Key Points
-
Urinary extracellular vesicles comprise a wide range of biologically distinct structures with contents that are a snapshot of the life of a cell
-
Urine is a dynamic biofluid, which changes over hours and days within an individual; therefore, at present, no single approach for the isolation of urinary extracellular vesicles is likely to comprehensively distinguish between healthy and disease states
-
Alterations in the composition of urinary extracellular vesicles are useful experimentally and may provide information about disease pathophysiology as well as provide diagnostic end points for the study of renal disease
-
Perhaps the greatest promise of this 'extracellular organelle' is to open a window for science into a greater understanding of cellular therapeutics
Abstract
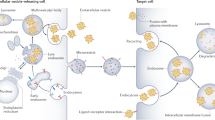
Urine is a valuable diagnostic medium and, with the discovery of urinary extracellular vesicles, is viewed as a dynamic bioactive fluid. Extracellular vesicles are lipid-enclosed structures that can be classified into three categories: exosomes, microvesicles (or ectosomes) and apoptotic bodies. This classification is based on the mechanisms by which membrane vesicles are formed: fusion of multivesicular bodies with the plasma membranes (exosomes), budding of vesicles directly from the plasma membrane (microvesicles) or those shed from dying cells (apoptotic bodies). During their formation, urinary extracellular vesicles incorporate various cell-specific components (proteins, lipids and nucleic acids) that can be transferred to target cells. The rigour needed for comparative studies has fueled the search for optimal approaches for their isolation, purification, and characterization. RNA, the newest extracellular vesicle component to be discovered, has received substantial attention as an extracellular vesicle therapeutic, and compelling evidence suggests that ex vivo manipulation of microRNA composition may have uses in the treatment of kidney disorders. The results of these studies are building the case that urinary extracellular vesicles act as mediators of renal pathophysiology. As the field of extracellular vesicle studies is burgeoning, this Review focuses on primary data obtained from studies of human urine rather than on data from studies of laboratory animals or cultured immortalized cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Wolf, P. The nature and significance of platelet products in human plasma. Br. J. Haematol. 13, 269–288 (1967).
Taylor, D. D. & Doellgast, G. J. Quantitation of peroxidase-antibody binding to membrane fragments using column chromatography. Anal. Biochem. 98, 53–59 (1979).
Ratajczak, J., Wysoczynski, M., Hayek, F., Janowska-Wieczorek, A. & Ratajczak, M. Z. Membrane-derived microvesicles: important and underappreciated mediators of cell-to-cell communication. Leukemia 20, 1487–1495 (2006).
Street, J. M. et al. Exosomal transmission of functional aquaporin 2 in kidney cortical collecting duct cells. J. Physiol. 589, 6119–6127 (2011).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Simpson, R. J., Kalra, H. & Mathivanan, S. ExoCarta as a resource for exosomal research. J. Extracell. Vesicles 1, 18374 (2012).
Mathivanan, S., Fahner, C. J., Reid, G. E. & Simpson, R. J. ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res. 40, D1241–D1244 (2012).
Kalra, H. et al. Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 10, e1001450 (2012).
Kim, D. K. et al. EVpedia: a community web portal for extracellular vesicles research. Bioinformatics 31, 933–939 (2015).
Kim, D. K. et al. EVpedia: an integrated database of high-throughput data for systemic analyses of extracellular vesicles. J. Extracell. Vesicles 2, 20384 (2013).
Dear, J. W., Street, J. M. & Bailey, M. A. Urinary exosomes: a reservoir for biomarker discovery and potential mediators of intrarenal signalling. Proteomics 13, 1572–1580 (2013).
Thery, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 9, 581–593 (2009).
Simpson, R. J. & Mathivanan, S. Extracellular microvesicles: the need for internationally recognised nomenclature and stringent purification criteria. J. Proteomics Bioinf. 5, 1–2 (2012).
Simpson, R. J., Jensen, S. S. & Lim, J. W. Proteomic profiling of exosomes: current perspectives. Proteomics 8, 4083–4099 (2008).
Lotvall, J. et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 3, 26913 (2014).
Witwer, K. W. et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2, 20360 (2013).
Gould, S. J. & Raposo, G. As we wait: coping with an imperfect nomenclature for extracellular vesicles. J. Extracell. Vesicles 2, 20389 (2013).
