Key Points
-
Major epigenetic reprogramming occurs during pre-implantation development. The precise functions of these changes in cell fate allocation remain to be addressed.
-
Genetic approaches in the past have uncovered important principles and players involved in lineage allocation during pre-implantation development.
-
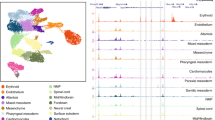
Recently, single-cell expression profiling and novel microscopy techniques have provided new insights into transcriptional and chromatin-regulated events that are responsible for lineage allocation.
-
Global chromatin mobility, as well as differential expression of chromatin modifiers in single cells, might promote cell fate allocation in the early embryo.
-
Future research focusing on single cells will provide key insights into the mechanisms that drive and enforce cell fate allocation decisions.
Abstract
Following fertilization, gametes undergo epigenetic reprogramming in order to revert to a totipotent state. How embryonic cells subsequently acquire their fate and the role of chromatin dynamics in this process are unknown. Genetic and experimental embryology approaches have identified some of the players and morphological changes that are involved in early mammalian development, but the exact events underlying cell fate allocation in single embryonic cells have remained elusive. Experimental and technological advances have recently provided novel insights into chromatin dynamics and nuclear architecture in single cells; these insights have reshaped our understanding of the mechanisms underlying cell fate allocation and plasticity in early mammalian development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lehtonen, E. Changes in cell dimensions and intercellular contacts during cleavage-stage cell cycles in mouse embryonic cells. J. Embryol. Exp. Morphol. 58, 231–249 (1980).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Vermeulen, M. et al. Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142, 967–980 (2010).
Musselman, C. A., Lalonde, M. E., Cote, J. & Kutateladze, T. G. Perceiving the epigenetic landscape through histone readers. Nature Struct. Mol. Biol. 19, 1218–1227 (2012).
Tropberger, P. et al. Regulation of transcription through acetylation of H3K122 on the lateral surface of the histone octamer. Cell 152, 859–872 (2013).
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Morgan, H. D., Santos, F., Green, K., Dean, W. & Reik, W. Epigenetic reprogramming in mammals. Hum. Mol. Genet. 14, R47–R58 (2005).
Burton, A. & Torres-Padilla, M. E. Epigenetic reprogramming and development: a unique heterochromatin organization in the preimplantation mouse embryo. Brief Funct. Genom. 9, 444–454 (2011).
Hemberger, M., Dean, W. & Reik, W. Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington's canal. Nature Rev. Mol. Cell Biol. 10, 526–537 (2009).
Leitch, H. G., Tang, W. W. & Surani, M. A. Primordial germ-cell development and epigenetic reprogramming in mammals. Curr. Top. Dev. Biol. 104, 149–187 (2013).
Rossant, J. & Tam, P. P. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Johnson, M. H. & Ziomek, C. A. The foundation of two distinct cell lineages within the mouse morula. Cell 24, 71–80 (1981).
Johnson, M. H. From mouse egg to mouse embryo: polarities, axes, and tissues. Annu. Rev. Cell Dev. Biol. 25, 483–512 (2009).
Suwinska, A., Czolowska, R., Ozdzenski, W. & Tarkowski, A. K. Blastomeres of the mouse embryo lose totipotency after the fifth cleavage division: expression of Cdx2 and Oct4 and developmental potential of inner and outer blastomeres of 16- and 32-cell embryos. Dev. Biol. 322, 133–144 (2008).
Morris, S. A. et al. Origin and formation of the first two distinct cell types of the inner cell mass in the mouse embryo. Proc. Natl Acad. Sci. USA 107, 6364–6369 (2010).
Dietrich, J. E. & Hiiragi, T. Stochastic patterning in the mouse pre-implantation embryo. Development 134, 4219–4231 (2007).
Niwa, H. et al. Interaction between Oct3/4 and Cdx2 determines trophectoderm differentiation. Cell 123, 917–929 (2005).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000).
Nishioka, N. et al. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech. Dev. 125, 270–283 (2008).
Strumpf, D. et al. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132, 2093–2102 (2005).
Palmieri, S. L., Peter, W., Hess, H. & Scholer, H. R. Oct-4 transcription factor is differentially expressed in the mouse embryo during establishment of the first two extraembryonic cell lineages involved in implantation. Dev. Biol. 166, 259–267 (1994).
