Abstract
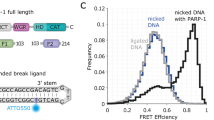
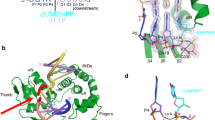
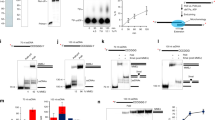
XRCC1 functions in the repair of single-strand DNA breaks in mammalian cells and forms a repair complex with β-Pol, ligase III and PARP. Here we describe the NMR solution structure of the XRCC1 N-terminal domain (XRCC1 NTD). The structural core is a β-sandwich with β-strands connected by loops, three helices and two short two-stranded β-sheets at each connection side. We show, for the first time, that the XRCC1 NTD specifically binds single-strand break DNA (gapped and nicked). We also show that the XRCC1 NTD binds a gapped DNA–β-Pol complex. The DNA binding and β-Pol binding surfaces were mapped by NMR and found to be well suited for interaction with single-strand gap DNA containing a 90° bend, and for simultaneously making contacts with the palm-thumb of β-Pol in a ternary complex. The findings suggest a mechanism for preferential binding of the XRCC1 NTD to flexible single-strand break DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Caldecott, K.W., Tucker, J.D., Stanker, L.H. & Thompson, L.H. Characterization of the XRCC1–DNA ligase III complex in vitro and its absence from mutant hamster cells. Nucleic Acids Res. 23, 4836–4843 (1995).
Caldecott, K.W., Aoufouchi, S., Johnson, P. & Shall, S. XRCC1 polypeptide interacts with DNA polymerase β and possibly poly (ADP-ribose) polymerase, and DNA ligase III is a novel molecular 'nick-sensor' in vitro . Nucleic Acids Res. 24, 4387– 4394 (1996).
Masson, M. et al. XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol. Cell. Biol. 18, 3563–3571 (1998).
Kubota, Y. et al. Reconstitution of DNA base excision-repair with purified human proteins: interaction between DNA polymerase β and the XRCC1 protein. EMBO J. 15, 6662–6670 (1996).
Thompson, L.H., Brookman, K.W., Jones, N.J., Allen, S.A. & Carrano, A.V. Molecular cloning of the human XRCC1 gene, which corrects defective DNA strand break repair and sister chromatid exchange. Mol. Cell. Biol. 10, 6160– 6171 (1990).
Shen, M,R., Zdzienicka, M.Z., Mohrenweiser, H., Thompson, L.H. & Thelen, M.P. Mutations in hamster single-strand break repair gene XRCC1 causing defective DNA repair. Nucleic Acids Res. 26, 1032–1037 (1998).
Mohrenweiser, H.W. et al. Refined mapping of the three DNA repair genes, ERCC1, ERCC2, and XRCC1, on human chromosome 19. Cytogenet. Cell Genet. 52, 11–14 ( 1989).
Price, E.A. et al. Rare microsatellite polymorphisms in the DNA repair genes XRCC1, XRCC3 and XRCC5 associated with cancer in patients of varying radiosensitivity. Somat. Cell Mol. Genet. 23, 237– 247 (1997).
Singhal, R.K, Prasad, R. & Wilson, S.H. DNA polymerase beta conducts the gap-filling step in uracil-initiated base excision repair in a bovine testis nuclear extract. J. Biol. Chem. 270, 949– 957 (1995).
Singhal, R.K. & Wilson, S.H. Short gap-filling synthesis by DNA polymerase beta is processive. J. Biol. Chem. 268 , 15906–15911 (1993).
Matsumoto, Y. & Kim, K. Excision of deoxyribose phosphate residues by DNA polymerase β during DNA repair. Science 269, 699–702 (1995).
Mullen, G.P. & Wilson, S.H. Repair activity in DNA polymerases: a structurally conserved helix-hairpin-helix motif in base excision repair enzymes and in DNA polymerase β. In Base excision repair (ed. Hickson, I.D.) 121–135 (Landes Bioscience, Austin, Texas; 1997).
