Key Points
-
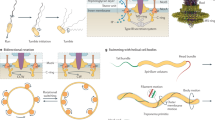
Prokaryotic cells have evolved numerous machineries to swim through liquid or crawl over surfaces. Perhaps the most common of these are the well-studied bacterial flagella and the unrelated archaeal flagella, which both function as rotary propellers. Extension and retraction of type IV pili allow movement over surfaces, as do a range of apparently unrelated gliding motors. In addition, prokaryotes can move passively by floating and sliding.
-
The bacterial flagellum is the best understood prokaryotic motility structure. It consists of a motor and a basal body that are embedded in the cell envelope and a long filament that usually extends from the cell. Rotation of the flagellum is driven by gradients of protons or sodium ions across the cytoplasmic membrane.
-
Bacterial flagella assemble by an unusual process that involves transport of many of the component proteins via a type III secretion system through the core of the basal body and filament before they are added to the tip of the growing structure. Expression of the genes that encode flagellar proteins is highly regulated in a hierarchical manner.
-
The flagellar filaments of spirochaetes are present entirely within the periplasm of the cell. Rotation of these periplasmic flagella is thought to result in movement of the cytoplasmic and outer membranes and attached structures in opposite directions, and results in cell movements.
-
Although bacterial flagella are usually involved in swimming in liquid, some bacteria express numerous flagella and use these to swim or crawl over moist surfaces in a process that is known as swarming.
-
Flagella are the motility structures that are responsible for swimming in the Archaea. Although these structures rotate, in common with bacterial flagella, the archaeal flagella lack a central channel, which means that they must assemble differently from bacterial flagella. Instead, archaeal flagella seem to share similarities in structure and assembly to bacterial type IV pili.
-
Some bacteria swim without using flagella. The wall-less spiroplasmas seem to use their well-developed cytoskeletons to alter their cell shape, which results in cell movement. Although some unicellular marine cyanobacteria swim, the mechanism (or mechanisms) that underlies this movement remains undefined.
-
Bacteria have various non-flagellar mechanisms that are used for crawling over surfaces. Type IV pilus extension and retraction pulls cells over moist surfaces by twitching motility. Flavobacterium spp. gliding motility involves a motor embedded in the cell envelope that moves adhesins along the cell surface. Two models have recently been proposed for Myxococcus spp. adventurous gliding: first, extrusion of polysaccharides, and second, a focal adhesion model that involves movement of cell surface adhesins by cytoplasmic motor proteins that interact with the cytoskeleton. Mycoplasma spp. might glide by 'inchworm' movements that involve the cytoskeleton or by 'centipede' movements of multiple leg-like structures on the cell surface.
Abstract
Prokaryotic cells move through liquids or over moist surfaces by swimming, swarming, gliding, twitching or floating. An impressive diversity of motility mechanisms has evolved in prokaryotes. Movement can involve surface appendages, such as flagella that spin, pili that pull and Mycoplasma 'legs' that walk. Internal structures, such as the cytoskeleton and gas vesicles, are involved in some types of motility, whereas the mechanisms of some other types of movement remain mysterious. Regardless of the type of motility machinery that is employed, most motile microorganisms use complex sensory systems to control their movements in response to stimuli, which allows them to migrate to optimal environments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Macnab, R. M. Type III flagellar protein export and flagellar assembly. Biochim. Biophys. Acta 1694, 207–217 (2004).
Macnab, R. M. How bacteria assemble flagella. Annu. Rev. Microbiol. 57, 77–100 (2003).
Aldridge, P. & Hughes, K. T. Regulation of flagellar assembly. Curr. Opin. Microbiol. 5, 160–165 (2002).
Apel, D. & Surette, M. G. Bringing order to a complex molecular machine: the assembly of the bacterial flagella. Biochim. Biophys. Acta 24 July 2007 (doi: 10.1016/j.bbamem.2007.07.005).
Kaiser, D. Bacterial swarming: a re-examination of cell-movement patterns. Curr. Biol. 17, R561–R570 (2007).
Lowe, G., Meister, M. & Berg, H. C. Rapid rotation of flagellar bundles in swimming bacteria. Nature 325, 637–640 (1987).
Lambert, C. et al. Characterizing the flagellar filament and the role of motility in bacterial prey-penetration by Bdellovibrio bacteriovorus. Mol. Microbiol. 60, 274–286 (2006).