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010). Demonstrated for the first time the use of microfluidics to quantify gene expression patterns in the mouse pre-implantation embryo.
Tolkunova, E. et al. The caudal-related protein cdx2 promotes trophoblast differentiation of mouse embryonic stem cells. Stem Cells 24, 139–144 (2006).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014). Applied single-cell RNA-seq analysis of hybrid embryos to reveal allelic expression patterns.
Miyanari, Y. & Torres-Padilla, M. E. Control of ground-state pluripotency by allelic regulation of Nanog. Nature 483, 470–473 (2012).
Terranova, R. et al. Polycomb group proteins Ezh2 and Rnf2 direct genomic contraction and imprinted repression in early mouse embryos. Dev. Cell 15, 668–679 (2008).
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000).
Dean, W. et al. Conservation of methylation reprogramming in mammalian development: aberrant reprogramming in cloned embryos. Proc. Natl Acad. Sci. USA 98, 13734–13738 (2001).
Beaujean, N. et al. Non-conservation of mammalian preimplantation methylation dynamics. Curr. Biol. 14, R266–R267 (2004).
Smallwood, S. A. et al. Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nature Genet. 43, 811–814 (2011).
Kobayashi, H. et al. Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet. 8, e1002440 (2012).
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 10, 475–478 (2000).
Santos, F., Hendrich, B., Reik, W. & Dean, W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev. Biol. 241, 172–182 (2002).
Wossidlo, M. et al. Dynamic link of DNA demethylation, DNA strand breaks and repair in mouse zygotes. EMBO J. 29, 1877–1888 (2010).
Wossidlo, M. et al. 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nature Commun. 2, 241 (2011).
Hajkova, P. et al. Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway. Science 329, 78–82 (2010).
Smith, Z. D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Liu, H., Kim, J. M. & Aoki, F. Regulation of histone H3 lysine 9 methylation in oocytes and early pre-implantation embryos. Development 131, 2269–2280 (2004).
Santos, F., Peters, A. H., Otte, A. P., Reik, W. & Dean, W. Dynamic chromatin modifications characterise the first cell cycle in mouse embryos. Dev. Biol. 280, 225–236 (2005).
Puschendorf, M. et al. PRC1 and Suv39h specify parental asymmetry at constitutive heterochromatin in early mouse embryos. Nature Genet. 40, 411–420 (2008).
Santenard, A. et al. Heterochromatin formation in the mouse embryo requires critical residues of the histone variant H3.3. Nature Cell Biol. 12, 853–862 (2010).
Kourmouli, N. et al. Heterochromatin and tri-methylated lysine 20 of histone H4 in animals. J. Cell Sci. 117, 2491–2501 (2004).
Lepikhov, K. & Walter, J. Differential dynamics of histone H3 methylation at positions K4 and K9 in the mouse zygote. BMC Dev. Biol. 4, 12 (2004).
Daujat, S. et al. H3K64 trimethylation marks heterochromatin and is dynamically remodeled during developmental reprogramming. Nature Struct. Mol. Biol. 16, 777–781 (2009).
Ooga, M. et al. Changes in H3K79 methylation during preimplantation development in mice. Biol. Reprod. 78, 413–424 (2008).
Brykczynska, U. et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nature Struct. Mol. Biol. 17, 679–687 (2010).
Hammoud, S. S. et al. Distinctive chromatin in human sperm packages genes for embryo development. Nature 460, 473–478 (2009).
Potok, M. E., Nix, D. A., Parnell, T. J. & Cairns, B. R. Reprogramming the maternal zebrafish genome after fertilization to match the paternal methylation pattern. Cell 153, 759–772 (2013).
Erkek, S. et al. Molecular determinants of nucleosome retention at CpG-rich sequences in mouse spermatozoa. Nature Struct. Mol. Biol. 20, 868–875 (2013).
van der Heijden, G. W. et al. Asymmetry in histone H3 variants and lysine methylation between paternal and maternal chromatin of the early mouse zygote. Mech. Dev. 122, 1008–1022 (2005).
Torres-Padilla, M. E., Bannister, A. J., Hurd, P. J., Kouzarides, T. & Zernicka-Goetz, M. Dynamic distribution of the replacement histone variant H3.3 in the mouse oocyte and preimplantation embryos. Int. J. Dev. Biol. 50, 455–461 (2006).
