Key Points
-
The development of in vitro and in vivo modelling systems that accurately reflect the diverse mutational profiles of urological diseases is crucial
-
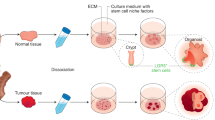
Patient-derived organoids in 3D culture systems enable the culturing of stem or progenitor cells for urological organs and they also mimic the in vivo microenvironment and stromal interactions of cancer cells
-
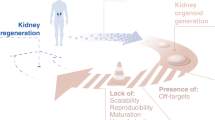
Nephron progenitor cells derived from human pluripotent stem cells have been successfully used to form complex multicellular kidney organoids, which have potential in disease modelling and drug screening
-
Organoids can be used to model disease; use of gene editing tools enables the study of the mechanisms of tumorigenesis and restoration of the functional defects caused by genetic lesions
Abstract
Technical advances in the development of organoid systems enable cell lines, primary adult cells, or stem or progenitor cells to develop into diverse, multicellular entities, which can self-renew, self-organize, and differentiate. These 3D organoid cultures have proven to be of value in increasing our understanding of the biology of disease and offer the potential of regenerative and genetic therapies. The successful application of 3D organoids derived from adult tissue into urological cancer research can further our understanding of these diseases and could also provide preclinical cancer models to realize the precision medicine paradigm by therapeutic screening of individual patient samples ex vivo. Kidney organoids derived from induced pluripotent stem cells provide personalized biomarkers, which can be correlated with genetic and clinical information. Organoid models can also improve our comprehension of aspects of particular diseases; for example, in prostate cancer, 3D organoids can aid in the identification of tumour-initiating cells from an epithelial cell lineage. Furthermore, kidney organoid differentiation from human pluripotent stem cells enables gene editing to model disease in kidney tubular epithelial cells. State-of-the-art human organoid cultures have potential as tools in basic and clinical research in renal, bladder, and prostatic diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Cancer Genome Atlas Research Network et al. Comprehensive molecular characterization of papillary renal-cell carcinoma. N. Engl. J. Med. 374, 135–145 (2016).
Davis, C. F. et al. The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell 26, 319–330 (2014).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Solit, D. B. et al. BRAF mutation predicts sensitivity to MEK inhibition. Nature 439, 358–362 (2006).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Cheon, D. J. & Orsulic, S. Mouse models of cancer. Annu. Rev. Pathol. 6, 95–119 (2011).
Hayflick, L. & Moorhead, P. S. The serial cultivation of human diploid cell strains. Exp. Cell Res. 25, 585–621 (1961).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
McKay, R. Stem cells in the central nervous system. Science 276, 66–71 (1997).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Liu, X. et al. ROCK inhibitor and feeder cells induce the conditional reprogramming of epithelial cells. Am. J. Pathol. 180, 599–607 (2012).
Chapman, S., Liu, X., Meyers, C., Schlegel, R. & McBride, A. A. Human keratinocytes are efficiently immortalized by a Rho kinase inhibitor. J. Clin. Invest. 120, 2619–2626 (2010).
Harrison, R. G., Greenman, M. J., Mall, F. P. & Jackson, C. M. Observations of the living developing nerve fiber. Anat. Rec. (Hoboken) 1, 116–128 (1907).
Gattazzo, F., Urciuolo, A. & Bonaldo, P. Extracellular matrix: a dynamic microenvironment for stem cell niche. Biochim. Biophys. Acta 1840, 2506–2519 (2014).
Shamir, E. R. & Ewald, A. J. Three-dimensional organotypic culture: experimental models of mammalian biology and disease. Nat. Rev. Mol. Cell Biol. 15, 647–664 (2014).
Rookmaaker, M. B., Schutgens, F., Verhaar, M. C. & Clevers, H. Development and application of human adult stem or progenitor cell organoids. Nat. Rev. Nephrol. 11, 546–554 (2015).
O'Brien, L. E. et al. Rac1 orientates epithelial apical polarity through effects on basolateral laminin assembly. Nat. Cell Biol. 3, 831–838 (2001).
Bryant, D. M. et al. A molecular network for de novo generation of the apical surface and lumen. Nat. Cell Biol. 12, 1035–1045 (2010).