Trucco, C. et al. A dual approach in the study of poly (ADP-ribose) polymerase: in vitro random mutagenesis and generation of deficient mice. Mol. Cell. Biochem. 193, 53–60 (1999).
Nash, R.A., Caldecott, K.W., Barnes, D.E. & Lindahl, T. XRCC1 protein interacts with one of two distinct forms of DNA ligase III. Biochemistry 36, 5207– 5211 (1997).
Cappelli, E. et al. Involvement of XRCC1 and DNA ligase III gene products in DNA base excision repair. J. Biol. Chem. 272, 23970–23975 (1997).
Bennett, R.A.O., Wilson, D.M., Wong, D. & Demple, B. Interaction of human apurinic endonuclease and DNA polymerase β in the base excision repair pathway. Proc. Natl. Acad. Sci. USA 94, 7166–7169 (1997).
Wilson, D.M. & Thompson, L.H. Life without DNA repair. Proc. Natl. Acad. Sci. USA 94, 12754– 12757 (1997).
Wilson, S.H. Mammalian base excision repair and DNA polymerase β. Mutation Res. 407, 203–215 ( 1998).
Kim, K., Biade, S. & Matsumoto, Y. Involvement of flap endonuclease 1 in base excision DNA repair. J. Biol. Chem. 273, 8842– 8848 (1998).
Klungland, A. & Lindahl, T. Second pathway for completion of human DNA base excision-repair: reconstitution with purified proteins and requirement for DNase IV (FEN1). EMBO J. 16, 3341–3348 (1997).
Dianov, G.L., Prasad, R., Wilson, S.H. & Bohr, V.A. Role of DNA polymerase beta in the excision step of long patch mammalian base excision repair. J. Biol. Chem. 274, 13741–13743 (1999).
Bork, P. et al. A superfamily of conserved domains in DNA damage-responsive cell cycle checkpoint proteins. FASEB J. 11, 68–76 (1997).
Zhang, X. et al. Structure of an XRCC1 BRCT domain: a new protein–protein interaction module. EMBO J. 17, 6404– 6411 (1998).
Liu, D-J., DeRose, E.F., Prasad, R., Wilson, S.H. & Mullen, G.P. Three-dimensional solution structure of the N-terminal domain of DNA polymerase β and mapping of the ssDNA interaction interface. Biochemistry 35, 6188– 6200 (1996).
Sawaya, M.R., Prasad, R., Wilson, S.H., Kraut, J. & Pelletier, H. Crystal structures of DNA polymerase β complexed with gapped and nicked DNA: evidence for an induced fit mechanism. Biochemistry 36, 11205–11215 (1997).
Doublie, S., Tabor, S., Long, A.M., Richardson, C.C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 A resolution. Nature 391, 251–258 (1998).
Li, Y., Korolev, S. & Waksman, G. Crystal structures of open and closed forms of binary and ternary complexes of the large fragment of Thermus aquaticus DNA polymerase I: structural basis for nucleotide incorporation. EMBO J. 17, 7514–25 ( 1998).
Cho, Y., Gorina, S., Jeffrey, P.D. & Pavletich, N.P. Crystal structure of a p53 tumor suppressor–DNA complex: understanding tumorigenic mutations. Science 265, 346– 355 (1994).
Zhou, P., Sun, L.J., Dotsch, V., Wagner, G. & Verdine, G.L. Solution structure of the core NFATC1/DNA complex. Cell 92, 687–696 (1998).
Gaskell, A., Crennell, S. & Taylor, G. The three domains of a bacterial sialidase: a β-propeller, an immunoglobulin module and a galactose-binding jelly-roll. Structure 3, 1197–1205 ( 1995).
Ito, N., Phillips, S.E., Yadav, K.D. & Knowles, P.F. Crystal structure of a free radical enzyme, galactose oxidase. J. Mol. Biol. 238, 794–814 (1994).
Bork, P., Holm, L. & Sander, C. The immunoglobulin fold. Structural classification, sequence patterns and common core. J. Mol. Biol. 242, 309–320 (1994).