Ziegler-Birling, C., Helmrich, A., Tora, L. & Torres-Padilla, M. E. Distribution of p53 binding protein 1 (53BP1) and phosphorylated H2A. X during mouse preimplantation development in the absence of DNA damage. Int. J. Dev. Biol. 53, 1003–1011 (2009).
Boskovic, A. et al. Analysis of active chromatin modifications in early mammalian embryos reveals uncoupling of H2A.Z acetylation and H3K36 trimethylation from embryonic genome activation. Epigenetics 7, 747–757 (2012).
Nashun, B., Yukawa, M., Liu, H., Akiyama, T. & Aoki, F. Changes in the nuclear deposition of histone H2A variants during pre-implantation development in mice. Development 137, 3785–3794 (2010).
Faast, R. et al. Histone variant H2A. Z is required for early mammalian development. Curr. Biol. 11, 1183–1187 (2001).
Rangasamy, D., Berven, L., Ridgway, P. & Tremethick, D. J. Pericentric heterochromatin becomes enriched with H2A.Z during early mammalian development. EMBO J. 22, 1599–1607 (2003).
Ahmed, K. et al. Global chromatin architecture reflects pluripotency and lineage commitment in the early mouse embryo. PLoS ONE 5, e10531 (2010).
Aguirre-Lavin, T. et al. 3D-FISH analysis of embryonic nuclei in mouse highlights several abrupt changes of nuclear organization during preimplantation development. BMC Dev. Biol. 12, 30 (2012).
Probst, A. V., Santos, F., Reik, W., Almouzni, G. & Dean, W. Structural differences in centromeric heterochromatin are spatially reconciled on fertilisation in the mouse zygote. Chromosoma 116, 403–415 (2007).
Burns, K. H. et al. Roles of NPM2 in chromatin and nucleolar organization in oocytes and embryos. Science 300, 633–636 (2003).
Jachowicz, J. W., Santenard, A., Bender, A., Muller, J. & Torres-Padilla, M. E. Heterochromatin establishment at pericentromeres depends on nuclear position. Genes Dev. 27, 2427–2432 (2013).
Ostrup, O. et al. Nuclear and nucleolar reprogramming during the first cell cycle in bovine nuclear transfer embryos. Clon. Stem Cells 11, 367–375 (2009).
Martin, C. et al. Genome restructuring in mouse embryos during reprogramming and early development. Dev. Biol. 292, 317–332 (2006).
Pichugin, A. et al. Dynamics of constitutive heterochromatin: two contrasted kinetics of genome restructuring in early cloned bovine embryos. Reproduction 139, 129–137 (2010).
Watanabe, D., Suetake, I., Tada, T. & Tajima, S. Stage- and cell-specific expression of Dnmt3a and Dnmt3b during embryogenesis. Mech. Dev. 118, 187–190 (2002).
Erhardt, S. et al. Consequences of the depletion of zygotic and embryonic enhancer of zeste 2 during preimplantation mouse development. Development 130, 4235–4248 (2003).
Sarmento, O. F. et al. Dynamic alterations of specific histone modifications during early murine development. J. Cell Sci. 117, 4449–4459 (2004).
Rugg-Gunn, P. J., Cox, B. J., Ralston, A. & Rossant, J. Distinct histone modifications in stem cell lines and tissue lineages from the early mouse embryo. Proc. Natl Acad. Sci. USA 107, 10783–10790 (2010).
Alder, O. et al. Ring1B and Suv39h1 delineate distinct chromatin states at bivalent genes during early mouse lineage commitment. Development 137, 2483–2492 (2010).
Surani, M. A., Hayashi, K. & Hajkova, P. Genetic and epigenetic regulators of pluripotency. Cell 128, 747–762 (2007).
Li, E. Chromatin modification and epigenetic reprogramming in mammalian development. Nature Rev. Genet. 3, 662–673 (2002).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
O'Carroll, D. et al. The polycomb-group gene Ezh2 is required for early mouse development. Mol. Cell. Biol. 21, 4330–4336 (2001).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Ng, R. K. et al. Epigenetic restriction of embryonic cell lineage fate by methylation of Elf5. Nature Cell Biol. 10, 1280–1290 (2008).