Yagi, S., Matsuda, M. & Kiyokawa, E. Suppression of Rac1 activity at the apical membrane of MDCK cells is essential for cyst structure maintenance. EMBO Rep. 13, 237–243 (2012).
Smith, Y. C., Grande, K. K., Rasmussen, S. B. & O'Brien, A. D. Novel three-dimensional organoid model for evaluation of the interaction of uropathogenic Escherichia coli with terminally differentiated human urothelial cells. Infect. Immun. 74, 750–757 (2006).
Unsworth, B. R. & Lelkes, P. I. Growing tissues in microgravity. Nat. Med. 4, 901–907 (1998).
Tyson, D. R., Inokuchi, J., Tsunoda, T., Lau, A. & Ornstein, D. K. Culture requirements of prostatic epithelial cell lines for acinar morphogenesis and lumen formation in vitro: role of extracellular calcium. Prostate 67, 1601–1613 (2007).
Lang, S. H. et al. Experimental prostate epithelial morphogenesis in response to stroma and three-dimensional matrigel culture. Cell Growth Differ. 12, 631–640 (2001).
Garraway, L. A. et al. Intermediate basal cells of the prostate: in vitro and in vivo characterization. Prostate 55, 206–218 (2003).
Xin, L., Lukacs, R. U., Lawson, D. A., Cheng, D. & Witte, O. N. Self-renewal and multilineage differentiation in vitro from murine prostate stem cells. Stem Cells 25, 2760–2769 (2007).
Rinkevich, Y. et al. In vivo clonal analysis reveals lineage-restricted progenitor characteristics in mammalian kidney development, maintenance, and regeneration. Cell Rep. 7, 1270–1283 (2014).
Pinto, D., Gregorieff, A., Begthel, H. & Clevers, H. Canonical Wnt signals are essential for homeostasis of the intestinal epithelium. Genes Dev. 17, 1709–1713 (2003).
Kim, K. A. et al. Mitogenic influence of human R-spondin1 on the intestinal epithelium. Science 309, 1256–1259 (2005).
Howe, J. R. et al. Mutations in the SMAD4/DPC4 gene in juvenile polyposis. Science 280, 1086–1088 (1998).
Howe, J. R. et al. Germline mutations of the gene encoding bone morphogenetic protein receptor 1A in juvenile polyposis. Nat. Genet. 28, 184–187 (2001).
Haramis, A. P. et al. De novo crypt formation and juvenile polyposis on BMP inhibition in mouse intestine. Science 303, 1684–1686 (2004).
Konturek, J. W., Bielanski, W., Konturek, S. J., Bogdal, J. & Oleksy, J. Distribution and release of epidermal growth factor in man. Gut 30, 1194–1200 (1989).
Dignass, A. U. & Sturm, A. Peptide growth factors in the intestine. Eur. J. Gastroenterol. Hepatol. 13, 763–770 (2001).
Schwank, G. et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13, 653–658 (2013).
Rock, J. R. et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl Acad. Sci. USA 106, 12771–12775 (2009).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology 141, 1762–1772 (2011).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Barker, N. et al. Lgr5(+ve) stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Xia, Y. et al. Directed differentiation of human pluripotent cells to ureteric bud kidney progenitor-like cells. Nat. Cell Biol. 15, 1507–1515 (2013).
Takasato, M. et al. Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nat. Cell Biol. 16, 118–126 (2014).
Matulay, J. T. et al. Genetic mutations in patient-derived bladder tumor organoids mimic parental tumor samples [abstract PD38-07]. J. Urol. 195, e926 (2016).
Karthaus, W. R. et al. Identification of multipotent luminal progenitor cells in human prostate organoid cultures. Cell 159, 163–175 (2014).
Drost, J. et al. Organoid culture systems for prostate epithelial and cancer tissue. Nat. Protoc. 11, 347–358 (2016).
Gao, D. et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 159, 176–187 (2014).
Batchelder, C. A., Martinez, M. L., Duru, N., Meyers, F. J. & Tarantal, A. F. Three dimensional culture of human renal cell carcinoma organoids. PLoS ONE 10, e0136758 (2015).