Minamino, T. & Namba, K. Self-assembly and type III protein export of the bacterial flagellum. J. Mol. Microbiol. Biotechnol. 7, 5–17 (2004).
Thomas, D. R., Francis, N. R., Xu, C. & DeRosier, D. J. The three-dimensional structure of the flagellar rotor from a clockwise-locked mutant of Salmonella enterica serovar Typhimurium. J. Bacteriol. 188, 7039–7048 (2006).
Berg, H. C. & Anderson, R. A. Bacteria swim by rotating their flagellar filaments. Nature 245, 380–382 (1973).
McCarter, L. L. Dual flagellar systems enable motility under different circumstances. J. Mol. Microbiol. Biotechnol. 7, 18–29 (2004).
Kojima, S. & Blair, D. F. Conformational change in the stator of the bacterial flagellar motor. Biochemistry 40, 13041–13050 (2001).
Blair, D. F. Flagellar movement driven by proton translocation. FEBS Lett. 545, 86–95 (2003).
Chen, X. & Berg, H. C. Torque-speed relationship of the flagellar rotary motor of Escherichia coli. Biophys. J. 78, 1036–1041 (2000).
Magariyama, Y. et al. Very fast flagellar rotation. Nature 371, 752 (1994).
Aldridge, P. D. et al. The flagellar-specific transcription factor, σ28, is the type III secretion chaperone for the flagellar-specific anti-σ28 factor FlgM. Genes Dev. 20, 2315–2326 (2006).
Evans, L. D., Stafford, G. P., Ahmed, S., Fraser, G. M. & Hughes, C. An escort mechanism for cycling of export chaperones during flagellum assembly. Proc. Natl Acad. Sci. USA 103, 17474–17479 (2006). Showed that FliJ has a novel chaperone-escort function in flagellum assembly by specifically recruiting unladen chaperones for the minor filament-type subunits (hook-associated proteins) and transferring them to their cognate subunits. Because FliJ does not escort the chaperone for flagellin, these data suggest a mechanism that would favour the export of the minor subunits to aid the early formation of the hook-filament junction and the capping structure.
Stafford, G. P. et al. Sorting of early and late flagellar subunits after docking at the membrane ATPase of the type III export pathway. J. Mol. Biol. 374, 877–882 (2007).
Paul, K., Erhardt, M., Hirano, T., Blair, D. F. & Hughes, K. T. Energy source of flagellar type III secretion. Nature 451, 489–492 (2008). The energy source for the transport of flagellin subunits through the T3SS, originally thought to be supplied by the hydrolysis of ATP by FliI, was shown instead to be dependent on the proton motive force, with ATP hydrolysis not being essential for secretion.
Yonekura, K. et al. The bacterial flagellar cap as the rotary promoter of flagellin self-assembly. Science 290, 2148–2152 (2000).
Lee, H. J. & Hughes, K. T. Posttranscriptional control of the Salmonella enterica flagellar hook protein FlgE. J. Bacteriol. 188, 3308–3316 (2006).
McCarter, L. L. Regulation of flagella. Curr. Opin. Microbiol. 9, 180–186 (2006).
Aldridge, P., Karlinsey, J. E., Becker, E., Chevance, F. F. & Hughes, K. T. Flk prevents premature secretion of the anti-σ factor FlgM into the periplasm. Mol. Microbiol. 60, 630–643 (2006).
Aldridge, P., Karlinsey, J. & Hughes, K. T. The type III secretion chaperone FlgN regulates flagellar assembly via a negative feedback loop containing its chaperone substrates FlgK and FlgL. Mol. Microbiol. 49, 1333–1345 (2003).
Ferris, H. U. & Minamino, T. Flipping the switch: bringing order to flagellar assembly. Trends Microbiol. 14, 519–526 (2006).
Waters, R. C., O'Toole, P. W. & Ryan, K. A. The FliK protein and flagellar hook-length control. Protein Sci. 16, 769–780 (2007).
Journet, L., Agrain, C., Broz, P. & Cornelis, G. R. The needle length of bacterial injectisomes is determined by a molecular ruler. Science 302, 1757–1760 (2003).
Shibata, S. et al. FliK regulates flagellar hook length as an internal ruler. Mol. Microbiol. 64, 1404–1415 (2007).
Ferris, H. U. et al. FlhB regulates ordered export of flagellar components via autocleavage mechanism. J. Biol. Chem. 280, 41236–41242 (2005).
