A., Hughson, F.M., Skehel, J.J. & Wiley, D.C. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371, 37–43 (1994).
Yin, H.S., Wen, X., Paterson, R.G., Lamb, R.A. & Jardetzky, T.S. Structure of the parainfluenza virus 5 F protein in its metastable, prefusion conformation. Nature 439, 38–44 (2006).
Gibbons, D.L. et al. Conformational change and protein-protein interactions of the fusion protein of Semliki Forest virus. Nature 427, 320–325 (2004).
Modis, Y., Ogata, S., Clements, D. & Harrison, S.C. Structure of the dengue virus envelope protein after membrane fusion. Nature 427, 313–319 (2004).
Heldwein, E.E. et al. Crystal structure of glycoprotein B from herpes simplex virus 1. Science 313, 217–220 (2006).
Roche, S., Bressanelli, S., Rey, F.A. & Gaudin, Y. Crystal structure of the low-pH form of the vesicular stomatitis virus glycoprotein G. Science 313, 187–191 (2006).
Roche, S., Rey, F.A., Gaudin, Y. & Bressanelli, S. Structure of the prefusion form of the vesicular stomatitis virus glycoprotein G. Science 315, 843–848 (2007).
Blissard, G.W. & Rohrmann, G.F. Baculovirus diversity and molecular biology. Annu. Rev. Entomol. 35, 127–155 (1990).
Blissard, G.W. & Wenz, J.R. Baculovirus gp64 envelope glycoprotein is sufficient to mediate pH-dependent membrane fusion. J. Virol. 66, 6829–6835 (1992).
Hefferon, K.L., Oomens, A.G., Monsma, S.A., Finnerty, C.M. & Blissard, G.W. Host cell receptor binding by baculovirus gp64 and kinetics of virion entry. Virology 258, 455–468 (1999).
Ho, Y.C. et al. Baculovirus transduction of human mesenchymal stem cell-derived progenitor cells: variation of transgene expression with cellular differentiation states. Gene Ther. 13, 1471–1479 (2006).
Zeng, J., Du, J., Zhao, Y., Palanisamy, N. & Wang, S. Baculoviral vector-mediated transient and stable transgene expression in human embryonic stem cells. Stem Cells 25, 1055–1061 (2007).
Kost, T.A., Condreay, J.P. & Jarvis, D.L. Baculovirus as versatile vectors for protein expression in insect and mammalian cells. Nat. Biotechnol. 23, 567–575 (2005).
Boublik, Y., Di Bonito, P. & Jones, I.M. Eukaryotic virus display: engineering the major surface glycoprotein of the Autographa californica nuclear polyhedrosis virus (AcNPV) for the presentation of foreign proteins on the virus surface. Bio/Technology 13, 1079–1084 (1995).
Zhou, J. & Blissard, G.W. Mapping the conformational epitope of a neutralizing antibody (AcV1) directed against the AcMNPV gp64 protein. Virology 352, 427–437 (2006).
Summers, M.D. & Volkman, L.E. Comparison of biophysical and morphological properties of occluded and extracellular nonoccluded baculovirus from in vivo and in vitro host systems. J. Virol. 17, 962–972 (1976).
Zhang, Y., Kaufmann, B., Chipman, P.R., Kuhn, R.J. & Rossmann, M.G. Structure of immature West Nile virus. J. Virol. 81, 6141–6145 (2007).
Plonsky, I. & Zimmerberg, J. The initial fusion pore induced by baculovirus gp64 is large and forms quickly. J. Cell Biol. 135, 1831–1839 (1996).
Brown, J.H., Cohen, C. & Parry, D.A. Heptad breaks in α-helical coiled coils: stutters and stammers. Proteins 26, 134–145 (1996).
Oomens, A.G., Monsma, S.A. & Blissard, G.W. The baculovirus gp64 envelope fusion protein: synthesis, oligomerization, and processing. Virology 209, 592–603 (1995).
Kampmann, T., Mueller, D.S., Mark, A.E., Young, P.R. & Kobe, B. The role of histidine residues in low-pH-mediated viral membrane fusion. Structure 14, 1481–1487 (2006).
Fredericksen, B.L. & Whitt, M.A. Vesicular stomatitis virus glycoprotein mutations that affect membrane fusion activity and abolish virus infectivity. J. Virol. 69, 1435–1443 (1995).
Hannah, B.P., Heldwein, E.E., Bender, F.C., Cohen, G.H. & Eisenberg, R.J. Mutational evidence of internal fusion loops in herpes simplex virus glycoprotein B. J. Virol. 81, 4858–4865 (2007).
Backovic, M., Jardetzky, T.S. & Longnecker, R. Hydrophobic residues that form putative fusion loops of Epstein-Barr virus glycoprotein B are critical for fusion activity. J. Virol. 81, 9596–9600 (2007).
Monsma, S.A. & Blissard, G.W. Identification of a membrane fusion domain and an oligomerization domain in the baculovirus gp64 envelope fusion protein. J. Virol. 69, 2583–2595 (1995).
Zhou, J. & Blissard, G.W. Identification of a gp64 subdomain involved in receptor binding by budded virions of the baculovirus AcMNPV. J. Virol. (2008).
Tani, H., Nishijima, M., Ushijima, H., Miyamura, T. & Matsuura, Y. Characterization of cell-surface determinants important for baculovirus infection. Virology 279, 343–353 (2001).