Thongboonkerd, V., McLeish, K. R., Arthur, J. M. & Klein, J. B. Proteomic analysis of normal human urinary proteins isolated by acetone precipitation or ultracentrifugation. Kidney Int. 62, 1461–1469 (2002).
Pisitkun, T., Shen, R. F. & Knepper, M. A. Identification and proteomic profiling of exosomes in human urine. Proc. Natl Acad. Sci. USA 101, 13368–13373 (2004).
Gonzales, P. A. et al. Large-scale proteomics and phosphoproteomics of urinary exosomes. J. Am. Soc. Nephrol. 20, 363–379 (2009).
Moon, P. G., You, S., Lee, J. E., Hwang, D. & Baek, M. C. Urinary exosomes and proteomics. Mass Spectrom. Rev. 30, 1185–1202 (2011).
Dear, J. W. Urinary exosomes join the fight against infection. J. Am. Soc. Nephrol. 25, 1889–1891 (2014).
Borges, F. T. et al. TGF-β1-containing exosomes from injured epithelial cells activate fibroblasts to initiate tissue regenerative responses and fibrosis. J. Am. Soc. Nephrol. 24, 385–392 (2013).
Raimondo, F., Morosi, L., Chinello, C., Magni, F. & Pitto, M. Advances in membranous vesicle and exosome proteomics improving biological understanding and biomarker discovery. Proteomics 11, 709–720 (2011).
Pan, B. T., Teng, K., Wu, C., Adam, M. & Johnstone, R. M. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. J. Cell Biol. 101, 942–948 (1985).
Raposo, G. et al. B lymphocytes secrete antigen-presenting vesicles. J. Exp. Med. 183, 1161–1172 (1996).
Thery, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
van Niel, G. et al. Intestinal epithelial cells secrete exosome-like vesicles. Gastroenterology 121, 337–349 (2001).
Thery, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 30, 3.22.1–3.22.29 (2006).
Fevrier, B. & Raposo, G. Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr. Opin. Cell Biol. 16, 415–421 (2004).
Mathivanan, S., Ji, H. & Simpson, R. J. Exosomes: extracellular organelles important in intercellular communication. J. Proteomics 73, 1907–1920 (2010).
Williams, R. L. & Urbe, S. The emerging shape of the ESCRT machinery. Nat. Rev. Mol. Cell Biol. 8, 355–368 (2007).
Hurley, J. H. ESCRT complexes and the biogenesis of multivesicular bodies. Curr. Opin. Cell Biol. 20, 4–11 (2008).
Raiborg, C. & Stenmark, H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 458, 445–452 (2009).
Keller, S., Sanderson, M. P., Stoeck, A. & Altevogt, P. Exosomes: from biogenesis and secretion to biological function. Immunol. Lett. 107, 102–108 (2006).
Laulagnier, K. et al. Mast cell- and dendritic cell-derived exosomes display a specific lipid composition and an unusual membrane organization. Biochem. J. 380, 161–171 (2004).
Skotland, T., Sandvig, K. & Llorente, A. Lipids in exosomes: current knowledge and the way forward. Prog. Lipid Res. 66, 30–41 (2017).
Record, M., Carayon, K., Poirot, M. & Silvente-Poirot, S. Exosomes as new vesicular lipid transporters involved in cell-cell communication and various pathophysiologies. Biochim. Biophys. Acta 1841, 108–120 (2014).
Record, M., Poirot, M. & Silvente-Poirot, S. Emerging concepts on the role of exosomes in lipid metabolic diseases. Biochimie 96, 67–74 (2014).
Bard, M. P. et al. Proteomic analysis of exosomes isolated from human malignant pleural effusions. Am. J. Respir. Cell Mol. Biol. 31, 114–121 (2004).
Escola, J. M. et al. Selective enrichment of tetraspan proteins on the internal vesicles of multivesicular endosomes and on exosomes secreted by human B-lymphocytes. J. Biol. Chem. 273, 20121–20127 (1998).
Chaput, N. et al. The potential of exosomes in immunotherapy of cancer. Blood Cells Mol. Dis. 35, 111–115 (2005).
Thery, C. et al. Proteomic analysis of dendritic cell-derived exosomes: a secreted subcellular compartment distinct from apoptotic vesicles. J. Immunol. 166, 7309–7318 (2001).