Wolfe, S.A. et al. Unusual Rel-like architecture in the DNA-binding domain of the transcription factor NFATc. Nature 385, 172–176 (1997).
Chen, L., Glover, J.N., Hogan, P.G., Rao, A. & Harrison, S.C. Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 392, 42–48 ( 1998).
Ghosh, G., van Duyne, G., Ghosh, S. & Sigler, P.B. Structure of NFκB p50 homodimer bound to a kappa B site. Nature 373, 303–310 (1995).
Müller, C.W., Rey, F.A., Sodeoka, M., Verdine, G.L. & Harrison, S.C. Structure of the NFκB p50 homodimer bound to DNA. Nature 373, 311–317 (1995).
Osheroff, W.P., Jung, H.K., Beard, W.A., Wilson, S.H. & Kunkel, T.A. The fidelity of DNA polymerase β during distributive and processive DNA synthesis. J. Biol. Chem. 274, 3642–3650 (1999).
Canitrot, Y. et al. Overexpression of DNA polymerase beta in cells results in a mutator phenotype and a decreased sensitivity to anticancer drugs. Proc. Natl. Acad. Sci. USA 95, 12586– 12590 (1998).
Mohrenweiser, H.W. & Jones, I.M. Variation in DNA repair is a factor in cancer susceptibility: a paradigm for promises and perils of individual and population risk estimation? Mutation Res. 400, 15–24 (1998).
Prasad, R. et al. Functional analysis of the amino-terminal 8-kDa domain of DNA polymerase β as revealed by site-directed mutagenesis: DNA binding and 5′-deoxyribose phosphate lyase activities. J. Biol. Chem. 273, 11121–11126 ( 1998).
Bartels, C., Xia, T., Billeter, M., Güntert, P. & Wütrich, K. The program XEASY for computer-supported NMR spectral analysis of biological macromolecules. J. Biomol. NMR 6, 1–10 (1995).
Marintchev, A., Maciejewski, M.W. & Mullen, G.P. 1H, 15N, and 13C resonance assignments for the N-terminal 20 kDa domain of the DNA single-strand break repair protein XRCC1. J. Biomol. NMR 13, 393–394 (1999).
Güntert, P., Mumenthaler, C. & Wüthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Mumenthaler, C., Güntert, P., Braun, W. & Wüthrich, K. Automated combined assignment of NOESY spectra and three-dimensional protein structure determination. J. Biomol. NMR 10, 351–362 (1997).
Brünger, A.T. X-PLOR: a system for X-ray crystallography and NMR (Yale University Press, New Haven, Connecticut; 1992).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graphics 14, 51– 55 (1996).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Acknowledgements
We thank A. Robertson and S. Wilson for providing the overexpression clone of the palm-thumb domain of β-Pol. We thank M. Santos for synthesis of the oligonucleotides used for NMR studies. This research was supported by an NIH grant to G.P.M. and by NIH postdoctoral fellowship grants to M.W.M. and to B.P.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marintchev, A., Mullen, M., Maciejewski, M. et al. Solution structure of the single-strand break repair protein XRCC1 N-terminal domain. Nat Struct Mol Biol 6, 884–893 (1999). https://doi.org/10.1038/12347
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/12347
This article is cited by
-
Structural dynamics of DNA strand break sensing by PARP-1 at a single-molecule level
Nature Communications (2022)
-
XRCC1 Arg194Trp polymorphism and thyroid cancer
Journal of Endocrinological Investigation (2020)
-
Identification of an XRCC1 DNA binding activity essential for retention at sites of DNA damage
Scientific Reports (2019)
-
Association of DNA repair and xenobiotic pathway gene polymorphisms with genetic susceptibility to gastric cancer patients in West Bengal, India
Tumor Biology (2016)
-
New paradigms in the repair of oxidative damage in human genome: mechanisms ensuring repair of mutagenic base lesions during replication and involvement of accessory proteins
Cellular and Molecular Life Sciences (2015)