Key Points
-
Cellular 'phenotypic' screening is a powerful unbiased method for elucidating and developing drugs for poorly understood processes such as neurodegeneration
-
Neurons and glial cells can now be generated from patient-derived induced pluripotent stem cells (iPSCs)
-
Commensurate progress in our ability to manipulate ('edit') human genomes paves the way for iPSC technology to be used for unbiased genetic and small-molecule screening of patient-derived cells
-
The main roadblock for the neurodegeneration field is the identification of robust and disease-relevant cellular phenotypes
-
A path toward screening of patient-derived cells and target identification can be suggested by investigations in model organisms, including the most screenable eukaryotic cell, baker's yeast
Abstract
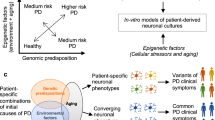
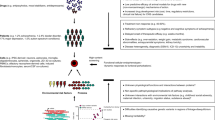
In the absence of a single preventive or disease-modifying strategy, neurodegenerative diseases are becoming increasingly prevalent in our ageing population. The mechanisms underlying neurodegeneration are poorly understood, making the target-based drug screening strategies that are employed by the pharmaceutical industry fraught with difficulty. However, phenotypic screening in neurons and glia derived from patients is now conceivable through unprecedented developments in reprogramming, transdifferentiation, and genome editing. We outline progress in this nascent field, but also consider the formidable hurdles to identifying robust, disease-relevant and screenable cellular phenotypes in patient-derived cells. We illustrate how analysis in the simple baker's yeast cell Saccharaomyces cerevisiae is driving discovery in patient-derived neurons, and how approaches in this model organism can establish a paradigm to guide the development of stem cell-based phenotypic screens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
10 facts on dementia. World Health Organization [online], (2015).
Lowe, J., Mirra, S. S., Hyman, B. T. & Dickson, D. W. in Greenfield's Neuropathology 8th edn (eds Love, S. et al.) 1031–1152 (CRC Press, 2008).
Goedert, M., Spillantini, M. G., Del Tredici, K. & Braak, H. 100 years of Lewy pathology. Nat. Rev. Neurol. 9, 13–24 (2013).
Yamanaka, S. Induced pluripotent stem cells: past, present, and future. Cell Stem Cell 10, 678–684 (2012).
Yang, N. et al. Generation of oligodendroglial cells by direct lineage conversion. Nat. Biotechnol. 31, 434–439 (2013).
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010).
Swinney, D. C. & Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 10, 507–519 (2011).
Lee, B.-H., Finley, D. & King, R. W. A high-throughput screening method for identification of inhibitors of the deubiquitinating enzyme USP14. Curr. Protoc. Chem. Biol. 4, 311–330 (2012).
Lee, B.-H. et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 467, 179–184 (2010).
Taipale, M. et al. Chaperones as thermodynamic sensors of drug-target interactions reveal kinase inhibitor specificities in living cells. Nat. Biotechnol. 31, 630–637 (2013).
Treusch, S. et al. Functional links between Aβ toxicity, endocytic trafficking, and Alzheimer's disease risk factors in yeast. Science 334, 1241–1245 (2011).
Volpicelli-Daley, L. A. et al. Exogenous α-synuclein fibrils induce Lewy body pathology leading to synaptic dysfunction and neuron death. Neuron 72, 57–71 (2011).
Chung, C. Y. et al. Identification and rescue of α-synuclein toxicity in Parkinson patient-derived neurons. Science 342, 983–987 (2013).
Khurana, V. & Lindquist, S. Modelling neurodegeneration in Saccharomyces cerevisiae: why cook with baker's yeast? Nat. Rev. Neurosci. 11, 436–449 (2010).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat. Genet. 45, 1452–1458 (2013).
Guerreiro, R., Brás, J. & Hardy, J. SnapShot: Genetics of ALS and FTD. Cell 160, 798–798e1 (2015).
Brás, J., Guerreiro, R. & Hardy, J. SnapShot: enetics of Parkinson's disease. Cell 160, 570–570e1 (2015).