Takahashi, A. et al. Exosomes maintain cellular homeostasis by excreting harmful DNA from cells. Nat. Commun. 8, 15287 (2017).
Heijnen, H. F., Schiel, A. E., Fijnheer, R., Geuze, H. J. & Sixma, J. J. Activated platelets release two types of membrane vesicles: microvesicles by surface shedding and exosomes derived from exocytosis of multivesicular bodies and α-granules. Blood 94, 3791–3799 (1999).
Hugel, B., Martinez, M. C., Kunzelmann, C. & Freyssinet, J. M. Membrane microparticles: two sides of the coin. Physiology (Bethesda) 20, 22–27 (2005).
McConnell, R. E. et al. The enterocyte microvillus is a vesicle-generating organelle. J. Cell Biol. 185, 1285–1298 (2009).
Kowal, J. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl Acad. Sci. USA 113, E968–E977 (2016).
van der Pol, E., Boing, A. N., Harrison, P., Sturk, A. & Nieuwland, R. Classification, functions, and clinical relevance of extracellular vesicles. Pharmacol. Rev. 64, 676–705 (2012).
Barry, O. P. & FitzGerald, G. A. Mechanisms of cellular activation by platelet microparticles. Thromb. Haemost. 82, 794–800 (1999).
Redman, C. W. & Sargent, I. L. Microparticles and immunomodulation in pregnancy and pre-eclampsia. J. Reprod. Immunol. 76, 61–67 (2007).
Miyazaki, Y. et al. High shear stress can initiate both platelet aggregation and shedding of procoagulant containing microparticles. Blood 88, 3456–3464 (1996).
Horstman, L. L., Jy, W., Jimenez, J. J., Bidot, C. & Ahn, Y. S. New horizons in the analysis of circulating cell-derived microparticles. Keio J. Med. 53, 210–230 (2004).
Cocucci, E., Racchetti, G. & Meldolesi, J. Shedding microvesicles: artefacts no more. Trends Cell Biol. 19, 43–51 (2009).
Pilzer, D., Gasser, O., Moskovich, O., Schifferli, J. A. & Fishelson, Z. Emission of membrane vesicles: roles in complement resistance, immunity and cancer. Springer Semin. Immunopathol. 27, 375–387 (2005).
Dean, W. L., Lee, M. J., Cummins, T. D., Schultz, D. J. & Powell, D. W. Proteomic and functional characterisation of platelet microparticle size classes. Thromb. Haemost. 102, 711–718 (2009).
Hunter, M. P. et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS ONE. 3, e3694 (2008).
D'Souza-Schorey, C. & Chavrier, P. ARF proteins: roles in membrane traffic and beyond. Nat. Rev. Mol. Cell Biol. 7, 347–358 (2006).
Donaldson, J. G. Multiple roles for Arf6: sorting, structuring, and signaling at the plasma membrane. J. Biol. Chem. 278, 41573–41576 (2003).
Beyer, C. & Pisetsky, D. S. The role of microparticles in the pathogenesis of rheumatic diseases. Nat. Rev. Rheumatol. 6, 21–29 (2010).
Elmore, S. Apoptosis: a review of programmed cell death. Toxicol. Pathol. 35, 495–516 (2007).
Turturici, G., Tinnirello, R., Sconzo, G. & Geraci, F. Extracellular membrane vesicles as a mechanism of cell-to-cell communication: advantages and disadvantages. Am. J. Physiol. Cell Physiol. 306, C621–C633 (2014).
Hristov, M., Erl, W., Linder, S. & Weber, P. C. Apoptotic bodies from endothelial cells enhance the number and initiate the differentiation of human endothelial progenitor cells in vitro. Blood 104, 2761–2766 (2004).
Turiak, L. et al. Proteomic characterization of thymocyte-derived microvesicles and apoptotic bodies in BALB/c mice. J. Proteomics 74, 2025–2033 (2011).
Poon, I. K., Lucas, C. D., Rossi, A. G. & Ravichandran, K. S. Apoptotic cell clearance: basic biology and therapeutic potential. Nat. Rev. Immunol. 14, 166–180 (2014).
Bobrie, A., Colombo, M., Raposo, G. & Thery, C. Exosome secretion: molecular mechanisms and roles in immune responses. Traffic 12, 1659–1668 (2011).