Minamino, T., Ferris, H. U., Moriya, N., Kihara, M. & Namba, K. Two parts of the T3S4 domain of the hook-length control protein FliK are essential for the substrate specificity switching of the flagellar type III export apparatus. J. Mol. Biol. 362, 1148–1158 (2006).
Charon, N. W. & Goldstein, S. F. Genetics of motility and chemotaxis of a fascinating group of bacteria: the spirochetes. Annu. Rev. Genet. 36, 47–73 (2002).
England, J. C. & Gober, J. W. Cell cycle control of cell morphogenesis in Caulobacter. Curr. Opin. Microbiol. 4, 674–680 (2001).
Wolgemuth, C. W., Charon, N. W., Goldstein, S. F. & Goldstein, R. E. The flagellar cytoskeleton of the spirochetes. J. Mol. Microbiol. Biotechnol. 11, 221–227 (2006).
Murphy, G. E., Matson, E. G., Leadbetter, J. R., Berg, H. C. & Jensen, G. J. Novel ultrastructures of Treponema primitia and their implications for motility. Mol. Microbiol. 67, 1184–1195 (2008).
Murphy, G. E., Leadbetter, J. R. & Jensen, G. J. In situ structure of the complete Treponema primitia flagellar motor. Nature 442, 1062–1064 (2006).
Harshey, R. M. Bacterial motility on a surface: many ways to a common goal. Annu. Rev. Microbiol. 57, 249–273 (2003).
Merino, S., Shaw, J. G. & Tomas, J. M. Bacterial lateral flagella: an inducible flagella system. FEMS Microbiol. Lett. 263, 127–135 (2006).
Atsumi, T., McCarter, L. & Imae, Y. Polar and lateral flagellar motors of marine Vibrio are driven by different ion-motive forces. Nature 355, 182–184 (1992).
Thomas, N. A., Bardy, S. L. & Jarrell, K. F. The archaeal flagellum: a different kind of prokaryotic motility structure. FEMS Microbiol. Rev. 25, 147–174 (2001).
Jarrell, K. F., Ng, S. Y. & Chaban, B. in Archaea: Molecular and Cellular Biology (ed. Cavicchioli, R.) 385–410 (ASM, Washington DC, 2007).
Alam, M., Claviez, M., Oesterhelt, D. & Kessel, M. Flagella and motility behaviour of square bacteria. EMBO J. 3, 2899–2903 (1984).
Marwan, W., Alam, M. & Oesterhelt, D. Rotation and switching of the flagellar motor assembly in Halobacterium halobium. J. Bacteriol. 173, 1971–1977 (1991).
Kupper, J. et al. The flagellar bundle of Halobacterium salinarium is inserted into a distinct polar cap structure. J. Bacteriol. 176, 5184–5187 (1994).
Metlina, A. L. Bacterial and archaeal flagella as prokaryotic motility organelles. Biochemistry (Mosc.) 69, 1203–1212 (2004).
Jarrell, K. F. & Koval, S. F. Ultrastructure and biochemistry of Methanococcus voltae. Crit. Rev. Microbiol. 17, 53–87 (1989).
Faguy, D. M., Jarrell, K. F., Kuzio, J. & Kalmokoff, M. L. Molecular analysis of archael flagellins: similarity to the type IV pilin-transport superfamily widespread in bacteria. Can. J. Microbiol. 40, 67–71 (1994).
Bardy, S. L. & Jarrell, K. F. Cleavage of preflagellins by an aspartic acid signal peptidase is essential for flagellation in the archaeon Methanococcus voltae. Mol. Microbiol. 50, 1339–1347 (2003).
Bardy, S. L. & Jarrell, K. F. FlaK of the archaeon Methanococcus maripaludis possesses preflagellin peptidase activity. FEMS Microbiol. Lett. 208, 53–59 (2002).
Albers, S. V., Szabo, Z. & Driessen, A. J. Archaeal homolog of bacterial type IV prepilin signal peptidases with broad substrate specificity. J. Bacteriol. 185, 3918–3925 (2003).
Ng, S. Y., Chaban, B. & Jarrell, K. F. Archaeal flagella, bacterial flagella and type IV pili: a comparison of genes and posttranslational modifications. J. Mol. Microbiol. Biotechnol. 11, 167–191 (2006).
Faguy, D. M. & Jarrell, K. F. A twisted tale: the origin and evolution of motility and chemotaxis in prokaryotes. Microbiology 145, 279–281 (1999).