Huang, Z. et al. A functional variomics tool for discovering drug-resistance genes and drug targets. Cell Rep. 3, 577–585 (2013).
Narayan, P., Ehsani, S. & Lindquist, S. Combating neurodegenerative disease with chemical probes and model systems. Nat. Chem. Biol. 10, 911–920 (2014).
Yeger-Lotem, E. et al. Bridging high-throughput genetic and transcriptional data reveals cellular responses to alpha-synuclein toxicity. Nat. Genet. 41, 316–323 (2009).
Cooper, A. A. et al. α-Synuclein blocks ER–Golgi traffic and Rab1 rescues neuron loss in Parkinson's models. Science 313, 324–328 (2006).
Gitler, A. D. et al. α-Synuclein is part of a diverse and highly conserved interaction network that includes PARK9 and manganese toxicity. Nat. Genet. 41, 308–315 (2009).
Treusch, S., Cyr, D. M. & Lindquist, S. Amyloid deposits: protection against toxic protein species? Cell Cycle 8, 1668–1674 (2009).
Gitler, A. D. et al. The Parkinson's disease protein α-synuclein disrupts cellular Rab homeostasis. Proc. Natl Acad. Sci. USA 105, 145–150 (2008).
Soper, J. H., Kehm, V., Burd, C. G., Bankaitis, V. A. & Lee, V. M. Aggregation of α-synuclein in S. cerevisiae is associated with defects in endosomal trafficking and phospholipid biosynthesis. J. Mol. Neurosci. 43, 391–405 (2011).
Thayanidhi, N. et al. α-Synuclein delays endoplasmic reticulum (ER)-to-Golgi transport in mammalian cells by antagonizing ER/Golgi SNAREs. Mol. Biol. Cell 21, 1850–1863 (2010).
Mazzulli, J. R. et al. Gaucher disease glucocerebrosidase and α-synuclein form a bidirectional pathogenic loop in synucleinopathies. Cell 146, 37–52 (2011).
Macleod, D. A. et al. RAB7L1 interacts with LRRK2 to modify intraneuronal protein sorting and Parkinson's disease risk. Neuron 77, 425–439 (2013).
Matlack, K. E. et al. Clioquinol promotes the degradation of metal-dependent amyloid-β (Aβ) oligomers to restore endocytosis and ameliorate Aβ toxicity. Proc. Natl Acad. Sci. USA 111, 4013–4018 (2014).
Dhungel, N. et al. Parkinson's disease genes VPS35 and EIF4G1 interact genetically and converge on α-synuclein. Neuron 85, 76–87 (2015).
Elden, A. C. et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466, 1069–1075 (2010).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Krencik, R., Weick, J. P., Liu, Y., Zhang, Z.-J. & Zhang, S.-C. Specification of transplantable astroglial subtypes from human pluripotent stem cells. Nat. Biotechnol. 29, 528–534 (2011).
Wang, S. et al. Human iPSC-derived oligodendrocyte progenitor cells can myelinate and rescue a mouse model of congenital hypomyelination. Cell Stem Cell 12, 252–264 (2013).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Okita, K. et al. An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells 31, 458–466 (2013).
Zhou, T. et al. Generation of human induced pluripotent stem cells from urine samples. Nat. Protoc. 7, 2080–2089 (2012).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Yu, J. et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 324, 797–801 (2009).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Sandoe, J. & Eggan, K. Opportunities and challenges of pluripotent stem cell neurodegenerative disease models. Nat. Neurosci. 16, 780–789 (2013).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Jack, C. R. Jr et al. Tracking pathophysiological processes in Alzheimer's disease: an updated hypothetical model of dynamic biomarkers. Lancet Neurol. 12, 207–216 (2013).
Shulman, J. M., De Jager, P. L. & Feany, M. B. Parkinson's disease: genetics and pathogenesis. Annu. Rev. Pathol. 6, 193–222 (2011).
Dickson, D. W. et al. Neuropathology of non-motor features of Parkinson disease. Parkinsonism Relat. Disord. 15 (Suppl. 3), S1–S5 (2009).