Choi, D. S., Kim, D. K., Kim, Y. K. & Gho, Y. S. Proteomics of extracellular vesicles: exosomes and ectosomes. Mass Spectrom. Rev. 34, 474–490 (2014).
Simons, M. & Raposo, G. Exosomes — vesicular carriers for intercellular communication. Curr. Opin. Cell Biol. 21, 575–581 (2009).
Musante, L., Saraswat, M., Ravida, A., Byrne, B. & Holthofer, H. Recovery of urinary nanovesicles from ultracentrifugation supernatants. Nephrol. Dial. Transplant. 28, 1425–1433 (2013).
Alvarez, M. L., Khosroheidari, M., Kanchi Ravi, R. & DiStefano, J. K. Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int. 82, 1024–1032 (2012).
Gonzales, P. A. et al. Isolation and purification of exosomes in urine. Methods Mol. Biol. 641, 89–99 (2010).
Merchant, M. L. et al. Microfiltration isolation of human urinary exosomes for characterization by MS. Proteomics Clin. Appl. 4, 84–96 (2010).
Rood, I. M. et al. Comparison of three methods for isolation of urinary microvesicles to identify biomarkers of nephrotic syndrome. Kidney Int. 78, 810–816 (2010).
Hiemstra, T. F. et al. Human urinary exosomes as innate immune effectors. J. Am. Soc. Nephrol. 25, 2017–2027 (2014).
Chen, C. Y., Hogan, M. C. & Ward, C. J. Purification of exosome-like vesicles from urine. Methods Enzymol. 524, 225–241 (2013).
Hogan, M. C. et al. Characterization of PKD protein-positive exosome-like vesicles. J. Am. Soc. Nephrol. 20, 278–288 (2009).
Hogan, M. C. et al. Subfractionation, characterization, and in-depth proteomic analysis of glomerular membrane vesicles in human urine. Kidney Int. 85, 1225–1237 (2013).
Miranda, K. C. et al. Nucleic acids within urinary exosomes/microvesicles are potential biomarkers for renal disease. Kidney Int. 78, 191–199 (2010).
Prunotto, M. et al. Proteomic analysis of podocyte exosome-enriched fraction from normal human urine. J. Proteomics 82, 193–229 (2013).
Hildonen, S., Skarpen, E., Halvorsen, T. G. & Reubsaet, L. Isolation and mass spectrometry analysis of urinary extraexosomal proteins. Sci. Rep. 6, 36331 (2016).
Salih, M. et al. Proteomics of urinary vesicles links plakins and complement to polycystic kidney disease. J. Am. Soc. Nephrol. 27, 3079–3092 (2016).
Salih, M., Fenton, R. A., Zietse, R. & Hoorn, E. J. Urinary extracellular vesicles as markers to assess kidney sodium transport. Curr. Opin. Nephrol. Hypertens. 25, 67–72 (2016).
Cheruvanky, A. et al. Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am. J. Physiol. Renal Physiol. 292, F1657–F1661 (2007).
Zubiri, I. et al. Diabetic nephropathy induces changes in the proteome of human urinary exosomes as revealed by label-free comparative analysis. J. Proteomics 96, 92–102 (2014).
Channavajjhala, S. K. et al. Optimizing the purification and analysis of miRNAs from urinary exosomes. Clin. Chem. Lab. Med. 52, 345–354 (2014).
Rider, M. A., Hurwitz, S. N. & Meckes, D. G. Jr. ExtraPEG: a polyethylene glycol-based method for enrichment of extracellular vesicles. Sci. Rep. 6, 23978 (2016).
Wang, D. & Sun, W. Urinary extracellular microvesicles: isolation methods and prospects for urinary proteome. Proteomics 14, 1922–1932 (2014).
Salih, M., Zietse, R. & Hoorn, E. J. Urinary extracellular vesicles and the kidney: biomarkers and beyond. Am. J. Physiol. Renal Physiol. 306, F1251–F1259 (2014).
Gardiner, C. et al. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J. Extracell. Vesicles 5, 32945 (2016).
Bobrie, A., Colombo, M., Krumeich, S., Raposo, G. & Thery, C. Diverse subpopulations of vesicles secreted by different intracellular mechanisms are present in exosome preparations obtained by differential ultracentrifugation. J. Extracell. Vesicles 1, 18397 (2012).