Torres-Padilla, M. E., Parfitt, D. E., Kouzarides, T. & Zernicka-Goetz, M. Histone arginine methylation regulates pluripotency in the early mouse embryo. Nature 445, 214–218 (2007).
Piotrowska-Nitsche, K., Perea-Gomez, A., Haraguchi, S. & Zernicka-Goetz, M. Four-cell stage mouse blastomeres have different developmental properties. Development 132, 479–490 (2005).
Burton, A. et al. Single-cell profiling of epigenetic modifiers identifies PRDM14 as an inducer of cell fate in the mammalian embryo. Cell Rep. 5, 687–701 (2013).
Yamaji, M. et al. Critical function of Prdm14 for the establishment of the germ cell lineage in mice. Nature Genet. 40, 1016–1022 (2008).
Kurimoto, K. et al. An improved single-cell cDNA amplification method for efficient high-density oligonucleotide microarray analysis. Nucleic Acids Res. 34, e42 (2006).
Bultman, S. J. et al. Maternal BRG1 regulates zygotic genome activation in the mouse. Genes Dev. 20, 1744–1754 (2006).
Posfai, E. et al. Polycomb function during oogenesis is required for mouse embryonic development. Genes Dev. 26, 920–932 (2012).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Clarke, H. J., Oblin, C. & Bustin, M. Developmental regulation of chromatin composition during mouse embryogenesis: somatic histone H1 is first detectable at the 4-cell stage. Development 115, 791–799 (1992).
Fu, G. et al. Mouse oocytes and early embryos express multiple histone H1 subtypes. Biol. Reprod. 68, 1569–1576 (2003).
O'Neill, L. P., VerMilyea, M. D. & Turner, B. M. Epigenetic characterization of the early embryo with a chromatin immunoprecipitation protocol applicable to small cell populations. Nature Genet. 38, 835–841 (2006).
VerMilyea, M. D., O'Neill, L. P. & Turner, B. M. Transcription-independent heritability of induced histone modifications in the mouse preimplantation embryo. PLoS ONE 4, e6086 (2009).
Dahl, J. A., Reiner, A. H., Klungland, A., Wakayama, T. & Collas, P. Histone H3 lysine 27 methylation asymmetry on developmentally-regulated promoters distinguish the first two lineages in mouse preimplantation embryos. PLoS ONE 5, e9150 (2010).
Fadloun, A. et al. Chromatin signatures and retrotransposon profiling in mouse embryos reveal regulation of LINE-1 by RNA. Nature Struct. Mol. Biol. 20, 332–338 (2013).
Borgel, J. et al. Targets and dynamics of promoter DNA methylation during early mouse development. Nature Genet. 42, 1093–1100 (2010). Used MeDIP followed by microarray analysis to demonstrate selective promoter remethylation before implantation for the first time.
Waddington, C. H. The Strategy of the Genes (George Allen & Unwin, 1953).
MacArthur, B. D. & Lemischka, I. R. Statistical mechanics of pluripotency. Cell 154, 484–489 (2013).
Ohnishi, Y. et al. Cell-to-cell expression variability followed by signal reinforcement progressively segregates early mouse lineages. Nature Cell Biol. 16, 27–37 (2014).
Torres-Padilla, M. E. & Chambers, I. Transcription factor heterogeneity in pluripotent stem cells: a stochastic advantage. Development 141, 2173–2181 (2014).
Tang, F. et al. Deterministic and stochastic allele specific gene expression in single mouse blastomeres. PLoS ONE 6, e21208 (2010).
Lorthongpanich, C., Doris, T. P., Limviphuvadh, V., Knowles, B. B. & Solter, D. Developmental fate and lineage commitment of singled mouse blastomeres. Development 139, 3722–3731 (2012).
Chazaud, C., Yamanaka, Y., Pawson, T. & Rossant, J. Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev. Cell 10, 615–624 (2006).
Plusa, B., Piliszek, A., Frankenberg, S., Artus, J. & Hadjantonakis, A. K. Distinct sequential cell behaviours direct primitive endoderm formation in the mouse blastocyst. Development 135, 3081–3091 (2008).
Yan, L. et al. Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells. Nature Struct. Mol. Biol. 20, 1131–1139 (2013).
Xue, Z. et al. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500, 593–597 (2013).
Plessy, C. et al. Linking promoters to functional transcripts in small samples with nanoCAGE and CAGEscan. Nature Methods 7, 528–534 (2010).
