Abstract
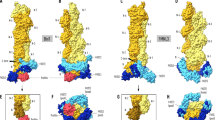
Gelation factor (ABP120) is one of the principal actin-cross-linking proteins of Dictyostelium discoideum. The extended molecule has an N-terminal 250-residue actin-binding domain and a rod constructed from six 100-residue repeats that have an Ig fold. The ability to dimerize is crucial to the actin cross-linking function of gelation factor and is mediated by the rod in which the two chains are arranged in an antiparallel fashion. We report the 2.2 Å resolution crystal structure of rod domains 5 and 6, which shows that dimerization is mediated primarily by rod domain 6 and is the result of a double edge-to-edge extension of β-sheets. Thus, contrary to earlier proposals, the chains of the dimeric gelation factor molecule overlap only within domain 6, and domains 1–5 do not pair with domains from the other chain. This information allows construction of a model of the gelation factor molecule and suggests how the chains in the related molecule filamin (ABP280) may interact.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Carlier, M.-F. & Pantaloni, D. J. Mol. Biol. 269, 459–467 (1997).
Steinmetz, M.O., Stoffer, D., Hoenger, A., Bremer, A. & Aebi, U. J. Struct. Biol. 119, 295– 320 (1997).
Puius, Y.A., Mahoney, N.M. & Almo, S.C. Curr. Opin. Cell Biol. 10, 23–34 (1998).
Fucini, P., Renner, C., Herberhold, C., Noegel, A.A. & Holak, T.A. Nature Struct. Biol. 4, 223–230 (1997).
McGough, A. Curr. Opin. Struct. Biol. 8, 166–176 (1998).
Rivero, F. et al. J. Cell Sci. 109, 2679– 2691 (1996).
Wachsstock, D.H., Schwarz, W.H. & Pollard, T.P. Biophys. J. 66, 801– 809 (1994).
Goldsmith, S.C. et al. Nature Struct. Biol. 4, 708– 712 (1997).
Brink, M. et al. J. Cell. Biol. 111, 1477– 1489 (1990).
Cox, D. et al. J. Cell Biol. 116, 943–955 (1992).
Cox, D., Risdale, J.A., Condeelis, J. & Hartwig, J. J. Cell Biol. 128, 819–835 (1995).
Witke, W., Schleicher, M. & Noegel, A.A. Cell, 68, 53– 62 (1992).
Gorlin, J.B. et al. J. Cell Biol. 111, 1089– 1105 (1990).
Cunningham, C.C. et al. Science 255, 325– 327 (1992).
Zhang, W., Han, S.W., McKeel, D.W., Goate, A. & Wu, J.Y. J. Neurosci. 18, 914–922 (1998).
Condeelis, J., Vahey, M., Carboni, J.M., DeMey, J. & Ogihara, S. J. Cell Biol. 99, 119s– 126s (1984).
Noegel, A.A., Rapp, S., Lottspeich, F., Schleicher, M., & Stewart, M. J. Cell. Biol. 109, 607– 618 (1989).
Bresnick, A.R., Warren, V. & Condeelis, J. J. Biol. Chem. 265, 9236– 9240 (1990).
Hartwig, J.H. In Protein profile: spectrin superfamily, Vol. 1 (ed. Hartwig, J.H.) 739–746 (Academic Press, New York; 1995).
Fucini, P., McCoy, A.J., Gomez-Ortiz, M., Schleicher, M., & Noegel, A.A. J. Struct. Biol. 120 , 192–195 (1997).
Jones, S. & Thornton, J.M. Proteins 21, 30–39 (1995).
Williams, A.F. & Barclay, A.N. Annu. Rev. Immunol. 6, 381–405 ( 1988).
Harpaz, Y. & Chothia, C. J. Mol. Biol. 238, 528–539 (1994).
Hamilton, J.A. et al. J. Biol. Chem. 268, 2416– 2424 (1993).
Colman, P.M. Adv. Immunol. 43, 99–132 (1988).
Improta, S. et al. J. Mol. Biol. 284, 761– 777 (1998).
Collaborative computational Project, Number 4. Acta Crystallogr. D 50, 760– 763 (1994).
de La Fortelle, E. & Bricogne, G. Methods Enzymol. 276, 472–493 ( 1997).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 ( 1991).
Esnouf, R.M. J. Mol. Graphics 15, 132–136 (1997).
Richardson, J.S. Adv. Protein Chem. 34, 167–339 (1981).
Acknowledgements
We thank A.Thompson, G. Leonard and A. Baker for assistance in collecting MAD data at BL14 at ESRF, Grenoble; R. Muller and H. Kent for assistance with protein preparation and light scattering; A. Murzin, E. de la Fortelle and J. Irwin for advice; and our colleagues in Martinsried and Cambridge, especially M. Schleicher, for their assistance. A.J.M. is a Howard Florey Fellow of the Royal Society of London. P.F. held a short-term EMBO Fellowship, and this work was supported in part by EC CHRX-CT93-0250 and a grant to A.N. from the Centre for Molecular Medicine, Cologne.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
McCoy, A., Fucini, P., Noegel, A. et al. Structural basis for dimerization of the Dictyostelium gelation factor (ABP120) rod. Nat Struct Mol Biol 6, 836–841 (1999). https://doi.org/10.1038/12296
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/12296
This article is cited by
-
Interactions between nascent proteins and the ribosome surface inhibit co-translational folding
Nature Chemistry (2021)
-
1H, 15N and 13C assignments of domain 5 of Dictyostelium discoideum gelation factor (ABP-120) in its native and 8M urea-denatured states
Biomolecular NMR Assignments (2009)
-
Actin binding domains direct actin-binding proteins to different cytoskeletal locations
BMC Cell Biology (2008)
-
Molecular basis of the C-terminal tail-to-tail assembly of the sarcomeric filament protein myomesin
The EMBO Journal (2008)
-
Structure of three tandem filamin domains reveals auto-inhibition of ligand binding
The EMBO Journal (2007)