Nutsch, T., Oesterhelt, D., Gilles, E. D. & Marwan, W. A quantitative model of the switch cycle of an archaeal flagellar motor and its sensory control. Biophys. J. 89, 2307–2323 (2005).
Cohen-Krausz, S. & Trachtenberg, S. The structure of the archeabacterial flagellar filament of the extreme halophile Halobacterium salinarum R1M1 and its relation to eubacterial flagellar filaments and type IV pili. J. Mol. Biol. 321, 383–395 (2002).
Cohen-Krausz, S. & Trachtenberg, S. The flagellar filament structure of the extreme acidothermophile Sulfolobus shibatae B12 suggests that archaeabacterial flagella have a unique and common symmetry and design. J. Mol. Biol. 375, 1113–1124 (2008). Analysed two phylogenetically distant archaea to show that their archaeal flagellar filament structures were unique compared with bacterial filaments. Also confirmed the lack of a central channel in both flagella filaments, a feature that is crucial for assembly models.
Trachtenberg, S. & Cohen-Krausz, S. The archaeabacterial flagellar filament: a bacterial propeller with a pilus-like structure. J. Mol. Microbiol. Biotechnol. 11, 208–220 (2006).
Jarrell, K. F., Bayley, D. P. & Kostyukova, A. S. The archaeal flagellum: a unique motility structure. J. Bacteriol. 178, 5057–5064 (1996).
Chaban, B. et al. Systematic deletion analyses of the fla genes in the flagella operon identify several genes essential for proper assembly and function of flagella in the archaeon, Methanococcus maripaludis. Mol. Microbiol. 66, 596–609 (2007).
Bardy, S. L., Mori, T., Komoriya, K., Aizawa, S. & Jarrell, K. F. Identification and localization of flagellins FlaA and FlaB3 within flagella of Methanococcus voltae. J. Bacteriol. 184, 5223–5233 (2002).
Beznosov, S. N., Pyatibratov, M. G. & Fedorov, O. V. On the multicomponent nature of Halobacterium salinarum flagella. Microbiology (Russ.) 76, 435–441 (2007).
Logan, S. M. Flagellar glycosylation — a new component of the motility repertoire? Microbiology 152, 1249–1262 (2006).
Lechner, J. & Wieland, F. Structure and biosynthesis of prokaryotic glycoproteins. Annu. Rev. Biochem. 58, 173–194 (1989).
Sumper, M. Halobacterial glycoprotein biosynthesis. Biochim. Biophys. Acta 906, 69–79 (1987).
Voisin, S. et al. Identification and characterization of the unique N-linked glycan common to the flagellins and S-layer glycoprotein of Methanococcus voltae. J. Biol. Chem. 280, 16586–16593 (2005).
Chaban, B., Voisin, S., Kelly, J., Logan, S. M. & Jarrell, K. F. Identification of genes involved in the biosynthesis and attachment of Methanococcus voltae N-linked glycans: insight into N-linked glycosylation pathways in Archaea. Mol. Microbiol. 61, 259–268 (2006). Identified genes that are involved in N -linked glycosylation in archaea. Also showed that underglycosylated flagellins are either not incorporated or incorporated only poorly into filaments, thereby demonstrating an essential role for glycosylation in the assembly of archaeal flagella.
Wolgemuth, C. W. & Charon, N. W. The kinky propulsion of Spiroplasma. Cell 122, 827–828 (2005).
Kurner, J., Frangakis, A. S. & Baumeister, W. Cryo-electron tomography reveals the cytoskeletal structure of Spiroplasma melliferum. Science 307, 436–438 (2005).
Trachtenberg, S. The cytoskeleton of Spiroplasma: a complex linear motor. J. Mol. Microbiol. Biotechnol. 11, 265–283 (2006).
Shaevitz, J. W., Lee, J. Y. & Fletcher, D. A. Spiroplasma swim by a processive change in body helicity. Cell 122, 941–945 (2005).
Waterbury, J. B., Willey, J. M., Franks, D. G., Valois, F. W. & Watson, S. W. A cyanobacterium capable of swimming motility. Science 230, 74–76 (1985).
Willey, J. M., Waterbury, J. B. & Greenberg, E. P. Sodium-coupled motility in a swimming cyanobacterium. J. Bacteriol. 169, 3429–3434 (1987).
McCarren, J. & Brahamsha, B. SwmB, a 1.12-megadalton protein that is required for nonflagellar swimming motility in Synechococcus. J. Bacteriol. 189, 1158–1162 (2007).