Tarkowski, A. K. & Wroblewska, J. Development of blastomeres of mouse eggs isolated at the 4- and 8-cell stage. J. Embryol. Exp. Morphol. 18, 155–180 (1967).
Lorthongpanich, C. et al. Single-cell DNA-methylation analysis reveals epigenetic chimerism in preimplantation embryos. Science 341, 1110–1112 (2013). Analysed single-cell DNA methylation and applied it to studying defects in epigenetic inheritance.
Coulon, A., Chow, C. C., Singer, R. H. & Larson, D. R. Eukaryotic transcriptional dynamics: from single molecules to cell populations. Nature Rev. Genet. 14, 572–584 (2013).
Boskovic, A. et al. Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev. 28, 1042–1047 (2014).
Plachta, N., Bollenbach, T., Pease, S., Fraser, S. E. & Pantazis, P. Oct4 kinetics predict cell lineage patterning in the early mammalian embryo. Nature Cell Biol. 13, 117–123 (2011). Demonstrated that the chromatin-binding patterns of a transcription factor correlate with and precede lineage allocation.
Kaur, G. et al. Probing transcription factor diffusion dynamics in the living mammalian embryo with photoactivatable fluorescence correlation spectroscopy. Nature Commun. 4, 1637 (2013).
Miyanari, Y., Ziegler-Birling, C. & Torres-Padilla, M. E. Live visualization of chromatin dynamics with fluorescent TALEs. Nature Struct. Mol. Biol. 20, 1321–1324 (2013).
de Laat, W. & Duboule, D. Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502, 499–506 (2013).
Kind, J. & van Steensel, B. Genome-nuclear lamina interactions and gene regulation. Curr. Opin. Cell Biol. 22, 320–325 (2010).
Filion, G. J. et al. Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143, 212–224 (2010).
Pinheiro, I. et al. Prdm3 and Prdm16 are H3K9me1 methyltransferases required for mammalian heterochromatin integrity. Cell 150, 948–960 (2012).
Finlan, L. E. et al. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 4, e1000039 (2008).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Kind, J. et al. Single-cell dynamics of genome–nuclear lamina interactions. Cell 153, 178–192 (2013). Describes the mapping of LADs in single cells over time using a modified Dam-ID protocol.
Feng, S., Jacobsen, S. E. & Reik, W. Epigenetic reprogramming in plant and animal development. Science 330, 622–627 (2010).
Gurdon, J. B., Byrne, J. A. & Simonsson, S. Nuclear reprogramming and stem cell creation. Proc. Natl Acad. Sci. USA 100 (Suppl. 1), 11819–11822 (2003).
Pasque, V., Jullien, J., Miyamoto, K., Halley-Stott, R. P. & Gurdon, J. B. Epigenetic factors influencing resistance to nuclear reprogramming. Trends Genet. 27, 516–525 (2011).
Narbonne, P., Miyamoto, K. & Gurdon, J. B. Reprogramming and development in nuclear transfer embryos and in interspecific systems. Curr. Opin. Genet. Dev. 22, 450–458 (2012).
Huangfu, D. et al. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nature Biotech. 26, 795–797 (2008).
Antony, J., Oback, F., Chamley, L. W., Oback, B. & Laible, G. Transient JMJD2B-mediated reduction of H3K9me3 levels improves reprogramming of embryonic stem cells into cloned embryos. Mol. Cell. Biol. 33, 974–983 (2013).
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nature Genet. 45, 34–42 (2013).
Yang, C. X. et al. Heterochromatin reprogramming in rabbit embryos after fertilization, intra-, and inter-species SCNT correlates with preimplantation development. Reproduction 145, 149–159 (2013).
Jullien, J. et al. HIRA dependent H3.3 deposition is required for transcriptional reprogramming following nuclear transfer to Xenopus oocytes. Epigenet. Chromatin 5, 17 (2012).
Wen, D. et al. Histone variant H3.3 is an essential maternal factor for oocyte reprogramming. Proc. Natl Acad. Sci. USA 111, 7325–7330 (2014).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Tsukamoto, T. et al. Visualization of gene activity in living cells. Nature Cell Biol. 2, 871–878 (2000).
Heun, P., Laroche, T., Shimada, K., Furrer, P. & Gasser, S. M. Chromosome dynamics in the yeast interphase nucleus. Science 294, 2181–2186 (2001).