Jacquillet, G., Hoorn, E. J., Vilasi, A. & Unwin, R. J. Urinary vesicles: in splendid isolation. Nephrol. Dial. Transplant. 28, 1332–1335 (2013).
Lv, L. L. et al. Isolation and quantification of microRNAs from urinary exosomes/microvesicles for biomarker discovery. Int. J. Biol. Sci. 9, 1021–1031 (2013).
Fernandez-Llama, P. et al. Tamm-Horsfall protein and urinary exosome isolation. Kidney Int. 77, 736–742 (2010).
Musante, L. et al. Biochemical and physical characterisation of urinary nanovesicles following CHAPS treatment. PLoS ONE 7, e37279 (2012).
Cheng, L., Sun, X., Scicluna, B. J., Coleman, B. M. & Hill, A. F. Characterization and deep sequencing analysis of exosomal and non-exosomal miRNA in human urine. Kidney Int. 86, 433–444 (2014).
Gonzales, P., Pisitkun, T. & Knepper, M. A. Urinary exosomes: is there a future? Nephrol. Dial. Transplant. 23, 1799–1801 (2008).
Woo, H. K. et al. Exodisc for rapid, size-selective, and efficient isolation and analysis of nanoscale extracellular vesicles from biological samples. ACS Nano 11, 1360–1370 (2017).
Liang, L. G. et al. An integrated double-filtration microfluidic device for isolation, enrichment and quantification of urinary extracellular vesicles for detection of bladder cancer. Sci. Rep. 7, 46224 (2017).
Taylor, D. D., Zacharias, W. & Gercel-Taylor, C. Exosome isolation for proteomic analyses and RNA profiling. Methods Mol. Biol. 728, 235–246 (2011).
Albertsson, P. A. & Frick, G. Partition of virus particles in a liquid two-phase system. Biochim. Biophys. Acta 37, 230–237 (1960).
van der Pol, E. et al. Particle size distribution of exosomes and microvesicles determined by transmission electron microscopy, flow cytometry, nanoparticle tracking analysis, and resistive pulse sensing. J. Thromb. Haemost. 12, 1182–1192 (2014).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Rupert, D. L., Claudio, V., Lasser, C. & Bally, M. Methods for the physical characterization and quantification of extracellular vesicles in biological samples. Biochim. Biophys. Acta 1861, 3164–3179 (2017).
Szatanek, R. et al. The methods of choice for extracellular vesicles (EVs) characterization. Int. J. Mol. Sci. 18, E1153 (2017).
Erdbrugger, U. & Lannigan, J. Analytical challenges of extracellular vesicle detection: a comparison of different techniques. Cytometry A 89, 123–134 (2016).
Coumans, F. A. W. et al. Methodological guidelines to study extracellular vesicles. Circ. Res. 120, 1632–1648 (2017).
Daaboul, G. G. et al. Digital detection of exosomes by interferometric imaging. Sci. Rep. 6, 37246 (2016).
Faez, S. et al. Fast, label-free tracking of single viruses and weakly scattering nanoparticles in a nanofluidic optical fiber. ACS Nano 9, 12349–12357 (2015).
McLeish, K. R., Merchant, M. L., Klein, J. B. & Ward, R. A. Technical note: proteomic approaches to fundamental questions about neutrophil biology. J. Leukoc. Biol. 94, 683–692 (2013).
Erdbrugger, U. & Le, T. H. Extracellular vesicles in renal diseases: more than novel biomarkers? J. Am. Soc. Nephrol. 27, 12–26 (2016).
Zhang, W. et al. Extracellular vesicles in diagnosis and therapy of kidney diseases. Am. J. Physiol. Renal Physiol. 311, F844–F851 (2016).
Smalley, D. M., Sheman, N. E., Nelson, K. & Theodorescu, D. Isolation and identification of potential urinary microparticle biomarkers of bladder cancer. J. Proteome Res. 7, 2088–2096 (2008).
Stamer, W. D., Hoffman, E. A., Luther, J. M., Hachey, D. L. & Schey, K. L. Protein profile of exosomes from trabecular meshwork cells. J. Proteomics 74, 796–804 (2011).
Moon, P. G. et al. Proteomic analysis of urinary exosomes from patients of early IgA nephropathy and thin basement membrane nephropathy. Proteomics 11, 2459–2475 (2011).