Cooper, O. et al. Pharmacological rescue of mitochondrial deficits in iPSC-derived neural cells from patients with familial Parkinson's disease. Sci. Transl. Med. 4, 141ra90 (2012).
Nguyen, H. N. et al. LRRK2 mutant iPSC-derived DA neurons demonstrate increased susceptibility to oxidative stress. Cell Stem Cell 8, 267–280 (2011).
Ryan, S. D. et al. Isogenic human iPSC Parkinson's model shows nitrosative stress-induced dysfunction in MEF2-PGC1α transcription. Cell 155, 1351–1364 (2013).
Miller, J. D. et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell Stem Cell 13, 691–705 (2013).
Lee, G. et al. Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature 461, 402–406 (2009).
Lee, G. et al. Large-scale screening using familial dysautonomia induced pluripotent stem cells identifies compounds that rescue IKBKAP expression. Nat. Biotechnol. 30, 1244–1248 (2012).
Ku, S. et al. Friedreich's ataxia induced pluripotent stem cells model intergenerational GAA·TTC triplet repeat instability. Cell Stem Cell 7, 631–637 (2010).
Hick, A. et al. Neurons and cardiomyocytes derived from induced pluripotent stem cells as a model for mitochondrial defects in Friedreich's ataxia. Dis. Model. Mech. 6, 608–621 (2013).
Almeida, S. et al. Induced pluripotent stem cell models of progranulin-deficient frontotemporal dementia uncover specific reversible neuronal defects. Cell Rep. 2, 789–798 (2012).
Sanders, L. H. et al. LRRK2 mutations cause mitochondrial DNA damage in iPSC-derived neural cells from Parkinson's disease patients: reversal by gene correction. Neurobiol. Dis. 62, 381–386 (2014).
Ebert, A. D. et al. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 457, 277–280 (2009).
Sendtner, M. Therapy development in spinal muscular atrophy. Nat. Neurosci. 13, 795–799 (2010).
Naryshkin, N. A. et al. Motor neuron disease. SMN2 splicing modifiers improve motor function and longevity in mice with spinal muscular atrophy. Science 345, 688–693 (2014).
Harper, S. Q. et al. RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model. Proc. Natl Acad. Sci. USA 102, 5820–5825 (2005).
Xia, H. et al. RNAi suppresses polyglutamine-induced neurodegeneration in a model of spinocerebellar ataxia. Nat. Med. 10, 816–820 (2004).
Keiser, M. S., Geoghegan, J. C., Boudreau, R. L., Lennox, K. A. & Davidson, B. L. RNAi or overexpression: alternative therapies for Spinocerebellar Ataxia Type 1. Neurobiol. Dis. 56, 6–13 (2013).
Sareen, D. et al. Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Sci. Transl. Med. 5, 208ra149 (2013).
Xu, X. et al. Prevention of β-amyloid induced toxicity in human iPS cell-derived neurons by inhibition of Cyclin-dependent kinases and associated cell cycle events. Stem Cell Res. 10, 213–227 (2013).
Yahata, N. et al. Anti-Aβ drug screening platform using human iPS cell-derived neurons for the treatment of Alzheimer's disease. PLoS ONE 6, e25788 (2011).
Kondo, T. et al. Modeling Alzheimer's disease with iPSCs reveals stress phenotypes associated with Intracellular Aβ and differential drug responsiveness. Cell Stem Cell 12, 487–496 (2013).
Burkhardt, M. F. et al. A cellular model for sporadic ALS using patient-derived induced pluripotent stem cells. Mol. Cell. Neurosci. 56, 355–364 (2013).
Barmada, S. J. et al. Autophagy induction enhances TDP43 turnover and survival in neuronal ALS models. Nat. Chem. Biol. 10, 677–685 (2014).
Skibinski, G. & Finkbeiner, S. Drug discovery in Parkinson's disease—update and developments in the use of cellular models. Int. J. High Throughput Screen. 2011, 15–25 (2011).
Egawa, N. et al. Drug screening for ALS using patient-specific induced pluripotent stem cells. Sci. Transl. Med. 4, 145ra104 (2012).
