Abstract
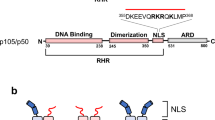
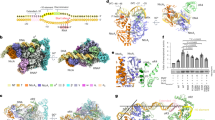
The crystal structure of the NFAT1 Rel homology region (RHR) bound to a pseudo-palindromic DNA site reveals an asymmetric dimer interaction between the RHR-C domains, unrelated to the contact seen in Rel dimers such as NFκB. Binding studies with a form of the NFAT1 RHR defective in the dimer contact show loss of cooperativity and demonstrate that the same interaction is present in solution. The structure we have determined may correspond to a functional NFAT binding mode at palindromic sites of genes induced during the anergic response to weak TCR signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Crabtree, G.R. & Olson, E.N. NFAT signaling: choreographing the social lives of cells. Cell 109 (suppl.), S67–S79 (2002).
Rao, A., Luo, C. & Hogan, P.G. Transcription factors of the NFAT family: regulation and function. Annu. Rev. Immunol. 15, 707–747 (1997).
Chytil, M. & Verdine, G.L. The Rel family of eukaryotic transcription factors. Curr. Opin. Struct. Biol. 6, 91–100 (1996).
Chen, L., Glover, J.N., Hogan, P.G., Rao, A. & Harrison, S.C. Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 392, 42–48 (1998).
Chen, L. Combinatorial gene regulation by eukaryotic transcription factors. Curr. Opin. Struct. Biol. 9, 48–55 (1999).
Giffin, M.J. et al. Structure of NFAT1 bound as a dimer to the HIV-1 LTR κB element. Nat. Struct. Biol. advance online publication, 31 August 2003 (doi:10.1038/nsb981).
Ghosh, G., van Duyne, G., Ghosh, S. & Sigler, P.B. Structure of NF-κB p50 homodimer bound to a κB site. Nature 373, 303–310 (1995).
Baltimore, D. & Beg, A.A. DNA-binding proteins. A butterfly flutters by. Nature 373, 287–288 (1995).
Muller, C.W., Rey, F.A., Sodeoka, M., Verdine, G.L. & Harrison, S.C. Structure of the NF-κB p50 homodimer bound to DNA. Nature 373, 311–317 (1995).
Seeman, N.C., Rosenberg, J.M. & Rich, A. Sequence-specific recognition of double helical nucleic acids by proteins. Proc. Natl. Acad. Sci. USA 73, 804–808 (1976).
Zhou, P., Sun, L.J., Dotsch, V., Wagner, G. & Verdine, G.L. Solution structure of the core NFATC1/DNA complex. Cell 92, 687–696 (1998).
Stroud, J.C., Lopez-Rodriguez, C., Rao, A. & Chen, L. Structure of a TonEBP-DNA complex reveals DNA encircled by a transcription factor. Nat. Struct. Biol. 9, 90–94 (2002).
Macian, F. et al. Transcriptional mechanisms underlying lymphocyte tolerance. Cell 109, 719–731 (2002).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Vagin, A. & Teplyakov, A. An approach to multi-copy search in molecular replacement. Acta Crystallogr. D 56, 1622–1624 (2000).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for the building of protein models in electron-density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Jain, J., Burgeon, E., Badalian, T.M., Hogan, P.G. & Rao, A. A similar DNA-binding motif in NFAT family proteins and the Rel homology region. J. Biol. Chem. 270, 4138–4145 (1995).
Acknowledgements
We acknowledge support from US National Institutes of Health grants to S.C.H. and to A.R. CHESS is supported by the US National Science Foundation. S.C.H is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Jin, L., Sliz, P., Chen, L. et al. An asymmetric NFAT1 dimer on a pseudo-palindromic κB-like DNA site. Nat Struct Mol Biol 10, 807–811 (2003). https://doi.org/10.1038/nsb975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb975
This article is cited by
-
The calcineurin/NFAT pathway is activated in diagnostic breast cancer cases and is essential to survival and metastasis of mammary cancer cells
Cell Death & Disease (2015)
-
NFATc1/αA: The other Face of NFAT Factors in Lymphocytes
Cell Communication and Signaling (2012)
-
NFAT proteins: key regulators of T-cell development and function
Nature Reviews Immunology (2005)
-
Structure of NFAT1 bound as a dimer to the HIV-1 LTR κB element
Nature Structural & Molecular Biology (2003)