Lobo, N. C. et al. Efficient generation of patient-matched malignant and normal primary cell cultures from clear cell renal cell carcinoma patients: clinically relevant models for research and personalized medicine. BMC Cancer 16, 485 (2016).
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Antonica, F. et al. Generation of functional thyroid from embryonic stem cells. Nature 491, 66–71 (2012).
Koehler, K. R., Mikosz, A. M., Molosh, A. I., Patel, D. & Hashino, E. Generation of inner ear sensory epithelia from pluripotent stem cells in 3D culture. Nature 500, 217–221 (2013).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Suga, H. et al. Self-formation of functional adenohypophysis in three-dimensional culture. Nature 480, 57–62 (2011).
Little, M. H. & McMahon, A. P. Mammalian kidney development: principles, progress, and projections. Cold Spring Harb. Perspect. Biol. 4, a008300 (2012).
Ader, M. & Tanaka, E. M. Modeling human development in 3D culture. Curr. Opin. Cell Biol. 31, 23–28 (2014).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015).
Takasato, M. & Little, M. H. The origin of the mammalian kidney: implications for recreating the kidney in vitro. Development 142, 1937–1947 (2015).
Takasato, M., Er, P. X., Chiu, H. S. & Little, M. H. Generation of kidney organoids from human pluripotent stem cells. Nat. Protoc. 11, 1681–1692 (2016).
Mugford, J. W., Sipila, P., McMahon, J. A. & McMahon, A. P. Osr1 expression demarcates a multi-potent population of intermediate mesoderm that undergoes progressive restriction to an Osr1-dependent nephron progenitor compartment within the mammalian kidney. Dev. Biol. 324, 88–98 (2008).
Calderon-Gierszal, E. L. & Prins, G. S. Directed differentiation of human embryonic stem cells into prostate organoids in vitro and its perturbation by low-dose bisphenol A exposure. PLoS ONE 10, e0133238 (2015).
Simon, J. Ectropia vesicæ; (absence, of the anterior walls of the bladder and pubic abdominal parietes); operation for directing the orifices of the ureters into the rectum; temporary success; subsequent death; autopsy. Lancet 60, 568–570 (1852).
van Hemelrijck, M., Thorstenson, A., Smith, P., Adolfsson, J. & Akre, O. Risk of in-hospital complications after radical cystectomy for urinary bladder carcinoma: population-based follow-up study of 7608 patients. BJU Int. 112, 1113–1120 (2013).
Osborn, S. L. et al. Induction of human embryonic and induced pluripotent stem cells into urothelium. Stem Cells Transl Med. 3, 610–619 (2014).
Kang, M., Kim, H. H. & Han, Y. M. Generation of bladder urothelium from human pluripotent stem cells under chemically defined serum- and feeder-free system. Int. J. Mol. Sci. 15, 7139–7157 (2014).
O'Driscoll, L. et al. Phenotypic and global gene expression profile changes between low passage and high passage MIN-6 cells. J. Endocrinol. 191, 665–676 (2006).
Beltran, H. et al. Molecular characterization of neuroendocrine prostate cancer and identification of new drug targets. Cancer Discov. 1, 487–495 (2011).
Scher, H. I. et al. Circulating tumor cell biomarker panel as an individual-level surrogate for survival in metastatic castration-resistant prostate cancer. J. Clin. Oncol. 33, 1348–1355 (2015).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Shin, K. et al. Hedgehog/Wnt feedback supports regenerative proliferation of epithelial stem cells in bladder. Nature 472, 110–114 (2011).
Ohishi, T., Koga, F. & Migita, T. Bladder cancer stem-like cells: their origin and therapeutic perspectives. Int. J. Mol. Sci. 17, 43 (2016).
Li, Z. et al. 3D culture supports long-term expansion of mouse and human nephrogenic progenitors. Cell Stem Cell 19, 516–529 (2016).
Pavlovich, C. P. & Schmidt, L. S. Searching for the hereditary causes of renal-cell carcinoma. Nat. Rev. Cancer 4, 381–393 (2004).
Lee, S. H. & Shen, M. M. Cell types of origin for prostate cancer. Curr. Opin. Cell Biol. 37, 35–41 (2015).