Brahamsha, B. An abundant cell-surface polypeptide is required for swimming by the nonflagellated marine cyanobacterium Synechococcus. Proc. Natl Acad. Sci. USA 93, 6504–6509 (1996).
Samuel, A. D., Petersen, J. D. & Reese, T. S. Envelope structure of Synechococcus sp. WH8113, a nonflagellated swimming cyanobacterium. BMC Microbiol. 1, 4 (2001).
Ehlers, K. M., Samuel, A. D., Berg, H. C. & Montgomery, R. Do cyanobacteria swim using traveling surface waves? Proc. Natl Acad. Sci. USA 93, 8340–8343 (1996).
Skerker, J. M. & Berg, H. C. Direct observation of extension and retraction of type IV pili. Proc. Natl Acad. Sci. USA 98, 6901–6904 (2001). Reported the direct observation of extension and retraction of fluorescently labelled T4P, which are associated with twitching motility.
Merz, A. J., So, M. & Sheetz, M. P. Pilus retraction powers bacterial twitching motility. Nature 407, 98–102 (2000).
Mattick, J. S. Type IV pili and twitching motility. Annu. Rev. Microbiol. 56, 289–314 (2002).
Duggan, P. S., Gottardello, P. & Adams, D. G. Molecular analysis of genes in Nostoc punctiforme involved in pilus biogenesis and plant infection. J. Bacteriol. 189, 4547–4551 (2007).
Varga, J. J. et al. Type IV pili-dependent gliding motility in the Gram-positive pathogen Clostridium perfringens and other Clostridia. Mol. Microbiol. 62, 680–694 (2006).
Carbonnelle, E., Helaine, S., Nassif, X. & Pelicic, V. A systematic genetic analysis in Neisseria meningitidis defines the Pil proteins required for assembly, functionality, stabilization and export of type IV pili. Mol. Microbiol. 61, 1510–1522 (2006).
Craig, L., Pique, M. E. & Tainer, J. A. Type IV pilus structure and bacterial pathogenicity. Nature Rev. Microbiol. 2, 363–378 (2004).
Burrows, L. L. Weapons of mass retraction. Mol. Microbiol. 57, 878–888 (2005).
Satyshur, K. A. et al. Crystal structures of the pilus retraction motor PilT suggest large domain movements and subunit cooperation drive motility. Structure 15, 363–376 (2007).
Crowther, L. J., Anantha, R. P. & Donnenberg, M. S. The inner membrane subassembly of the enteropathogenic Escherichia coli bundle-forming pilus machine. Mol. Microbiol. 52, 67–79 (2004).
Craig, L. et al. Type IV pilus structure by cryo-electron microscopy and crystallography: implications for pilus assembly and functions. Mol. Cell 23, 651–662 (2006).
Winther-Larsen, H. C. et al. A conserved set of pilin-like molecules controls type IV pilus dynamics and organelle-associated functions in Neisseria gonorrhoeae. Mol. Microbiol. 56, 903–917 (2005).
Bradley, D. E. A function of Pseudomonas aeruginosa PAO polar pili: twitching motility. Can. J. Microbiol. 26, 146–154 (1980).
Sun, H., Zusman, D. R. & Shi, W. Type IV pilus of Myxococcus xanthus is a motility apparatus controlled by the frz chemosensory system. Curr. Biol. 10, 1143–1146 (2000).
Morand, P. C. et al. Type IV pilus retraction in pathogenic Neisseria is regulated by the PilC proteins. EMBO J. 23, 2009–2017 (2004).
McBride, M. J. Bacterial gliding motility: multiple mechanisms for cell movement over surfaces. Annu. Rev. Microbiol. 55, 49–75 (2001).
Braun, T. F., Khubbar, M. K., Saffarini, D. A. & McBride, M. J. Flavobacterium johnsoniae gliding motility genes identified by mariner mutagenesis. J. Bacteriol. 187, 6943–6952 (2005).
Nelson, S. S., Bollampalli, S. & McBride, M. J. SprB is a cell-surface component of the Flavobacterium johnsoniae gliding motility machinery. J. Bacteriol. 190, 2851–2857 (2008). Presented evidence that the 669 kDa protein SprB is a moving cell surface component of the F. johnsoniae gliding motility machinery.
Liu, J., McBride, M. J. & Subramaniam, S. Cell surface filaments of the gliding bacterium Flavobacterium johnsoniae revealed by cryo-electron tomography. J. Bacteriol. 189, 7503–7506 (2007).