Yang, Y. M. et al. A small molecule screen in stem-cell-derived motor neurons identifies a kinase inhibitor as a candidate therapeutic for ALS. Cell Stem Cell 12, 713–726 (2013).
Höing, S. et al. Discovery of inhibitors of microglial neurotoxicity acting through multiple mechanisms using a stem-cell-based phenotypic assay. Cell Stem Cell 11, 620–632 (2012).
Tardiff, D. F., Khurana, V., Chung, C. Y. & Lindquist, S. From yeast to patient neurons and back again: powerful new discovery platforms. Mov. Disord. 10, 1231–1240 (2014).
Tardiff, D. F. et al. Yeast reveal a “druggable” Rsp5/Nedd4 network that ameliorates α-synuclein toxicity in neurons. Science 342, 979–983 (2013).
Auluck, P. K., Caraveo, G. & Lindquist, S. α-Synuclein: membrane interactions and toxicity in Parkinson's disease. Annu. Rev. Cell Dev. Biol. 26, 211–233 (2010).
Hu, G. & Luo, J. A primer on using pooled shRNA libraries for functional genomic screens. Acta Biochim. Biophys. Sin. (Shanghai) 44, 103–112 (2012).
Hasson, S. A. et al. High-content genome-wide RNAi screens identify regulators of parkin upstream of mitophagy. Nature 504, 291–295 (2013).
Seibler, P. et al. Mitochondrial Parkin recruitment is impaired in neurons derived from mutant PINK1 induced pluripotent stem cells. J. Neurosci. 31, 5970–5976 (2011).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84 (2014).
Leeb, M. & Wutz, A. Derivation of haploid embryonic stem cells from mouse embryos. Nature 479, 131–134 (2011).
Elling, U. et al. Forward and reverse genetics through derivation of haploid mouse embryonic stem cells. Cell Stem Cell 9, 563–574 (2011).
Son, E. Y. et al. Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell 9, 205–218 (2011).
Gupta, R. M. & Musunuru, K. Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J. Clin. Invest. 124, 4154–4161 (2014).
Acknowledgements
V.K. has received an American Brain and Parkinson's Disease Foundations Clinician–Scientist Development Award, and is also funded by the Harvard NeuroDiscovery Center and the Multiple System Atrophy Coalition. D.F.T. is the recipient of National Research Service Award fellowship F32NS061419, and is also funded by the Thome Memorial Foundation. C.Y.C. is funded by the National Institute on Aging. S.L. is funded by the Howard Hughes Medical Institute, the JPB Foundation and the Eleanor Schwartz Charitable Foundation. We thank Linda Clayton for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
V.K., D.F.T. and C.Y.C. researched data for the article, and reviewed and edited the manuscript before submission. V.K., D.F.T. and S.L. made substantial contributions to discussions of the content. All authors contributed equally to writing the article.
Corresponding authors
Ethics declarations
Competing interests
V.K., D.F.T., C.Y.C. and S.L. are scientific co-founders of Yumanity Therapeutics, a company involved in neurodegenerative disease drug discovery.
Rights and permissions
About this article
Cite this article
Khurana, V., Tardiff, D., Chung, C. et al. Toward stem cell-based phenotypic screens for neurodegenerative diseases. Nat Rev Neurol 11, 339–350 (2015). https://doi.org/10.1038/nrneurol.2015.79
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneurol.2015.79
This article is cited by
-
A pesticide and iPSC dopaminergic neuron screen identifies and classifies Parkinson-relevant pesticides
Nature Communications (2023)
-
Revisiting a Telencephalic Extent of the Ascending Reticular Activating System
Cellular and Molecular Neurobiology (2023)
-
Clinical Trial-Ready Patient Cohorts for Multiple System Atrophy: Coupling Biospecimen and iPSC Banking to Longitudinal Deep-Phenotyping
The Cerebellum (2022)
-
A11, a novel diaryl acylhydrazone derivative, exerts neuroprotection against ischemic injury in vitro and in vivo
Acta Pharmacologica Sinica (2019)
-
Induced pluripotent stem cells in multiple system atrophy: recent developments and scientific challenges
Clinical Autonomic Research (2019)