Key Points
-
The human genome encodes only a small number of digestive glycoside hydrolases for the breakdown of sucrose, lactose and starch. Instead, the large diversity of complex polysaccharides in our diet is mainly digested by specialized enzymes encoded by the gut microbiome.
-
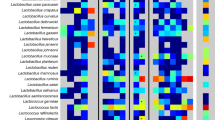
A model human microbiome was constructed from 177 microbial genomes in proportions that approximate their representation in the healthy adult gut, and this mini-microbiome was used to evaluate the diversity of carbohydrate-active enzymes (CAZymes) in the gut microbiota.
-
Gut bacteria from the phylum Bacteroidetes encode more CAZymes, and encode CAZymes from more families, than the other phyla represented in the model mini-microbiome. The large substrate range of these CAZymes is compatible with the diversity of the dietary plant cell wall polysaccharides that are presented to members of the microbiota.
Abstract
Descriptions of the microbial communities that live on and in the human body have progressed at a spectacular rate over the past 5 years, fuelled primarily by highly parallel DNA-sequencing technologies and associated advances in bioinformatics, and by the expectation that understanding how to manipulate the structure and functions of our microbiota will allow us to affect health and prevent or treat diseases. Among the myriad of genes that have been identified in the human gut microbiome, those that encode carbohydrate-active enzymes (CAZymes) are of particular interest, as these enzymes are required to digest most of our complex repertoire of dietary polysaccharides. In this Analysis article, we examine the carbohydrate-digestive capacity of a simplified but representative mini-microbiome in order to highlight the abundance and variety of bacterial CAZymes that are represented in the human gut microbiota.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hooper, L. V., Littman, D. R. & Macpherson, A. J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nature Commun. 3, 1245 (2012).
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009). Metagenomic and 16S rRNA sequencing suggest the existence of a functional core microbiome and that obesity is associated with reduced bacterial diversity and altered metabolic pathways.
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012). This huge metagenomic sequencing effort on the gut microbiota of 124 individuals has revealed a set of 3.3 million non-redundant genes. If 1.5–2% of these genes encode GHs and PLs (our model mini-microbiome has 1.75%), this would amount to some 50,000–60,000 GHs and PLs.
Ley, R. E. et al. Obesity alters gut microbial ecology. Proc. Natl Acad. Sci. USA 102, 11070–11075 (2005).
Paliy, O., Kenche, H., Abernathy, F. & Michail, S. High-throughput quantitative analysis of the human intestinal microbiota with a phylogenetic microarray. Appl. Environ. Microbiol. 75, 3572–3579 (2009).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Lagier, J. C., Million, M., Hugon, P., Armougom, F. & Raoult, D. Human gut microbiota: repertoire and variations. Front. Cell. Infect. Microbiol. 2, 136 (2012).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 8, e1002358 (2012).
Grabitske, H. A. & Slavin, J. L. Low-digestible carbohydrates in practice. J. Am. Diet. Assoc. 108, 1677–1681 (2008).
Topping, D. L. & Clifton, P. M. Short-chain fatty acids and human colonic function: roles of resistant starch and nonstarch polysaccharides. Physiol. Rev. 81, 1031–1064 (2001).
Lattimer, J. M. & Haub, M. D. Effects of dietary fiber and its components on metabolic health. Nutrients 2, 1266–1289 (2010).
Martens, E. C. et al. Recognition and degradation of plant cell wall polysaccharides by two human gut symbionts. PLoS Biol. 9, e1001221 (2011). This study shows that two species of gut bacteria, B. thetaiotaomicron and Bacteroides ovatus , are capable of utilizing nearly all the major plant and host complex glycans, including type II rhamnogalacturonan.
Cantarel, B. L., Lombard, V. & Henrissat, B. Complex carbohydrate utilization by the healthy human microbiome. PLoS ONE 7, e28742 (2012). This analysis of 520 metagenomic samples from five major body sites shows that even when the microbial community composition varies, the CAZyme profiles are very similar within a body site, suggesting that the observed functional profile and microbiota have adapted to the local carbohydrate composition.
Bernalier-Donadille, A. [Fermentative metabolism by the human gut microbiota]. Gastroenterol. Clin. Biol. 34 (Suppl. 1), S16–S22 (2010) (in French).
McNeil, N. I. The contribution of the large intestine to energy supplies in man. Am. J. Clin. Nutr. 39, 338–342 (1984).
Hijova, E. & Chmelarova, A. Short chain fatty acids and colonic health. Bratisl. Lek. Listy 108, 354–358 (2007).
Floch, M. H. & Hong-Curtiss, J. Probiotics and functional foods in gastrointestinal disorders. Curr. Treat. Opt. Gastroenterol. 5, 311–321 (2002).
Hamer, H. M. et al. Review article: the role of butyrate on colonic function. Aliment. Pharmacol. Ther. 27, 104–119 (2008).
Henrissat, B. A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 280, 309–316 (1991).
Koropatkin, N. M., Cameron, E. A. & Martens, E. C. How glycan metabolism shapes the human gut microbiota. Nature Rev. Microbiol. 10, 323–335 (2012).
Lombard, V. et al. A hierarchical classification of polysaccharide lyases for glycogenomics. Biochem. J. 432, 437–444 (2010).