Wall, D. & Kaiser, D. Type IV pili and cell motility. Mol. Microbiol. 32, 1–10 (1999).
Wolgemuth, C., Hoiczyk, E., Kaiser, D. & Oster, G. How myxobacteria glide. Curr. Biol. 12, 369–377 (2002).
Hoiczyk, E. & Baumeister, W. The junctional pore complex, a prokaryotic secretion organelle, is the molecular motor underlying gliding motility in cyanobacteria. Curr. Biol. 8, 1161–1168 (1998).
Yu, R. & Kaiser, D. Gliding motility and polarized slime secretion. Mol. Microbiol. 63, 454–467 (2007).
White, D. J. & Hartzell, P. L. AglU, a protein required for gliding motility and spore maturation of Myxococcus xanthus, is related to WD-repeat proteins. Mol. Microbiol. 36, 662–678 (2000).
Youderian, P., Burke, N., White, D. J. & Hartzell, P. L. Identification of genes required for adventurous gliding motility in Myxococcus xanthus with the transposable element mariner. Mol. Microbiol. 49, 555–570 (2003).
Mignot, T., Shaevitz, J. W., Hartzell, P. L. & Zusman, D. R. Evidence that focal adhesion complexes power bacterial gliding motility. Science 315, 853–856 (2007). Showed that the M. xanthus adventurous gliding motility protein AglZ remained fixed relative to the substratum as cells glided. These data suggest a model of cell movement that involves intracellular motors that exert force on the cytoskeleton and on cell surface adhesion complexes.
Heintzelman, M. B. Cellular and molecular mechanics of gliding locomotion in eukaryotes. Int. Rev. Cytol. 251, 79–129 (2006).
Miyata, M. Centipede and inchworm models to explain Mycoplasma gliding. Trends Microbiol. 16, 6–12 (2008).
Hasselbring, B. M. & Krause, D. C. Cytoskeletal protein P41 is required to anchor the terminal organelle of the wall-less prokaryote Mycoplasma pneumoniae. Mol. Microbiol. 63, 44–53 (2007).
Uenoyama, A. & Miyata, M. Identification of a 123-kilodalton protein (Gli123) involved in machinery for gliding motility of Mycoplasma mobile. J. Bacteriol. 187, 5578–5584 (2005).
Seto, S., Uenoyama, A. & Miyata, M. Identification of a 521-kilodalton protein (Gli521) involved in force generation or force transmission for Mycoplasma mobile gliding. J. Bacteriol. 187, 3502–3510 (2005).
Uenoyama, A. & Miyata, M. Gliding ghosts of Mycoplasma mobile. Proc. Natl Acad. Sci. USA 102, 12754–12758 (2005). Presented evidence that the energy source for M. mobile gliding is ATP. Cells were permeabilized with Triton X-100 to form non-motile and non-viable ghosts that regained motility when ATP was added exogenously.
Ohtani, N. & Miyata, M. Identification of a novel nucleoside triphosphatase from Mycoplasma mobile: a prime candidate motor for gliding motility. Biochem. J. 403, 71–77 (2007).
Seybert, A., Herrmann, R. & Frangakis, A. S. Structural analysis of Mycoplasma pneumoniae by cryo-electron tomography. J. Struct. Biol. 156, 342–354 (2006).
Henderson, G. P. & Jensen, G. J. Three-dimensional structure of Mycoplasma pneumoniae's attachment organelle and a model for its role in gliding motility. Mol. Microbiol. 60, 376–385 (2006).
Martinez, A., Torello, S. & Kolter, R. Sliding motility in mycobacteria. J. Bacteriol. 181, 7331–7338 (1999).
Stevens, J. M., Galyov, E. E. & Stevens, M. P. Actin-dependent movement of bacterial pathogens. Nature Rev. Microbiol. 4, 91–101 (2006).
Szurmant, H. & Ordal, G. W. Diversity in chemotaxis mechanisms among the bacteria and archaea. Microbiol. Mol. Biol. Rev. 68, 301–319 (2004).
Turner, L., Ryu, W. S. & Berg, H. C. Real-time imaging of fluorescent flagellar filaments. J. Bacteriol. 182, 2793–2801 (2000).
Rudolph, J. & Oesterhelt, D. Deletion analysis of the che operon in the archaeon Halobacterium salinarium. J. Mol. Biol. 258, 548–554 (1996).