Soutoglou, E. et al. Positional stability of single double-strand breaks in mammalian cells. Nature Cell Biol. 9, 675–682 (2007).
Thanisch, K. et al. Targeting and tracing of specific DNA sequences with dTALEs in living cells. Nucleic Acids Res. 42, e38 (2014).
Ma, H., Reyes-Gutierrez, P. & Pederson, T. Visualization of repetitive DNA sequences in human chromosomes with transcription activator-like effectors. Proc. Natl Acad. Sci. USA 110, 21048–21053 (2013).
Yuan, K., Shermoen, A. W. & O'Farrell, P. H. Illuminating DNA replication during Drosophila development using TALE-lights. Curr. Biol. 24, R144–R145 (2014).
Saad, H. et al. DNA dynamics during early double-strand break processing revealed by non-intrusive imaging of living cells. PLoS Genet. 10, e1004187 (2014).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Acknowledgements
M.E.T.-P. acknowledges funding from EpiGeneSys NoE, ERC-Stg 'NuclearPotency', the FP7 Marie-Curie Actions ITN Nucleosome4D, the EMBO YIP and the Fondation Schlumberger pour l'Education et la Recherche. A.B. was a recipient of a fellowship from the Fondation pour la Recherche Médicale.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Totipotent
-
Totipotent cells are unique to the early embryo and have an unlimited potential to differentiate to the three germ layers of the embryo as well as to the extra-embryonic tissues.
- Blastocyst
-
The stage in mammalian development in which the embryo contains a fluid-filled cavity called the blastocoel.
- Trophectoderm
-
The first differentiated cell type that forms the outer layer of the blastocyst, which gives rise to extra-embryonic tissues that support the developing embryo.
- Inner cell mass
-
(ICM). Cells in the interior of the blastocyst that give rise to all tissues of the embryo and are the source of embryonic stem cells. The ICM is completely surrounded by the trophectoderm cells.
- Pluripotent
-
Pluripotent cells have the potential to differentiate to the three germ layers of the embryo: the endoderm, ectoderm and mesoderm.
- Histone variants
-
Non-allelic variants of the canonical histone proteins that differ in their protein structure, possess 5′ and 3′ untranslated regions, and are not restricted in their expression and incorporation into chromatin to the S phase of the cell cycle.
- Cell plasticity
-
The ability of a cell to change state or fate, whether by differentiation, reprogramming or any other sort of transformation.
- Regulative
-
A term used to describe developmental progression in which cells remain plastic and their fates are not determined from an early stage.
- Blastomeres
-
Cells of the pre-implantation embryo.
- Blastocyst asymmetry
-
The two initial cell types that exist within the blastocyst: the trophectoderm cells forming the outer trophoblast layer that surrounds the cavity, and the inner cell mass.
- Epiblast
-
An embryonic compartment derived from the inner cell mass. It gives rise to the embryo proper and differentiates to form the three layers of the developing embryo: ectoderm, endoderm and mesoderm during gastrulation.
- Primitive endoderm
-
A cell lineage derived from the inner cell mass that generates primarily extra-embryonic tissues, which will constitute the embryonic part of the placenta.
- Embryonic stem cells
-
(ES cells). Pluripotent stem cells derived from the inner cell mass that can be cultured in vitro indefinitely and differentiated into all three germ layers.
- Trophoblast stem cells
-
Stem cells that are derived from the polar trophectoderm of pre-implantation embryos and retain the capacity to differentiate in vitro into all trophoblast derivatives of the placenta.
- RNA fluorescence in situ hybridization
-
(RNA-FISH). An approach for studying the localization of nascent transcripts. Single-molecule RNA-FISH is a quantitative adaptation of RNA-FISH.
- Bisulphite sequencing
-
A technique for analysing sequence-specific methylated cytosines, based on their specific resistance to bisulphite conversion.
- Reduced representation bisulphite sequencing
-
(RRBS). A variant of bisulphite sequencing that is used to analyse methylation patterns at specific loci with high CpG content.
- TET enzymes
-
Ten-eleven translocation methylcytosine dioxygenase enzymes that catalyse the conversion of 5-methylcytosine to 5-hydroxymethylcytosine by oxidation.