Gloster, T. M., Turkenburg, J. P., Potts, J. R., Henrissat, B. & Davies, G. J. Divergence of catalytic mechanism within a glycosidase family provides insight into evolution of carbohydrate metabolism by human gut flora. Chem. Biol. 15, 1058–1067 (2008).
Davies, G. & Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 3, 853–859 (1995).
Henrissat, B. & Bairoch, A. Updating the sequence-based classification of glycosyl hydrolases. Biochem. J. 316, 695–696 (1996).
Henrissat, B. & Bairoch, A. New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 293, 781–788 (1993).
Henrissat, B. & Davies, G. Structural and sequence-based classification of glycoside hydrolases. Curr. Opin. Struct. Biol. 7, 637–644 (1997).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2009).
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008).
Bjursell, M. K., Martens, E. C. & Gordon, J. I. Functional genomic and metabolic studies of the adaptations of a prominent adult human gut symbiont, Bacteroides thetaiotaomicron, to the suckling period. J. Biol. Chem. 281, 36269–36279 (2006).
Mirande, C. et al. Dietary fibre degradation and fermentation by two xylanolytic bacteria Bacteroides xylanisolvens XB1A and Roseburia intestinalis XB6B4 from the human intestine. J. Appl. Microbiol. 109, 451–460 (2010).
Flint, H. J., Bayer, E. A., Rincon, M. T., Lamed, R. & White, B. A. Polysaccharide utilization by gut bacteria: potential for new insights from genomic analysis. Nature Rev. Microbiol. 6, 121–131 (2008).
Robert, C., Del'Homme, C. & Bernalier-Donadille, A. Interspecies H2 transfer in cellulose degradation between fibrolytic bacteria and H2-utilizing microorganisms from the human colon. FEMS Microbiol. Lett. 205, 209–214 (2001).
Sonnenburg, J. L. et al. Glycan foraging in vivo by an intestine-adapted bacterial symbiont. Science 307, 1955–1959 (2005).
Markowitz, V. M. et al. The integrated microbial genomes system: an expanding comparative analysis resource. Nucleic Acids Res. 38, D382–D390 (2010).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Kuriki, T. & Imanaka, T. The concept of the α-amylase family: structural similarity and common catalytic mechanism. J. Biosci. Bioeng. 87, 557–565 (1999).
Stam, M. R., Danchin, E. G., Rancurel, C., Coutinho, P. M. & Henrissat, B. Dividing the large glycoside hydrolase family 13 into subfamilies: towards improved functional annotations of α-amylase-related proteins. Protein Eng. Des. Sel. 19, 555–562 (2006).
Leitch, E. C., Walker, A. W., Duncan, S. H., Holtrop, G. & Flint, H. J. Selective colonization of insoluble substrates by human faecal bacteria. Environ. Microbiol. 9, 667–679 (2007).
Vollmer, W., Joris, B., Charlier, P. & Foster, S. Bacterial peptidoglycan (murein) hydrolases. FEMS Microbiol. Rev. 32, 259–286 (2008).
Hehemann, J. H. et al. Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature 464, 908–912 (2010).
Kitahara, M., Sakamoto, M., Ike, M., Sakata, S. & Benno, Y. Bacteroides plebeius sp. nov. and Bacteroides coprocola sp. nov., isolated from human faeces. Int. J. Syst. Evol. Microbiol. 55, 2143–2147 (2005).
Hehemann, J. H., Kelly, A. G., Pudlo, N. A., Martens, E. C. & Boraston, A. B. Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes. Proc. Natl Acad. Sci. USA 109, 19786–19791 (2012).
Zoetendal, E. G., Plugge, C. M., Akkermans, A. D. & de Vos, W. M. Victivallis vadensis gen. nov., sp. nov., a sugar-fermenting anaerobe from human faeces. Int. J. Syst. Evol. Microbiol. 53, 211–215 (2003).
Ze, X., Duncan, S. H., Louis, P. & Flint, H. J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 6, 1535–1543 (2012).
Acknowledgements
A.E.K. was funded by La Fondation Infectiopôle Sud, France.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (table)
The genomes composing the mini-microbiome (PDF 304 kb)
Supplementary information S2 (table)
Abundance of each family of glycoside hydrolases (GH) and polysaccharide lyases (PL) in the bacterial strains composing the mini-microbiome (XLSX 85 kb)
Rights and permissions
About this article
Cite this article
Kaoutari, A., Armougom, F., Gordon, J. et al. The abundance and variety of carbohydrate-active enzymes in the human gut microbiota. Nat Rev Microbiol 11, 497–504 (2013). https://doi.org/10.1038/nrmicro3050
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3050
This article is cited by
-
Relationships between diet and gut microbiome in an Italian and Dutch cohort: does the dietary protein to fiber ratio play a role?
European Journal of Nutrition (2024)
-
What if gastrointestinal complications in endurance athletes were gut injuries in response to a high consumption of ultra-processed foods? Please take care of your bugs if you want to improve endurance performance: a narrative review
European Journal of Applied Physiology (2024)
-
Comparative analysis of intestinal microbiota composition and transcriptome in diploid and triploid Carassius auratus
BMC Microbiology (2023)
-
Strain-specific alterations in gut microbiome and host immune responses elicited by tolerogenic Bifidobacterium pseudolongum
Scientific Reports (2023)
-
Identifying glycan consumers in human gut microbiota samples using metabolic labeling coupled with fluorescence-activated cell sorting
Nature Communications (2023)