Rudolph, J., Tolliday, N., Schmitt, C., Schuster, S. C. & Oesterhelt, D. Phosphorylation in halobacterial signal transduction. EMBO J. 14, 4249–4257 (1995).
Zusman, D. R., Scott, A. E., Yang, Z. & Kirby, J. R. Chemosensory pathways, motility and development in Myxococcus xanthus. Nature Rev. Microbiol. 5, 862–872 (2007).
Mignot, T., Merlie, J. P. Jr & Zusman, D. R. Regulated pole-to-pole oscillations of a bacterial gliding motility protein. Science 310, 855–857 (2005).
Leonardy, S., Freymark, G., Hebener, S., Ellehauge, E. & Sogaard-Andersen, L. Coupling of protein localization and cell movements by a dynamically localized response regulator in Myxococcus xanthus. EMBO J. 26, 4433–4444 (2007).
Bonner, P. J. & Shimkets, L. J. Phospholipid directed motility of surface-motile bacteria. Mol. Microbiol. 61, 1101–1109 (2006).
Beard, S. J., Hayes, P. K., Pfeifer, F. & Walsby, A. E. The sequence of the major gas vesicle protein, GvpA, influences the width and strength of halobacterial gas vesicles. FEMS Microbiol. Lett. 213, 149–157 (2002).
Walsby, A. E. Gas vesicles. Microbiol. Rev. 58, 94–144 (1994).
Mlouka, A., Comte, K., Castets, A. M., Bouchier, C. & Tandeau de Marsac, N. The gas vesicle gene cluster from Microcystis aeruginosa and DNA rearrangements that lead to loss of cell buoyancy. J. Bacteriol. 186, 2355–2365 (2004).
Offner, S., Hofacker, A., Wanner, G. & Pfeifer, F. Eight of fourteen gvp genes are sufficient for formation of gas vesicles in halophilic archaea. J. Bacteriol. 182, 4328–4336 (2000).
DasSarma, S., Arora, P., Lin, F., Molinari, E. & Yin, L. R. Wild-type gas vesicle formation requires at least ten genes in the gvp gene cluster of Halobacterium halobium plasmid pNRC100. J. Bacteriol. 176, 7646–7652 (1994).
Scheuch, S. & Pfeifer, F. GvpD-induced breakdown of the transcriptional activator GvpE of halophilic archaea requires a functional p-loop and an arginine-rich region of GvpD. Microbiology 153, 947–958 (2007).
Offner, S., Ziese, U., Wanner, G., Typke, D. & Pfeifer, F. Structural characteristics of halobacterial gas vesicles. Microbiology 144, 1331–1342 (1998).
Walsby, A. E. & Dunton, P. G. Gas vesicles in actinomycetes? Trends Microbiol. 14, 99–100 (2006).
Pfeifer, F. et al. Gas vesicle formation in halophilic Archaea. Arch. Microbiol. 167, 259–268 (1997).
Dunton, P. G., Mawby, W. J., Shaw, V. A. & Walsby, A. E. Analysis of tryptic digests indicates regions of GvpC that bind to gas vesicles of Anabaena flos-aquae. Microbiology 152, 1661–1669 (2006).
Buchholz, B. E., Hayes, P. K. & Walsby, A. E. The distribution of the outer gas vesicle protein, GvpC, on the Anabaena gas vesicle, and its ratio to GvpA. J. Gen. Microbiol. 139, 2353–2363 (1993).
Shukla, H. D. & DasSarma, S. Complexity of gas vesicle biogenesis in Halobacterium sp. strain NRC-1: identification of five new proteins. J. Bacteriol. 186, 3182–3186 (2004).
Walsby, A. E., Ng, G., Dunn, C. & David, P. A. Comparison of the depth where Planktothrix rubescens stratifies and the depth where the daily insolation supports its neutral bouyancy. New Phytol. 162, 133–145 (2004).
Oliver, R. L. & Walsby, A. E. Direct evidence for the role of light-mediated gas vesicle collapse in the bouyancy regulation of Anabaena flos-aquae (cyanobacteria). Limnol. Oceanogr. 29, 879–886 (1984).
Macnab, R. M. The bacterial flagellum: reversible rotary propellor and type III export apparatus. J. Bacteriol. 181, 7149–7153 (1999).