Yin, H.S., Paterson, R.G., Wen, X., Lamb, R.A. & Jardetzky, T.S. Structure of the uncleaved ectodomain of the paramyxovirus (hPIV3) fusion protein. Proc. Natl. Acad. Sci. USA 102, 9288–9293 (2005).
Rossmann, M.G. & Argos, P. Exploring structural homology of proteins. J. Mol. Biol. 105, 75–95 (1976).
Stuart, D.I., Levine, M., Muirhead, H. & Stammers, D.K. Crystal structure of cat muscle pyruvate kinase at a resolution of 2.6. J. Mol. Biol. 134, 109–142 (1979).
Weissenhorn, W., Hinz, A. & Gaudin, Y. Virus membrane fusion. FEBS Lett. 581, 2150–2155 (2007).
Lu, W., Chapple, S.D., Lissini, O. & Jones, I.M. Characterization of a truncated soluble form of the baculovirus (AcMNPV) major envelope protein gp64. Protein Expr. Purif. 24, 196–201 (2002).
Walter, T.S. et al. Semi-automated microseeding of nanolitre crystallization experiments. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 64, 14–18 (2008).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
Abrahams, J.P. & Leslie, A.G.W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Cowtan, K.D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D Biol. Crystallogr. 52, 43–48 (1996).
Roversi, P., Blanc, E., Vonrhein, C., Evans, G. & Bricogne, G. Modelling prior distributions of atoms for macromolecular refinement and completion. Acta Crystallogr. D Biol. Crystallogr. 56, 1316–1323 (2000).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Li, H., Robertson, A.D. & Jensen, J.H. Very fast empirical prediction and rationalization of protein pKa values. Proteins 61, 704–721 (2005).
Bond, C.S. TopDraw: a sketchpad for protein structure topology cartoons. Bioinformatics 19, 311–312 (2003).
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F. & Higgins, D.G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Gouet, P., Courcelle, E., Stuart, D.I. & Metoz, F. ESPript: multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Strelkov, S.V. & Burkhard, P. Analysis of α-helical coiled coils with the program TWISTER reveals a structural mechanism for stutter compensation. J. Struct. Biol. 137, 54–64 (2002).
Acknowledgements
The work was supported by grants from the UK Medical Research Council to D.I.S. and I.M.J. J.K. was supported by a long term fellowship from the European Molecular Biology Organisation (EMBO). We thank M. Walsh for assistance during the data collection at the UK MAD beamline BM14 at the European Synchrotron Radiation Facility and R. Aricescu for help with syncytium-formation assays. We also thank L. Volkman (University of California) for the gift of the B12D5 monoclonal antibody and K. Harlos for the help with heavy-atom derivatives.
Author information
Authors and Affiliations
Contributions
S.L. and I.M.J. expressed and purified the protein; J.K. and N.G.A.A. obtained crystals, and solved and refined the structure; J.K. and S.L. performed syncytium-formation assays; J.K., D.I.S. and I.M.J. analyzed the structure and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 4395 kb)
Supplementary Video 1
Transition of dengue virus E domain III to HSV-1 gB domain IV, first superposed with program SHP. Structurally equivalent residues are shown in red, while areas where major changes occur are in green. Domain boundaries and orientation correspond to the ones in Figure 4. (MOV 1461 kb)
Supplementary Video 2
Transition of paramyxovirus F domain I to gp64 domain Ib, first superposed with program SHP. Structurally equivalent residues are shown in red, while areas where major changes occur are in green. Domain boundaries and orientation correspond to the ones in Figure 4. (MOV 807 kb)
Supplementary Video 3
Transition of dengue virus E domain IIa to VSV G domain Ib, first superposed with program SHP. Structurally equivalent residues are shown in red, while areas where major changes occur are in green. Domain boundaries and orientation correspond to the ones in Figure 4. (MOV 940 kb)
Supplementary Video 4
Transition of paramyxovirus F domain III to dengue virus E domain IIa, first superposed with program SHP. For clarity, the helical insertion of paramyxovirus F protein (56-239) is not shown, and residues 55 and 240 are connected. Structurally equivalent residues are shown in red, while areas where major changes occur are in green. Domain boundaries and orientation correspond to the ones in Figure 4. (MOV 1011 kb)
Rights and permissions
About this article
Cite this article
Kadlec, J., Loureiro, S., Abrescia, N. et al. The postfusion structure of baculovirus gp64 supports a unified view of viral fusion machines. Nat Struct Mol Biol 15, 1024–1030 (2008). https://doi.org/10.1038/nsmb.1484
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1484
This article is cited by
-
Exploring viral infections in honey bee colonies: insights from a metagenomic study in southern Brazil
Brazilian Journal of Microbiology (2023)
-
Recombinant AcMNPV-gp64-EGFP and synergist triphenyl phosphate, an effective combination against Spodoptera frugiperda
Biotechnology Letters (2022)
-
The structural basis of herpesvirus entry
Nature Reviews Microbiology (2021)
-
An efficient method for multigene co-interference by recombinant Bombyx mori nucleopolyhedrovirus
Molecular Genetics and Genomics (2019)
-
Novel GP64 envelope variants for improved delivery to human airway epithelial cells
Gene Therapy (2017)