Raj, D. A., Fiume, I., Capasso, G. & Pocsfalvi, G. A multiplex quantitative proteomics strategy for protein biomarker studies in urinary exosomes. Kidney Int. 81, 1263–1272 (2012).
Principe, S. et al. In-depth proteomic analyses of exosomes isolated from expressed prostatic secretions in urine. Proteomics 13, 1667–1671 (2013).
Fraser, K. B. et al. LRRK2 secretion in exosomes is regulated by 14-3-3. Hum. Mol. Genet. 22, 4988–5000 (2013).
Hogan, M. C. et al. Identification of biomarkers for PKD1 using urinary exosomes. J. Am. Soc. Nephrol. 26, 1661–1670 (2015).
Bourderioux, M. et al. A new workflow for proteomic analysis of urinary exosomes and assessment in cystinuria patients. J. Proteome Res. 14, 567–577 (2015).
Rood, I. M. et al. Increased expression of lysosome membrane protein 2 in glomeruli of patients with idiopathic membranous nephropathy. Proteomics 15, 3722–3730 (2015).
Zhou, H. et al. Collection, storage, preservation, and normalization of human urinary exosomes for biomarker discovery. Kidney Int. 69, 1471–1476 (2006).
Zhou, H. et al. Exosomal fetuin-A identified by proteomics: a novel urinary biomarker for detecting acute kidney injury. Kidney Int. 70, 1847–1857 (2006).
Zhou, H. et al. Urinary exosomal transcription factors, a new class of biomarkers for renal disease. Kidney Int. 74, 613–621 (2008).
Wang, Z., Hill, S., Luther, J. M., Hachey, D. L. & Schey, K. L. Proteomic analysis of urine exosomes by multidimensional protein identification technology (MudPIT). Proteomics 12, 329–338 (2012).
Gerlach, J. Q. et al. Surface glycosylation profiles of urine extracellular vesicles. PLoS ONE 8, e74801 (2013).
Corbetta, S. et al. Urinary exosomes in the diagnosis of Gitelman and Bartter syndromes. Nephrol. Dial. Transplant. 30, 621–630 (2015).
Esteva-Font, C. et al. Renal sodium transporters are increased in urinary exosomes of cyclosporine-treated kidney transplant patients. Am. J. Nephrol. 39, 528–535 (2014).
Saraswat, M. et al. N-linked (N−) glycoproteomics of urinary exosomes. [Corrected]. Mol. Cell. Proteomics 14, 263–276 (2015).
Kosanovic, M. & Jankovic, M. Isolation of urinary extracellular vesicles from Tamm- Horsfall protein-depleted urine and their application in the development of a lectin-exosome-binding assay. Biotechniques 57, 143–149 (2014).
Turco, A. E. et al. Specific renal parenchymal-derived urinary extracellular vesicles identify age-associated structural changes in living donor kidneys. J. Extracell. Vesicles 5, 29642 (2016).
Panich, T. et al. Urinary exosomal activating transcriptional factor 3 as the early diagnostic biomarker for sepsis-induced acute kidney injury. BMC Nephrol. 18, 10 (2017).
Wolley, M. J. et al. In primary aldosteronism, mineralocorticoids influence exosomal sodium-chloride cotransporter abundance. J. Am. Soc. Nephrol. 28, 56–63 (2017).
Lytvyn, Y. et al. Assessment of urinary microparticles in normotensive patients with type 1 diabetes. Diabetologia 60, 581–584 (2017).
Hogan, M. C. et al. Strategy and rationale for urine collection protocols employed in the NEPTUNE study. BMC Nephrol. 16, 190 (2015).
Thomas, C. E., Sexton, W., Benson, K., Sutphen, R. & Koomen, J. Urine collection and processing for protein biomarker discovery and quantification. Cancer Epidemiol. Biomarkers Prev. 19, 953–959 (2010).
Rodby, R. A. Timed urine collections for albumin and protein: “the king is dead, long live the king!”. Am. J. Kidney Dis. 68, 836–838 (2016).
Musante, L., Tataruch, D. E. & Holthofer, H. Use and isolation of urinary exosomes as biomarkers for diabetic nephropathy. Front. Endocrinol. (Lausanne) 5, 149 (2014).
Sir Elkhatim, R., Li, J. Y., Yong, T. Y. & Gleadle, J. M. Dipping your feet in the water: podocytes in urine. Expert Rev. Mol. Diagn. 14, 423–437 (2014).