Acknowledgements
The authors thank the members of their laboratories for helpful discussions and the researchers who generously supplied videos. Research in the authors laboratories is supported by grants from the National Science Foundation (MCB-0641366) to M.J.M. and from the Natural Sciences and Engineering Research Council of Canada to K.F.J.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary information S1 (movie)
Movement of SprB protein on cell surface of gliding cells of Flavobacterium johnsoniae. Protein G coated 0.5 μm polystyrene spheres with anti-SprB antibodies were added to cells of Flavobacterium johnsoniae on a glass slide and images were recorded using a phase-contrast microscope. Bar indicates 10 μm. (MOV 4932 kb)
Supplementary information S2 (movie)
Preparation and reactivation of Mycoplasma mobile ghosts. The cells gliding on a glass coverslip were treated with 0.01% Triton X 100, 1 mg/ml each of DNase and RNase, and ATP at the times indicated in the video. Reproduced with permission from: Uenoyama, A. & Miyata, M. Gliding ghosts of Mycoplasma mobile. Proc. Natl. Acad. Sci. USA 102, 12754–12758 (2005) © (2005) National Academy of Sciences. Video courtesy of Makoto Miyata. (MOV 8084 kb)
Supplementary information S3 (movie)
Digital microcinematography of cell-independent gliding of detached mutant MPN311 terminal organelles of Mycoplasma pnuemoniae. Reproduced with permission from: Hasselbring, B. M. & Krause, D. C. Cytoskeletal protein P41 is required to anchor the terminal organelle of the wall-less prokaryote Mycoplasma pneumoniae. Mol. Microbiol. 63, 44 53 (2007) © (2007) Blackwell Publishing. Video courtesy of Duncan Krause. (AVI 8406 kb)
Supplementary information S4 (movie)
Digital microcinematography of Mycoplasma pnuemoniae mutant MPN311 terminal organelle detachment after cell intersection. Reproduced with permission from: Hasselbring, B. M. & Krause, D. C. Cytoskeletal protein P41 is required to anchor the terminal organelle of the wall-less prokaryote Mycoplasma pneumoniae. Mol. Microbiol. 63, 44–53 (2007) © (2007) Blackwell Publishing. Video courtesy of Duncan Krause. (AVI 6431 kb)
Related links
Related links
DATABASES
Entrez Genome Project
Entrez Protein
FURTHER INFORMATION
Howard Berg's laboratory website (movies: bacteria swarming; type IV pili 1 and type IV pili 2)
Howard Berg's laboratory website (movies: bacteria swarming; type IV pili 1 and type IV pili 2)
Howard Berg's laboratory website (movies: bacteria swarming; type IV pili 1 and type IV pili 2)
Howard Berg's laboratory website (movies: bacteria swarming; type IV pili 1 and type IV pili 2)
Joshua Shaevitz's laboratory website (movie: Spiroplasma kinking)
Keiichi Namba's laboratory website (movie: flagella assembly)
Nyles Charon's laboratory website (movie: Borrelia swimming)
Glossary
- Proton motive force
-
A special case of an electrochemical potential. Proton motive force is the force that is created by the accumulation of protons on one side of a cell membrane. This concentration gradient is generated using energy sources, such as redox potential or ATP. Once established, the proton motive force can be used to carry out work, for example, to synthesize ATP or pump compounds across the membrane.
- Chemotaxis
-
Directed movement towards attractants or away from repellents.
- Halophile
-
A bacterium or archaeon that can grow in environments that contain high concentrations of salt (at least 2 M).
- Signal peptide
-
A short (3–60 amino acid long) peptide chain that directs the post-translational transport of a protein. Signal peptides are also known as targeting signals, signal sequences, transit peptides or localization signals.
Rights and permissions
About this article
Cite this article
Jarrell, K., McBride, M. The surprisingly diverse ways that prokaryotes move. Nat Rev Microbiol 6, 466–476 (2008). https://doi.org/10.1038/nrmicro1900
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1900
This article is cited by
-
Motility mediates satellite formation in confined biofilms
The ISME Journal (2023)
-
The Facile Synthesis of Eco-Friendly Zinc Magnesium Bimetal Nanoparticles and its Application in the Eradication of Xanthomonas oryzae pv. oryzae that Causes Leaf Blight Disease of Rice
Current Microbiology (2023)
-
Phylogenomics of the Liquorilactobacillus Genus
Current Microbiology (2023)
-
Kin discrimination drives territorial exclusion during Bacillus subtilis swarming and restrains exploitation of surfactin
The ISME Journal (2022)
-
Structural insights into the mechanism of archaellar rotational switching
Nature Communications (2022)