- Protamines
-
A group of small, highly basic proteins associated with the DNA, particularly in sperm, in place of histones.
- Polycomb repressive complex 2
-
(PRC2). A di- and trimethyl-transferase complex. Its substrate is Lys27 of histone H3, a mark of facultative heterochromatin.
- Major satellite
-
(Also known as a gamma satellite). An extensive region of tandem DNA repeats. Major satellites are normally found at pericentromeres and are mostly AT-rich.
- Embryonic genome activation
-
(EGA). The process by which the embryonic genome begins to transcribe the major portion of its genome.
- Chromocentres
-
Irregular, densely stained aggregations of DNA, consisting of heterochromatic, centromeric and pericentromeric regions.
- Nucleolar-like bodies
-
(NLBs). Spherical bodies of uncertain structure and function that are unique to the pre-implantation embryo of mammals and are thought to be the non-functional precursors of nucleoli.
- Bivalent domains
-
Regions of chromatin that contain both activating and repressive chromatin marks coincidently, notably trimethylated histone H3 Lys4 (H3K4me3) and H3K27me3.
- Carrier-chromatin immunoprecipitation
-
(CChIP). A chromatin immunoprecipitation approach of native chromatin (prepared by nuclease digestion as opposed to crosslinking) modified for low sample quantities.
- CpG islands
-
Regions in the genome that contain a high frequency of CpG dinucleotides. They are often found in gene promoters.
- Methylated DNA immunoprecipitation
-
(MeDIP). A genome-wide, high-resolution approach to quantifying DNA methylation. The antibody used for the precipitation recognizes 5-methylcytosine.
- Microfluidics
-
A technology that enables the analysis of a set of transcripts (typically 48 or 96) in single cells by quantitative PCR in nanolitre volumes, allowing for truly quantitative information on gene expression to be extracted.
- Cap analysis of gene expression
-
(CAGE). A technique for capturing mRNAs by the addition of linkers at their 5′ end. It provides information on the 5′ end of a transcript and therefore on its transcription start sites.
- Fluorescence recovery after photobleaching
-
(FRAP). This technique uses fluorescently tagged proteins of interest. The recovery of the fluorescent signal after bleaching can provide information on the mobility of the protein of interest.
- Fluorescence decay after photoactivation
-
(FDAP). This technique uses a photoactivatable fluorescent molecule tagged to the protein of interest. The loss of fluorescence after photoactivation is then measured over time to determine protein mobility parameters.
- Fluorescence correlation spectroscopy
-
(FCS). This is a correlation analysis of fluorescence fluctuations to study the concentration and dynamics of a fluorescently tagged molecule. In combination with photoactivation (paFCS) it provides information on the number of molecules analysed.
- Transcription activator-like effector
-
(TALE). A protein with hypervariable domains that recognize specific DNA bases, allowing sequence-specific targeting.
- Chromosome conformation capture
-
(3C). A technique for studying genomic organization, in which nuclei are fixed, the DNA is digested and chromosomal regions in physical proximity are ligated and identified by PCR.
- DNA adenine methyltransferase identification
-
(Dam-ID). A technique for mapping protein binding to DNA by fusing proteins to a bacterial adenine methylase, which is not endogenous to eukaryotes. Binding can be mapped based on the position of methylated adenines.
- Lamina-associated domains
-
(LADs). Regions of the genome that have been demonstrated to interact with the nuclear lamina.
- Hi-C
-
A variation of chromosome conformation capture in which all interacting regions of the genome can be mapped by high-throughput sequencing of the ligated products.
Rights and permissions
About this article
Cite this article
Burton, A., Torres-Padilla, ME. Chromatin dynamics in the regulation of cell fate allocation during early embryogenesis. Nat Rev Mol Cell Biol 15, 723–735 (2014). https://doi.org/10.1038/nrm3885
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3885
This article is cited by
-
Emergence of replication timing during early mammalian development
Nature (2024)
-
ATAC-seq and RNA-seq analysis unravel the mechanism of sex differentiation and infertility in sex reversal chicken
Epigenetics & Chromatin (2023)
-
Dynamic chromatin regulatory programs during embryogenesis of hexaploid wheat
Genome Biology (2023)
-
The composition dynamics of transposable elements in human blastocysts
Journal of Human Genetics (2023)
-
Epitranscriptomics in metabolic disease
Nature Metabolism (2023)