Lawson, D. A. et al. Basal epithelial stem cells are efficient targets for prostate cancer initiation. Proc. Natl Acad. Sci. USA 107, 2610–2615 (2010).
Stoyanova, T. et al. Prostate cancer originating in basal cells progresses to adenocarcinoma propagated by luminal-like cells. Proc. Natl Acad. Sci. USA 110, 20111–20116 (2013).
Wang, Z. A. et al. Lineage analysis of basal epithelial cells reveals their unexpected plasticity and supports a cell-of-origin model for prostate cancer heterogeneity. Nat. Cell Biol. 15, 274–283 (2013).
Choi, N., Zhang, B., Zhang, L., Ittmann, M. & Xin, L. Adult murine prostate basal and luminal cells are self-sustained lineages that can both serve as targets for prostate cancer initiation. Cancer Cell 21, 253–265 (2012).
Wang, X. et al. A luminal epithelial stem cell that is a cell of origin for prostate cancer. Nature 461, 495–500 (2009).
Chua, C. W. et al. Single luminal epithelial progenitors can generate prostate organoids in culture. Nat. Cell Biol. 16, 951–961 (2014).
Agarwal, S. et al. Identification of different classes of luminal progenitor cells within prostate tumors. Cell Rep. 13, 2147–2158 (2015).
Liu, X. et al. Low CD38 identifies progenitor-like inflammation-associated luminal cells that can initiate human prostate cancer and predict poor outcome. Cell Rep. 17, 2596–2606 (2016).
Park, J. W. et al. Prostate epithelial cell of origin determines cancer differentiation state in an organoid transformation assay. Proc. Natl Acad. Sci. USA 113, 4482–4487 (2016).
Sluka, P. & Davis, I. D. Cell mates: paracrine and stromal targets for prostate cancer therapy. Nat. Rev. Urol. 10, 441–451 (2013).
Gaggioli, C. et al. Fibroblast-led collective invasion of carcinoma cells with differing roles for RhoGTPases in leading and following cells. Nat. Cell Biol. 9, 1392–1400 (2007).
Scriven, S. D., Booth, C., Thomas, D. F., Trejdosiewicz, L. K. & Southgate, J. Reconstitution of human urothelium from monolayer cultures. J. Urol. 158, 1147–1152 (1997).
Varley, C. L. & Southgate, J. Organotypic and 3D reconstructed cultures of the human bladder and urinary tract. Methods Mol. Biol. 695, 197–211 (2011).
Wan, X. et al. Prostate cancer cell-stromal cell crosstalk via FGFR1 mediates antitumor activity of dovitinib in bone metastases. Sci. Transl Med. 6, 252ra122 (2014).
Fong, E. L. et al. A 3D in vitro model of patient-derived prostate cancer xenograft for controlled interrogation of in vivo tumor-stromal interactions. Biomaterials 77, 164–172 (2016).
Valta, M. P. et al. FGF-8 is involved in bone metastasis of prostate cancer. Int. J. Cancer 123, 22–31 (2008).
Åkerfelt, M. et al. Automated tracking of tumor-stroma morphology in microtissues identifies functional targets within the tumor microenvironment for therapeutic intervention. Oncotarget 6, 30035–30056 (2015).
Hahn, W. C. et al. Creation of human tumour cells with defined genetic elements. Nature 400, 464–468 (1999).
Johnson, P. J., Coussens, P. M., Danko, A. V. & Shalloway, D. Overexpressed pp60c-src can induce focus formation without complete transformation of NIH 3T3 cells. Mol. Cell. Biol. 5, 1073–1083 (1985).
Bonner, T. I. et al. Structure and biological activity of human homologs of the raf/mil oncogene. Mol. Cell. Biol. 5, 1400–1407 (1985).
Bello, D., Webber, M. M., Kleinman, H. K., Wartinger, D. D. & Rhim, J. S. Androgen responsive adult human prostatic epithelial cell lines immortalized by human papillomavirus 18. Carcinogenesis 18, 1215–1223 (1997).
Zhu, J., Nguyen, M. T., Nakamura, E., Yang, J. & Mackem, S. Cre-mediated recombination can induce apoptosis in vivo by activating the p53 DNA damage-induced pathway. Genesis 50, 102–111 (2012).
