Abstract
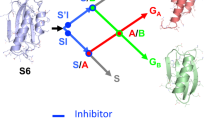
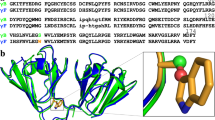
We use protein engineering and crystallography to simulate aspects of the early evolution of βγ-crystallins by observing how a single domain oligomerizes in response to changes in a sequence extension. The crystal structure of the C-terminal domain of γβ-crystallin with its four-residue C-terminal extension shows that the domain does not form a symmetric homodimer analogous to the two-domain pairing in γ-crystallins. Instead the C-terminal extension now forms heterologous interactions with other domains leading to the solvent exposure of the natural hydrophobic interface with a consequent loss in protein solubility. However, this domain truncated by just the C-terminal tyrosine forms a symmetric homodimer of domains in the crystal lattice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Dobson, C.M. Finding the right fold. Nature Struct. Biol. 2, 513–517 (1995).
Wistow, G.J. & Piatigorsky, J. Lens crystallins: The evolution and expression of proteins for a highly specialized tissue. Annu. Rev. Biochem. 57, 479–504 (1988).
Blundell, T. et al. The molecular structure and stability of the eye lens: X-ray analysis of γ-crystallin II. Nature 289, 771–777 (1981).
Bloemendal, H. & de Jong, W. W. Lens proteins and their genes. Prog. Nucleic Add Res. Mol. Biol. 41, 259–281 (1991).
Bax, B. et al. X-ray analysis of βB2-crystallin and evolution of oligomeric lens proteins. Nature 347, 776–780 (1990).
Lubsen, N.H., Aarts, H.J.M. & Schoenmakers, J.G.G. The evolution of lenticular proteins: the β- and γ-crystallin super gene family. Prog. Biophys. Molec. Biol. 51, 47–76 (1988).
Wistow, G. Evolution of a protein superfamily: relationships between vertebrate lens crystallins and microorganism dormancy proteins. J. Mol. Evol. 30, 140–145 (1990).
Bennett, M.J., Choe, S. & Eisenberg, D. Domain swapping: entangling alliances between proteins. Proc Natl. Acad. Sci. USA 91, 3127–3131 (1994).
Slingsby, C., Bateman, O.A. & Simpson, A. Motifs involved in protein-protein interactions. Mol. Biol. Rep. 17, 185–195 (1993).
Wang, J. et al. Structural basis of asymmetry in the human immunodeficiency virus type 1 reverse transcriptase heterodimer. Proc. Natl. Acad. Sci. USA 91, 7242–7246 (1994).
Fletterick, R.J. & Bazan, J.F. When one and one are not two. Nature Struct. Biol. 2, 721–723 (1995).
Bennett, M.J., Schlunegger, M.P. & Eisenberg, D. 3D Domain swapping—a mechanism for oligomer assembly. Protein Sci. 4, 2455–2468 (1995).
Mayr, E-M., Jaenicke, R. & Glockshuber, R. Domain interactions and connecting peptides in lens crystallins. J. Mol. Biol. 235, 84–88 (1994).
Rudolph, R., Siebendritt, R., Nesslauer, G., Sharma, A.K. & Jaenicke, R. Folding of an all-β protein: independent domain folding in γ||-crystallin from calf lens. Proc. Natl. Acad. Sci. USA 87, 4625–4629 (1990).
Magalhaes, A., Maigret, B., Hoflack, J., Gomes, J.N.F. & Scheraga, H.A. Contribution of unusual arginine-arginine short-range interactions to stabilization and recognition in proteins. J. Protein Chem. 13, 195–215 (1994).
De Jong, W.W., Lubsen, N.H. & Kraft, H.J. Molecular evolution of the eye lens. Prog. Ret. Eye Res. 13, 391–442 (1994).
Cooper, P.G., Carver, J.A., Aquilina, J.A., Ralston, G.B. &Truscott, R.J.W. A1 NMR spectroscopic comparison of γS and γB-crystallins. Exp. Eye Res. 59, 211–220 (1994).
Najmudin, S. et al. Structure of the bovine eye lens protein γB(gll)-crystallin at 1.47 Å. Acta Crystallogr. D49, 223–233 (1993).
Carver, J.A., Aquilina, J.A., Cooper, P.G., Williams, G.A. &Truscott, R.J.W. α-Crystallin: molecular chaperone and protein surfactant. Biochim. Biophys. Acta 1204, 195–206 (1994).
Otwinowski, Z. in Data Collection and Processing, Proceedings of the CCP4 Study Weekend, 29–30 January 1993 (ed. Sawyer, L, Isaacs, N. & Bailey, S) 56–62 (SERC, Daresbury Laboratory, UK, 1993).
Leslie, A.G.W., Brick, P. & Wonacott, A.T. Daresbury Laboratory information Quarterly for Protein Crystallography 18, 33–39 (S. E.R.C. Daresbury Laboratory, Warrington, U.K., 1986)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Messerschmidt, A. & Pflugrath, J.W. Crystal orientation and X-ray pattern prediction routines for area-detector diffractometer systems in macromolecular crystallography. J. Appl. Crystallogr. 20, 306–315 (1987).
Fox, G.C. & Holmes, K.C. An alternative method of solving the larger scaling equations of Hamilton, Rollett and Sparks. Acta Crystallogr. 20, 886–891 (1966).
French, G.S. & Wilson, K.S. On the treatment of negative intensity observations. Acta Crystallogr. A34, 517–525 (1978).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Navaza, J. AMoRe: an Automated Package for Molecular Replacement. Acta Crystallogr. A50, 157–163 (1994).
Crowther, R.A. in The Molecular Replacement Method (ed. Rossmann, M.G.) 173–178 (Gordon and Breach, New York, 1972)
Driessen, H.P.C. et al. Structure of oligomeric βB2 crystallin: an application of the T2 translation function to an asymmetric unit containing two dimers. Acta Crystallogr. B47, 987–997 (1991).
Wang, D., Driessen, H.P.C. & Tickle, I.J. MOLPACK: Molecular graphics for studying the packing of protein molecules in the crystallographic unit cell. J. Mol. Graph. 9, 28, 50–52 (1991).
Brünger, A.T., Kuryan, J. & Karplus, M. Crystallographic R-factor refinement by molecular-dynamics. Science 235, 458–460 (1987).
Jones, T.A. FRODO: a graphics model-building and refinement system for macromolecules. J. Appl. Crystallogr. 11, 268–272 (1978).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK-A program to check the stereochemical quality of protein structures J.Appl. Crystallogr. 26, 283–291 (1993).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for the building of protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47 110–119 (1991).
Lee, B. & Richards, F.M. The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55, 379–400 (1971).
Bailey, S. The CCP4 suite—programs for protein crystallography. Acta Crystallogr. D50 760–763 (1994).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Norledge, B., Mayr, EM., Glockshuber, R. et al. The X-ray structures of two mutant crystallin domains shed light on the evolution of multi-domain proteins. Nat Struct Mol Biol 3, 267–274 (1996). https://doi.org/10.1038/nsb0396-267
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0396-267
This article is cited by
-
The functional roles of the unstructured N- and C-terminal regions in αB-crystallin and other mammalian small heat-shock proteins
Cell Stress and Chaperones (2017)
-
Explosive Expansion of βγ-Crystallin Genes in the Ancestral Vertebrate
Journal of Molecular Evolution (2010)
-
Eye lens proteomics
Amino Acids (2006)
-
Structure of the crystallins
Eye (1999)
-
βγ-crystallin redux
Nature Structural & Molecular Biology (1996)