Fabian, M. R., Sonenberg, N. & Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 79, 351–379 (2010).
Trionfini, P., Benigni, A. & Remuzzi, G. MicroRNAs in kidney physiology and disease. Nat. Rev. Nephrol. 11, 23–33 (2015).
Schageman, J. et al. The complete exosome workflow solution: from isolation to characterization of RNA cargo. Biomed Res. Int. 2013, 253957 (2013).
Cantaluppi, V. et al. Microvesicles derived from endothelial progenitor cells protect the kidney from ischemia-reperfusion injury by microRNA-dependent reprogramming of resident renal cells. Kidney Int. 82, 412–427 (2012).
Gildea, J. J. et al. Exosomal transfer from human renal proximal tubule cells to distal tubule and collecting duct cells. Clin. Biochem. 47, 89–94 (2014).
Murakami, T. et al. Development of glomerulus-, tubule-, and collecting duct-specific mRNA assay in human urinary exosomes and microvesicles. PLoS ONE 9, e109074 (2014).
Chen, H. H. et al. Exosomal ATF3 RNA attenuates pro-inflammatory gene MCP-1 transcription in renal ischemia-reperfusion. J. Cell. Physiol. 229, 1202–1211 (2014).
Lv, L. L. et al. CD2AP mRNA in urinary exosome as biomarker of kidney disease. Clin. Chim. Acta 428, 26–31 (2014).
Lv, L. L. et al. MicroRNA-29c in urinary exosome/microvesicle as a biomarker of renal fibrosis. Am. J. Physiol. Renal Physiol. 305, F1220–F1227 (2013).
Ichii, O. et al. Decreased miR-26a expression correlates with the progression of podocyte injury in autoimmune glomerulonephritis. PLoS ONE 9, e110383 (2014).
Barutta, F. et al. Urinary exosomal microRNAs in incipient diabetic nephropathy. PLoS ONE 8, e73798 (2013).
Eissa, S., Matboli, M., Aboushahba, R., Bekhet, M. M. & Soliman, Y. Urinary exosomal microRNA panel unravels novel biomarkers for diagnosis of type 2 diabetic kidney disease. J. Diabetes Complications 30, 1585–1592 (2016).
Jia, Y. et al. miRNAs in urine extracellular vesicles as predictors of early-stage diabetic nephropathy. J. Diabetes Res. 2016, 7932765 (2016).
Rekker, K. et al. Comparison of serum exosome isolation methods for microRNA profiling. Clin. Biochem. 47, 135–138 (2014).
Nicolson, G. L. The fluid-mosaic model of membrane structure: still relevant to understanding the structure, function and dynamics of biological membranes after more than 40 years. Biochim. Biophys. Acta 1838, 1451–1466 (2014).
Kreimer, S. et al. Mass-spectrometry-based molecular characterization of extracellular vesicles: lipidomics and proteomics. J. Proteome Res. 14, 2367–2384 (2015).
Gyorgy, B. et al. Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell. Mol. Life Sci. 68, 2667–2688 (2011).
Trajkovic, K. et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319, 1244–1247 (2008).
Llorente, A. et al. Molecular lipidomics of exosomes released by PC-3 prostate cancer cells. Biochim. Biophys. Acta 1831, 1302–1309 (2013).
Subra, C., Laulagnier, K., Perret, B. & Record, M. Exosome lipidomics unravels lipid sorting at the level of multivesicular bodies. Biochimie 89, 205–212 (2007).
Muralidharan-Chari, V. et al. ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr. Biol. 19, 1875–1885 (2009).
Agmon, E. & Stockwell, B. R. Lipid homeostasis and regulated cell death. Curr. Opin. Chem. Biol. 39, 83–89 (2017).
Del Boccio, P. et al. A hyphenated microLC-Q-TOF-MS platform for exosomal lipidomics investigations: application to RCC urinary exosomes. Electrophoresis 33, 689–696 (2012).
Ichimura, T. et al. Kidney injury molecule–1 is a phosphatidylserine receptor that confers a phagocytic phenotype on epithelial cells. J. Clin. Invest. 118, 1657–1668 (2008).
Zhao, N. et al. MicroRNA miR145 regulates TGFBR2 expression and matrix synthesis in vascular smooth muscle cells. Circ. Res. 116, 23–34 (2015).