Matano, M. et al. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat. Med. 21, 256–262 (2015).
Drost, J. et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature 521, 43–47 (2015).
Freedman, B. S. et al. Modelling kidney disease with CRISPR-mutant kidney organoids derived from human pluripotent epiblast spheroids. Nat. Commun. 6, 8715 (2015).
Carver, B. S. et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat. Genet. 41, 619–624 (2009).
Yui, S. et al. Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nat. Med. 18, 618–623 (2012).
Toyohara, T. et al. Cell therapy using human induced pluripotent stem cell-derived renal progenitors ameliorates acute kidney injury in mice. Stem Cells Transl Med. 4, 980–992 (2015).
Imberti, B. et al. Renal progenitors derived from human iPSCs engraft and restore function in a mouse model of acute kidney injury. Sci. Rep. 5, 8826 (2015).
Kabadi, A. M., Ousterout, D. G., Hilton, I. B. & Gersbach, C. A. Multiplex CRISPR/Cas9-based genome engineering from a single lentiviral vector. Nucleic Acids Res. 42, e147 (2014).
Deiss, F. et al. Platform for high-throughput testing of the effect of soluble compounds on 3D cell cultures. Anal. Chem. 85, 8085–8094 (2013).
Hakanson, M., Textor, M. & Charnley, M. Engineered 3D environments to elucidate the effect of environmental parameters on drug response in cancer. Integr. Biol. (Camb.) 3, 31–38 (2011).
Weigelt, B., Lo, A. T., Park, C. C., Gray, J. W. & Bissell, M. J. HER2 signaling pathway activation and response of breast cancer cells to HER2-targeting agents is dependent strongly on the 3D microenvironment. Breast Cancer Res. Treat. 122, 35–43 (2010).
Cheung, K. J., Gabrielson, E., Werb, Z. & Ewald, A. J. Collective invasion in breast cancer requires a conserved basal epithelial program. Cell 155, 1639–1651 (2013).
Calon, A. et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat. Genet. 47, 320–329 (2015).
Wyatt, A. W. et al. Heterogeneity in the inter-tumor transcriptome of high risk prostate cancer. Genome Biol. 15, 426 (2014).
Boutros, P. C. et al. Spatial genomic heterogeneity within localized, multifocal prostate cancer. Nat. Genet. 47, 736–745 (2015).
Kumar, A. et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat. Med. 22, 369–378 (2016).
Dayyani, F., Gallick, G. E., Logothetis, C. J. & Corn, P. G. Novel therapies for metastatic castrate-resistant prostate cancer. J. Natl Cancer Inst. 103, 1665–1675 (2011).
Acknowledgements
We thank Margaret McPartland for providing editorial input. Funding was providing by the National Institutes of Health (5K08CA140946 and NIH Cancer Center Core Grant P30 CA008748, and SPORE in Prostate Cancer (5P50CA092629-15)), US Department of Defense (W81XWH-10-1-0197), the Geoffrey Beene Cancer Center, the STARR Cancer Consortium (I8-A722), a Stand Up To Cancer–Prostate Cancer Foundation Prostate Dream Team Translational Research Grant (SU2CAACR-DT0712), and a Prostate Cancer Foundation Movember Challenge Grant.
Author information
Authors and Affiliations
Contributions
All the authors researched data for the article, substantially contributed to discussion of the content, wrote and reviewed and/or edited the article before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Wang, S., Gao, D. & Chen, Y. The potential of organoids in urological cancer research. Nat Rev Urol 14, 401–414 (2017). https://doi.org/10.1038/nrurol.2017.65
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrurol.2017.65
This article is cited by
-
Novel organoid construction strategy for non-involuting congenital hemangioma for drug validation
Journal of Biological Engineering (2023)
-
Individualisierte Präzisionsmedizin
Die Urologie (2023)
-
Use and application of organ-on-a-chip platforms in cancer research
Journal of Cell Communication and Signaling (2023)
-
Expansion of mouse castration-resistant intermediate prostate stem cells in vitro
Stem Cell Research & Therapy (2022)
-
Patient-derived organoid (PDO) platforms to facilitate clinical decision making
Journal of Translational Medicine (2021)