Kelly, K. J. et al. Improved structure and function in autosomal recessive polycystic rat kidneys with renal tubular cell therapy. PLoS ONE 10, e0131677 (2015).
Collino, F. et al. AKI recovery induced by mesenchymal stromal cell-derived extracellular vesicles carrying microRNAs. J. Am. Soc. Nephrol. 26, 2349–2360 (2015).
Bruno, S. et al. Microvesicles derived from mesenchymal stem cells enhance survival in a lethal model of acute kidney injury. PLoS ONE 7, e33115 (2012).
Bruno, S. et al. Mesenchymal stem cell-derived microvesicles protect against acute tubular injury. J. Am. Soc. Nephrol. 20, 1053–1067 (2009).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02138331 (2014).
Gracia, T. et al. Urinary exosomes contain microRNAs capable of paracrine modulation of tubular transporters in kidney. Sci. Rep. 7, 40601 (2017).
Burger, D. et al. Urinary podocyte microparticles identify prealbuminuric diabetic glomerular injury. J. Am. Soc. Nephrol. 25, 1401–1407 (2014).
Colombo, M., Raposo, G. & Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 30, 255–289 (2014).
Khurana, R. et al. Identification of urinary exosomal noncoding RNAs as novel biomarkers in chronic kidney disease. RNA 23, 142–152 (2017).
Acknowledgements
J.B.K. and M.L.M. are supported in part by grants R01DK110077 (J.B.K.), R01DK091584 (M.L.M.), P50AA024337 (M.L.M.) and P20GM113226 (MLM) from the US NIH. I.M.R. is supported by an AGIKO grant from ZonMw-NWO (grant 92003587).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article. M.L.M, I.M.R, J.K.J.D. and J.B.K made substantial contributions to discussions of the content. M.L.M, I.M.R and J.K.J.D. wrote the article and M.L.M., I.M.R., J.K.J.D. and J.B.K. reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
DATABASES
FURTHER INFORMATION
Glossary
- mRNAs
-
Messenger RNAs, which are transcripts of DNA.
- MicroRNAs
-
(miRNAs). Small, non-coding RNAs that regulate gene expression post-transcriptionally by targeting specific mRNAs for inhibition or degradation through complementary base pairing.
- Exosomes
-
Extracellular vesicles that are formed by inward budding of the cell membrane, followed by fusion with a multivesicular body (MVB) and formation of intraluminal vesicles inside the MVB. The intraluminal vesicles that are released by fusion of the MVB with the cell plasma membrane are called exosomes.
- Microvesicles
-
Extracellular vesicles that are formed by direct budding from the cell plasma membrane.
- Apoptotic bodies
-
Extracellular vesicle that are released during the late stages of cell death.
- Transcriptomics
-
The study of the complete set of RNA transcripts (the transcriptome) that is encoded by the genome, underspecific circumstances or in a specific cell.
- Proteomics
-
Large scale studies of proteins involving the systematic identification and quantification of the complete set of proteins (the proteome) of a biological system (cell, tissue, organ, biological fluid or organism) at a specific point in time. Mass spectrometry is the technique most often used for proteomic analysis.
- Nephrotic syndrome
-
A constellation of symptoms characterized by heavy proteinuria (>3–3.5 g/24 h), hypoalbuminuria (<30 g/l), peripheral oedema and hyperlipidaemia.
Rights and permissions
About this article
Cite this article
Merchant, M., Rood, I., Deegens, J. et al. Isolation and characterization of urinary extracellular vesicles: implications for biomarker discovery. Nat Rev Nephrol 13, 731–749 (2017). https://doi.org/10.1038/nrneph.2017.148
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneph.2017.148
This article is cited by
-
Extracellular vesicle therapy for obesity-induced NAFLD: a comprehensive review of current evidence
Cell Communication and Signaling (2024)
-
Identification of novel snoRNA-based biomarkers for clear cell renal cell carcinoma from urine-derived extracellular vesicles
Biology Direct (2024)
-
Exosomes derived from programmed cell death: mechanism and biological significance
Cell Communication and Signaling (2024)
-
Ferroptosis: a promising candidate for exosome-mediated regulation in different diseases
Cell Communication and Signaling (2024)
-
Unlocking the Potential of Extracellular Vesicles as the Next Generation Therapy: Challenges and Opportunities
Tissue Engineering and Regenerative Medicine (2024)